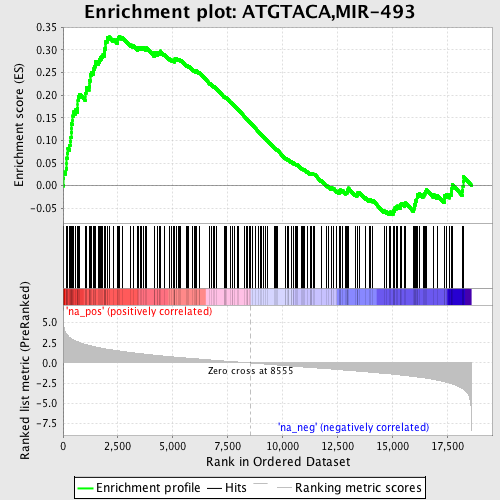
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | set04_transDMpreB_versus_DMpreB |
| Phenotype | NoPhenotypeAvailable |
| Upregulated in class | na_pos |
| GeneSet | ATGTACA,MIR-493 |
| Enrichment Score (ES) | 0.3306186 |
| Normalized Enrichment Score (NES) | 1.7167051 |
| Nominal p-value | 0.0 |
| FDR q-value | 0.06307589 |
| FWER p-Value | 0.363 |

| PROBE | GENE SYMBOL | GENE_TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|---|
| 1 | COBLL1 | 18 | 4.808 | 0.0158 | Yes | ||
| 2 | BCL2 | 39 | 4.461 | 0.0302 | Yes | ||
| 3 | GAD1 | 131 | 3.695 | 0.0381 | Yes | ||
| 4 | SVIL | 162 | 3.575 | 0.0489 | Yes | ||
| 5 | NAB1 | 167 | 3.555 | 0.0611 | Yes | ||
| 6 | UBE3A | 193 | 3.467 | 0.0718 | Yes | ||
| 7 | KIF1B | 217 | 3.381 | 0.0823 | Yes | ||
| 8 | PHF12 | 298 | 3.147 | 0.0889 | Yes | ||
| 9 | CUGBP2 | 320 | 3.105 | 0.0986 | Yes | ||
| 10 | RBBP6 | 352 | 3.032 | 0.1074 | Yes | ||
| 11 | UBE2Q1 | 370 | 3.002 | 0.1170 | Yes | ||
| 12 | HMGCS1 | 381 | 2.984 | 0.1268 | Yes | ||
| 13 | ARID1A | 388 | 2.976 | 0.1368 | Yes | ||
| 14 | E2F6 | 439 | 2.912 | 0.1442 | Yes | ||
| 15 | PIP5K3 | 446 | 2.900 | 0.1540 | Yes | ||
| 16 | KLF3 | 465 | 2.873 | 0.1630 | Yes | ||
| 17 | FBS1 | 559 | 2.742 | 0.1675 | Yes | ||
| 18 | MCL1 | 659 | 2.620 | 0.1713 | Yes | ||
| 19 | AXIN1 | 674 | 2.610 | 0.1796 | Yes | ||
| 20 | TBK1 | 676 | 2.610 | 0.1886 | Yes | ||
| 21 | BRD7 | 693 | 2.592 | 0.1968 | Yes | ||
| 22 | EPHA7 | 749 | 2.523 | 0.2025 | Yes | ||
| 23 | ANKRD17 | 1005 | 2.315 | 0.1967 | Yes | ||
| 24 | ANTXR1 | 1006 | 2.314 | 0.2048 | Yes | ||
| 25 | SPEN | 1047 | 2.290 | 0.2106 | Yes | ||
| 26 | CREBBP | 1062 | 2.275 | 0.2177 | Yes | ||
| 27 | BAZ2A | 1182 | 2.168 | 0.2188 | Yes | ||
| 28 | RSBN1 | 1202 | 2.156 | 0.2253 | Yes | ||
| 29 | MEF2C | 1205 | 2.154 | 0.2327 | Yes | ||
| 30 | TMEFF1 | 1235 | 2.130 | 0.2385 | Yes | ||
| 31 | UBE2V2 | 1236 | 2.130 | 0.2459 | Yes | ||
| 32 | BHLHB3 | 1295 | 2.085 | 0.2500 | Yes | ||
| 33 | ATP2A2 | 1370 | 2.038 | 0.2531 | Yes | ||
| 34 | DOCK9 | 1400 | 2.020 | 0.2586 | Yes | ||
