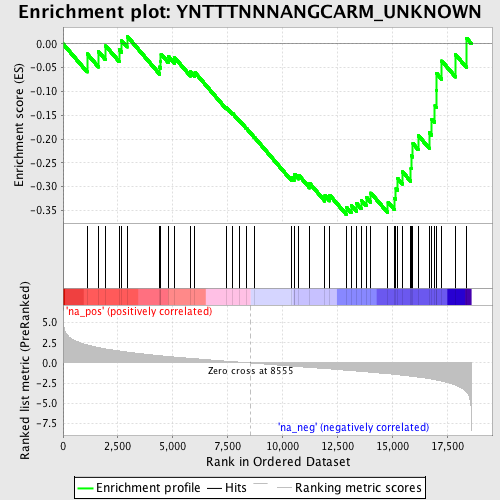
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | set04_transDMpreB_versus_DMpreB |
| Phenotype | NoPhenotypeAvailable |
| Upregulated in class | na_neg |
| GeneSet | YNTTTNNNANGCARM_UNKNOWN |
| Enrichment Score (ES) | -0.35835743 |
| Normalized Enrichment Score (NES) | -1.3578998 |
| Nominal p-value | 0.07692308 |
| FDR q-value | 0.6101154 |
| FWER p-Value | 1.0 |

| PROBE | GENE SYMBOL | GENE_TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|---|
| 1 | FGF14 | 1096 | 2.242 | -0.0208 | No | ||
| 2 | DMD | 1614 | 1.903 | -0.0162 | No | ||
| 3 | C1S | 1940 | 1.738 | -0.0041 | No | ||
| 4 | OMG | 2559 | 1.484 | -0.0121 | No | ||
| 5 | KLHL5 | 2671 | 1.440 | 0.0065 | No | ||
| 6 | LEMD2 | 2914 | 1.346 | 0.0164 | No | ||
| 7 | GABRA1 | 4403 | 0.898 | -0.0484 | No | ||
| 8 | MSX2 | 4452 | 0.883 | -0.0360 | No | ||
| 9 | GABRG1 | 4460 | 0.880 | -0.0213 | No | ||
| 10 | SYNPR | 4794 | 0.798 | -0.0257 | No | ||
| 11 | PDE4D | 5085 | 0.727 | -0.0289 | No | ||
| 12 | KCTD12 | 5805 | 0.569 | -0.0579 | No | ||
| 13 | NRF1 | 6010 | 0.521 | -0.0600 | No | ||
| 14 | KIF13A | 7436 | 0.212 | -0.1331 | No | ||
| 15 | TLL2 | 7720 | 0.162 | -0.1456 | No | ||
| 16 | NRK | 8037 | 0.102 | -0.1609 | No | ||
| 17 | B3GNT7 | 8354 | 0.043 | -0.1772 | No | ||
| 18 | VAX1 | 8736 | -0.038 | -0.1971 | No | ||
| 19 | EYA1 | 10412 | -0.397 | -0.2805 | No | ||
| 20 | LRRTM1 | 10529 | -0.418 | -0.2796 | No | ||
| 21 | EVX1 | 10547 | -0.422 | -0.2733 | No | ||
| 22 | SMPX | 10744 | -0.463 | -0.2760 | No | ||
| 23 | ETV5 | 11242 | -0.563 | -0.2932 | No | ||
| 24 | ETV6 | 11933 | -0.696 | -0.3185 | No | ||
| 25 | GPM6A | 12149 | -0.734 | -0.3175 | No | ||
| 26 | DNAH5 | 12908 | -0.901 | -0.3430 | Yes | ||
| 27 | GCDH | 13130 | -0.948 | -0.3387 | Yes | ||
| 28 | ELAVL2 | 13378 | -1.004 | -0.3349 | Yes | ||
| 29 | CNN3 | 13596 | -1.053 | -0.3287 | Yes | ||
| 30 | FGF12 | 13816 | -1.104 | -0.3216 | Yes | ||
| 31 | MAFB | 14025 | -1.154 | -0.3132 | Yes | ||
| 32 | SFRP2 | 14806 | -1.340 | -0.3323 | Yes | ||
| 33 | GNAI1 | 15109 | -1.414 | -0.3245 | Yes | ||
| 34 | FGD4 | 15170 | -1.430 | -0.3034 | Yes | ||
| 35 | VDR | 15254 | -1.452 | -0.2831 | Yes | ||
| 36 | HCRTR2 | 15469 | -1.524 | -0.2686 | Yes | ||
| 37 | GPR158 | 15852 | -1.646 | -0.2611 | Yes | ||
| 38 | CDX2 | 15867 | -1.650 | -0.2338 | Yes | ||
| 39 | GRK5 | 15943 | -1.676 | -0.2092 | Yes | ||
| 40 | KCNA2 | 16185 | -1.760 | -0.1922 | Yes | ||
| 41 | HOXC4 | 16696 | -1.962 | -0.1862 | Yes | ||
| 42 | DCX | 16804 | -2.008 | -0.1578 | Yes | ||
| 43 | HOXC5 | 16947 | -2.080 | -0.1300 | Yes | ||
| 44 | ADAMTS5 | 17015 | -2.113 | -0.0976 | Yes | ||
| 45 | PVRL1 | 17028 | -2.121 | -0.0621 | Yes | ||
| 46 | HOXB6 | 17258 | -2.263 | -0.0358 | Yes | ||
| 47 | IL10 | 17887 | -2.759 | -0.0226 | Yes | ||
| 48 | LDB2 | 18395 | -3.626 | 0.0119 | Yes |
