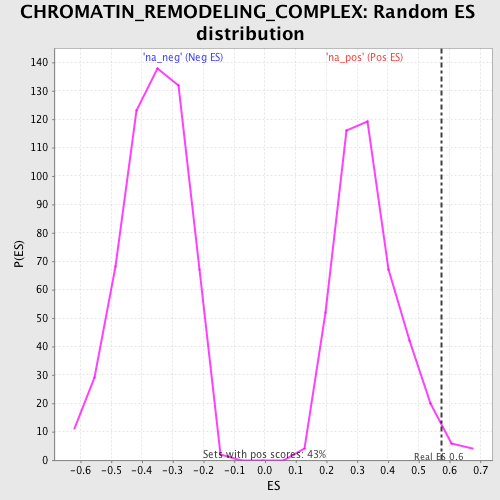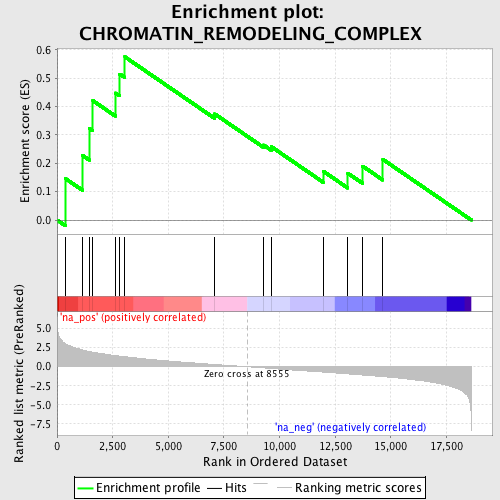
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | set04_transDMpreB_versus_DMpreB |
| Phenotype | NoPhenotypeAvailable |
| Upregulated in class | na_pos |
| GeneSet | CHROMATIN_REMODELING_COMPLEX |
| Enrichment Score (ES) | 0.57493377 |
| Normalized Enrichment Score (NES) | 1.7047026 |
| Nominal p-value | 0.023255814 |
| FDR q-value | 0.6627563 |
| FWER p-Value | 0.995 |

| PROBE | GENE SYMBOL | GENE_TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|---|
| 1 | ARID1A | 388 | 2.976 | 0.1453 | Yes | ||
| 2 | CHAF1A | 1146 | 2.195 | 0.2272 | Yes | ||
| 3 | RSF1 | 1446 | 1.998 | 0.3227 | Yes | ||
| 4 | NAP1L1 | 1607 | 1.907 | 0.4206 | Yes | ||
| 5 | SMARCD1 | 2619 | 1.457 | 0.4476 | Yes | ||
| 6 | NAP1L4 | 2816 | 1.383 | 0.5143 | Yes | ||
| 7 | NCR1 | 3046 | 1.307 | 0.5749 | Yes | ||
| 8 | ASF1A | 7085 | 0.284 | 0.3737 | No | ||
| 9 | CHAF1B | 9280 | -0.155 | 0.2644 | No | ||
| 10 | SMARCA5 | 9626 | -0.226 | 0.2585 | No | ||
| 11 | SMARCC1 | 11960 | -0.701 | 0.1722 | No | ||
| 12 | SMARCB1 | 13068 | -0.935 | 0.1649 | No | ||
| 13 | NAP1L3 | 13733 | -1.085 | 0.1898 | No | ||
| 14 | NAP1L2 | 14643 | -1.301 | 0.2136 | No |