| 35 | PMP22 | 1427 | 2.007 | 0.2641 | Yes | ||
| 36 | PHF2 | 1463 | 1.991 | 0.2692 | Yes | ||
| 37 | PTPN12 | 1484 | 1.981 | 0.2750 | Yes | ||
| 38 | ATAD2 | 1629 | 1.893 | 0.2737 | Yes | ||
| 39 | DACH1 | 1656 | 1.881 | 0.2789 | Yes | ||
| 40 | TLE4 | 1692 | 1.857 | 0.2834 | Yes | ||
| 41 | CHKB | 1765 | 1.824 | 0.2859 | Yes | ||
| 42 | HIC2 | 1816 | 1.801 | 0.2894 | Yes | ||
| 43 | PCDH18 | 1877 | 1.772 | 0.2923 | Yes | ||
| 44 | RNF38 | 1885 | 1.766 | 0.2981 | Yes | ||
| 45 | BCOR | 1903 | 1.760 | 0.3033 | Yes | ||
| 46 | CCNT2 | 1934 | 1.741 | 0.3077 | Yes | ||
| 47 | CDC14A | 1936 | 1.740 | 0.3137 | Yes | ||
| 48 | CARM1 | 1946 | 1.732 | 0.3192 | Yes | ||
| 49 | SORCS3 | 2004 | 1.710 | 0.3221 | Yes | ||
| 50 | WNT5A | 2012 | 1.708 | 0.3277 | Yes | ||
| 51 | PFN2 | 2097 | 1.676 | 0.3289 | Yes | ||
| 52 | CPNE8 | 2302 | 1.584 | 0.3234 | Yes | ||
| 53 | FBXO33 | 2482 | 1.511 | 0.3189 | Yes | ||
| 54 | VAMP2 | 2485 | 1.509 | 0.3240 | Yes | ||
| 55 | HNRPA0 | 2505 | 1.504 | 0.3282 | Yes | ||
| 56 | ITGB1 | 2557 | 1.485 | 0.3306 | Yes | ||
| 57 | SALL1 | 2691 | 1.431 | 0.3284 | No | ||
| 58 | ITSN2 | 3091 | 1.295 | 0.3112 | No | ||
| 59 | SOX21 | 3189 | 1.258 | 0.3103 | No | ||
| 60 | EIF4ENIF1 | 3401 | 1.191 | 0.3030 | No | ||
| 61 | SERTAD2 | 3432 | 1.181 | 0.3054 | No | ||
| 62 | DNAJC13 | 3512 | 1.155 | 0.3052 | No | ||
| 63 | FZD4 | 3581 | 1.132 | 0.3054 | No | ||
| 64 | RYBP | 3651 | 1.109 | 0.3055 | No | ||
| 65 | CLASP1 | 3765 | 1.072 | 0.3031 | No | ||
| 66 | RIMS3 | 3804 | 1.057 | 0.3047 | No | ||
| 67 | PPP1R10 | 4160 | 0.960 | 0.2888 | No | ||
| 68 | CRSP7 | 4162 | 0.959 | 0.2921 | No | ||
| 69 | ZMYND19 | 4166 | 0.958 | 0.2952 | No | ||
| 70 | BCL7A | 4296 | 0.927 | 0.2914 | No | ||
| 71 | NCOA1 | 4311 | 0.923 | 0.2939 | No | ||
| 72 | PDPK1 | 4391 | 0.901 | 0.2927 | No | ||
| 73 | GABRA1 | 4403 | 0.898 | 0.2953 | No | ||
| 74 | BTG1 | 4415 | 0.893 | 0.2978 | No | ||
| 75 | THRAP1 | 4604 | 0.845 | 0.2905 | No | ||
| 76 | TNFSF11 | 4842 | 0.785 | 0.2803 | No | ||
| 77 | KCNMA1 | 4936 | 0.763 | 0.2779 | No | ||
