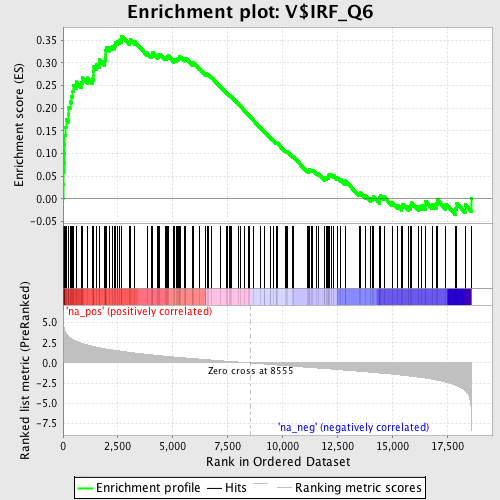
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | set04_transDMpreB_versus_DMpreB |
| Phenotype | NoPhenotypeAvailable |
| Upregulated in class | na_pos |
| GeneSet | V$IRF_Q6 |
| Enrichment Score (ES) | 0.3589613 |
| Normalized Enrichment Score (NES) | 1.778473 |
| Nominal p-value | 0.0 |
| FDR q-value | 0.073692024 |
| FWER p-Value | 0.232 |

| PROBE | GENE SYMBOL | GENE_TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|---|
| 1 | CCND1 | 2 | 6.319 | 0.0319 | Yes | ||
| 2 | LMO2 | 11 | 5.459 | 0.0592 | Yes | ||
| 3 | ITGB7 | 41 | 4.403 | 0.0800 | Yes | ||
| 4 | BCL11A | 59 | 4.158 | 0.1001 | Yes | ||
| 5 | ISGF3G | 64 | 4.085 | 0.1207 | Yes | ||
| 6 | LSP1 | 70 | 4.026 | 0.1408 | Yes | ||
| 7 | MAP3K11 | 104 | 3.838 | 0.1585 | Yes | ||
| 8 | DAPP1 | 133 | 3.682 | 0.1757 | Yes | ||
| 9 | USF1 | 258 | 3.239 | 0.1854 | Yes | ||
| 10 | ADAR | 263 | 3.223 | 0.2015 | Yes | ||
| 11 | PIP5K2B | 315 | 3.124 | 0.2146 | Yes | ||
| 12 | RRBP1 | 377 | 2.990 | 0.2265 | Yes | ||
| 13 | MS4A1 | 443 | 2.905 | 0.2377 | Yes | ||
| 14 | NFATC1 | 467 | 2.872 | 0.2510 | Yes | ||
| 15 | FCGR2B | 593 | 2.703 | 0.2579 | Yes | ||
| 16 | PTPRO | 838 | 2.449 | 0.2571 | Yes | ||
| 17 | IL18BP | 885 | 2.408 | 0.2669 | Yes | ||
| 18 | EIF4A2 | 1099 | 2.238 | 0.2667 | Yes | ||
| 19 | LGI1 | 1333 | 2.056 | 0.2645 | Yes | ||
| 20 | MOV10 | 1382 | 2.029 | 0.2722 | Yes | ||
| 21 | KYNU | 1386 | 2.028 | 0.2823 | Yes | ||
| 22 | ZFPM1 | 1404 | 2.019 | 0.2916 | Yes | ||
| 23 | TNFSF15 | 1513 | 1.960 | 0.2957 | Yes | ||
| 24 | TCIRG1 | 1644 | 1.887 | 0.2983 | Yes | ||
| 25 | PDGFRB | 1659 | 1.880 | 0.3071 | Yes | ||
| 26 | BCOR | 1903 | 1.760 | 0.3028 | Yes | ||
| 27 | UPF2 | 1913 | 1.752 | 0.3112 | Yes | ||
| 28 | ARHGEF6 | 1931 | 1.742 | 0.3191 | Yes | ||
| 29 | CCNT2 | 1934 | 1.741 | 0.3279 | Yes | ||
| 30 | TBX1 | 1970 | 1.722 | 0.3347 | Yes | ||
| 31 | TLR7 | 2131 | 1.658 | 0.3345 | Yes | ||
| 32 | TLK2 | 2242 | 1.609 | 0.3367 | Yes | ||
| 33 | TOP1 | 2356 | 1.560 | 0.3385 | Yes | ||
| 34 | EREG | 2388 | 1.544 | 0.3446 | Yes | ||
| 35 | PTK2 | 2477 | 1.514 | 0.3475 | Yes | ||
| 36 | KCNQ4 | 2563 | 1.483 | 0.3504 | Yes | ||
| 37 | STAT6 | 2672 | 1.439 | 0.3519 | Yes | ||
| 38 | LGALS8 | 2677 | 1.435 | 0.3590 | Yes | ||
| 39 | IL22RA1 | 3014 | 1.315 | 0.3474 | No | ||
| 40 | SEMA3A | 3074 | 1.299 | 0.3508 | No | ||
| 41 | BBX | 3268 | 1.231 | 0.3466 | No | ||
| 42 | PCDH7 | 3825 | 1.051 | 0.3219 | No | ||
| 43 | PRDM16 | 4025 | 0.997 | 0.3161 | No | ||
| 44 | MIA2 | 4078 | 0.982 | 0.3183 | No | ||
| 45 | PTMA | 4082 | 0.981 | 0.3231 | No | ||
| 46 | FLI1 | 4323 | 0.920 | 0.3148 | No | ||
| 47 | MLLT3 | 4362 | 0.909 | 0.3173 | No | ||
| 48 | PITX2 | 4409 | 0.896 | 0.3194 | No | ||
| 49 | SELL | 4649 | 0.837 | 0.3107 | No | ||
| 50 | NLGN2 | 4719 | 0.817 | 0.3111 | No | ||
| 51 | MYOG | 4738 | 0.811 | 0.3143 | No | ||
| 52 | SLC39A7 | 4783 | 0.799 | 0.3159 | No | ||
| 53 | DGKA | 5048 | 0.736 | 0.3054 | No | ||
| 54 | IRF2 | 5074 | 0.730 | 0.3077 | No | ||
| 55 | SLC40A1 | 5181 | 0.707 | 0.3056 | No | ||
| 56 | GATA6 | 5190 | 0.705 | 0.3087 | No | ||
| 57 | CASP7 | 5238 | 0.693 | 0.3097 | No | ||
| 58 | GSDMDC1 | 5283 | 0.680 | 0.3107 | No | ||
| 59 | B2M | 5299 | 0.677 | 0.3134 | No | ||
| 60 | HPCAL1 | 5369 | 0.658 | 0.3130 | No | ||
| 61 | PLXNC1 | 5534 | 0.625 | 0.3073 | No | ||
| 62 | THBS1 | 5577 | 0.615 | 0.3081 | No | ||
| 63 | HTR2C | 5595 | 0.611 | 0.3103 | No | ||
| 64 | DCAMKL1 | 5880 | 0.553 | 0.2977 | No | ||
| 65 | FMR1 | 5913 | 0.545 | 0.2987 | No | ||
| 66 | DUSP10 | 5933 | 0.541 | 0.3005 | No | ||
| 67 | SIX1 | 6214 | 0.478 | 0.2877 | No | ||
| 68 | EHD1 | 6491 | 0.414 | 0.2749 | No | ||
| 69 | DNASE1L3 | 6498 | 0.413 | 0.2766 | No | ||
| 70 | CDH3 | 6568 | 0.398 | 0.2749 | No | ||
| 71 | MUSK | 6624 | 0.386 | 0.2739 | No | ||
| 72 | TAPBP | 6762 | 0.356 | 0.2683 | No | ||
| 73 | CDK6 | 7151 | 0.272 | 0.2487 | No | ||
| 74 | ARF3 | 7429 | 0.213 | 0.2347 | No | ||
| 75 | SLC2A5 | 7514 | 0.197 | 0.2312 | No | ||
| 76 | HOXB3 | 7595 | 0.185 | 0.2278 | No | ||
| 77 | NCF1 | 7645 | 0.175 | 0.2260 | No | ||
| 78 | RORB | 7671 | 0.172 | 0.2256 | No | ||
| 79 | IFNB1 | 7971 | 0.113 | 0.2099 | No | ||
| 80 | ECM2 | 8102 | 0.091 | 0.2034 | No | ||
| 81 | NPEPPS | 8283 | 0.059 | 0.1939 | No | ||
| 82 | IFIT2 | 8428 | 0.026 | 0.1863 | No | ||
| 83 | TWIST1 | 8516 | 0.006 | 0.1816 | No | ||
| 84 | CHST1 | 8665 | -0.025 | 0.1737 | No | ||
| 85 | BHLHB5 | 8981 | -0.086 | 0.1571 | No | ||
| 86 | STAG2 | 9164 | -0.127 | 0.1479 | No | ||
| 87 | KLHL13 | 9193 | -0.133 | 0.1470 | No | ||
| 88 | PURA | 9436 | -0.185 | 0.1349 | No | ||
| 89 | ISL1 | 9457 | -0.189 | 0.1347 | No | ||
| 90 | IFI35 | 9583 | -0.217 | 0.1291 | No | ||
| 91 | GJC1 | 9710 | -0.244 | 0.1235 | No | ||
| 92 | HIPK1 | 9732 | -0.249 | 0.1236 | No | ||
| 93 | PSMA5 | 9751 | -0.255 | 0.1239 | No | ||
| 94 | DLL1 | 10122 | -0.335 | 0.1056 | No | ||
| 95 | SLC15A3 | 10167 | -0.343 | 0.1049 | No | ||
| 96 | HOXB4 | 10212 | -0.353 | 0.1044 | No | ||
| 97 | PSMB8 | 10463 | -0.406 | 0.0929 | No | ||
| 98 | SOCS1 | 10482 | -0.409 | 0.0940 | No | ||
| 99 | NEDD4 | 11123 | -0.540 | 0.0621 | No | ||
| 100 | MSX1 | 11182 | -0.551 | 0.0617 | No | ||
| 101 | UBE2L6 | 11220 | -0.559 | 0.0626 | No | ||
| 102 | ETV5 | 11242 | -0.563 | 0.0643 | No | ||
| 103 | CYBB | 11308 | -0.576 | 0.0637 | No | ||
| 104 | MYB | 11377 | -0.589 | 0.0630 | No | ||
| 105 | CD80 | 11565 | -0.625 | 0.0560 | No | ||
| 106 | DIO2 | 11640 | -0.641 | 0.0553 | No | ||
| 107 | FUT8 | 11916 | -0.693 | 0.0439 | No | ||
| 108 | ETV6 | 11933 | -0.696 | 0.0466 | No | ||
| 109 | RARG | 11991 | -0.706 | 0.0471 | No | ||
| 110 | PSMB10 | 12046 | -0.713 | 0.0477 | No | ||
| 111 | TCF7L2 | 12099 | -0.725 | 0.0486 | No | ||
| 112 | HAPLN1 | 12104 | -0.726 | 0.0521 | No | ||
| 113 | BST2 | 12139 | -0.733 | 0.0540 | No | ||
| 114 | TCF15 | 12232 | -0.754 | 0.0528 | No | ||
| 115 | LY86 | 12331 | -0.778 | 0.0514 | No | ||
| 116 | CNTFR | 12503 | -0.813 | 0.0463 | No | ||
| 117 | GABARAP | 12638 | -0.845 | 0.0433 | No | ||
| 118 | ZNF385 | 12860 | -0.891 | 0.0359 | No | ||
| 119 | LZTS2 | 12876 | -0.894 | 0.0396 | No | ||
| 120 | ZBP1 | 13485 | -1.028 | 0.0119 | No | ||
| 121 | ADORA3 | 13552 | -1.041 | 0.0136 | No | ||
| 122 | ZADH2 | 13784 | -1.094 | 0.0067 | No | ||
| 123 | CXCR4 | 14014 | -1.151 | 0.0001 | No | ||
| 124 | GRIA3 | 14099 | -1.169 | 0.0015 | No | ||
| 125 | SEMA6D | 14157 | -1.185 | 0.0044 | No | ||
| 126 | LGALS9 | 14421 | -1.252 | -0.0035 | No | ||
| 127 | MAPK10 | 14428 | -1.253 | 0.0025 | No | ||
| 128 | F3 | 14447 | -1.257 | 0.0079 | No | ||
| 129 | SLC11A2 | 14632 | -1.298 | 0.0046 | No | ||
| 130 | TNFSF13B | 14990 | -1.387 | -0.0077 | No | ||
| 131 | ARHGAP5 | 15242 | -1.448 | -0.0140 | No | ||
| 132 | EGFL6 | 15442 | -1.515 | -0.0171 | No | ||
| 133 | EOMES | 15486 | -1.529 | -0.0116 | No | ||
| 134 | CXCL10 | 15726 | -1.603 | -0.0164 | No | ||
| 135 | CBX4 | 15846 | -1.644 | -0.0145 | No | ||
| 136 | FGF9 | 15892 | -1.659 | -0.0086 | No | ||
| 137 | SERHL | 16204 | -1.767 | -0.0164 | No | ||
| 138 | LGP2 | 16352 | -1.817 | -0.0152 | No | ||
| 139 | ESR1 | 16505 | -1.877 | -0.0139 | No | ||
| 140 | ADAM12 | 16528 | -1.884 | -0.0055 | No | ||
| 141 | PSMA3 | 16833 | -2.024 | -0.0117 | No | ||
| 142 | CASK | 17004 | -2.109 | -0.0102 | No | ||
| 143 | EXT1 | 17046 | -2.132 | -0.0016 | No | ||
| 144 | ADAM15 | 17449 | -2.398 | -0.0112 | No | ||
| 145 | PRKAG1 | 17877 | -2.749 | -0.0204 | No | ||
| 146 | NT5C3 | 17945 | -2.828 | -0.0097 | No | ||
| 147 | RTN4 | 18322 | -3.432 | -0.0126 | No | ||
| 148 | PRKACA | 18600 | -5.618 | 0.0009 | No |