| 78 | POU4F1 | 5020 | 0.743 | 0.2760 | No | ||
| 79 | IRF2 | 5074 | 0.730 | 0.2757 | No | ||
| 80 | PDE4D | 5085 | 0.727 | 0.2777 | No | ||
| 81 | THRAP2 | 5094 | 0.725 | 0.2797 | No | ||
| 82 | VPS13D | 5098 | 0.725 | 0.2821 | No | ||
| 83 | EFTUD1 | 5167 | 0.709 | 0.2809 | No | ||
| 84 | SNRP70 | 5245 | 0.691 | 0.2791 | No | ||
| 85 | UNC50 | 5282 | 0.680 | 0.2795 | No | ||
| 86 | KCNIP4 | 5363 | 0.659 | 0.2774 | No | ||
| 87 | SV2B | 5642 | 0.603 | 0.2644 | No | ||
| 88 | BASP1 | 5656 | 0.600 | 0.2658 | No | ||
| 89 | DHX36 | 5732 | 0.588 | 0.2638 | No | ||
| 90 | OSBPL6 | 5908 | 0.546 | 0.2562 | No | ||
| 91 | DAB2IP | 5981 | 0.528 | 0.2541 | No | ||
| 92 | FOXO1A | 6042 | 0.516 | 0.2526 | No | ||
| 93 | SIN3A | 6072 | 0.509 | 0.2528 | No | ||
| 94 | ATP5A1 | 6077 | 0.508 | 0.2544 | No | ||
| 95 | PDZRN3 | 6213 | 0.478 | 0.2487 | No | ||
| 96 | RBM12 | 6221 | 0.476 | 0.2500 | No | ||
| 97 | MBNL2 | 6693 | 0.370 | 0.2257 | No | ||
| 98 | PDGFC | 6759 | 0.357 | 0.2234 | No | ||
| 99 | USP21 | 6863 | 0.333 | 0.2189 | No | ||
| 100 | DDX3X | 6889 | 0.328 | 0.2187 | No | ||
| 101 | NXPH1 | 6975 | 0.309 | 0.2152 | No | ||
| 102 | MTPN | 7332 | 0.236 | 0.1966 | No | ||
| 103 | IRF7 | 7369 | 0.227 | 0.1955 | No | ||
| 104 | CUL5 | 7380 | 0.224 | 0.1957 | No | ||
| 105 | NMT1 | 7420 | 0.215 | 0.1943 | No | ||
| 106 | PIK3R1 | 7425 | 0.214 | 0.1949 | No | ||
| 107 | SDC2 | 7649 | 0.175 | 0.1833 | No | ||
| 108 | SNX9 | 7702 | 0.167 | 0.1811 | No | ||
| 109 | HEMGN | 7707 | 0.165 | 0.1815 | No | ||
| 110 | RPE | 7823 | 0.143 | 0.1757 | No | ||
| 111 | TTC7B | 7933 | 0.120 | 0.1702 | No | ||
| 112 | SPRYD3 | 7973 | 0.113 | 0.1685 | No | ||
| 113 | PSD2 | 8279 | 0.060 | 0.1521 | No | ||
| 114 | DKK2 | 8348 | 0.045 | 0.1486 | No | ||
| 115 | TJP1 | 8381 | 0.039 | 0.1470 | No | ||
| 116 | PAIP1 | 8384 | 0.038 | 0.1470 | No | ||
| 117 | ARFGEF1 | 8488 | 0.011 | 0.1414 | No | ||
| 118 | AK5 | 8541 | 0.002 | 0.1386 | No | ||
| 119 | GABRA5 | 8613 | -0.014 | 0.1348 | No | ||
| 120 | JARID1B | 8777 | -0.045 | 0.1261 | No | ||
| 121 | SIPA1L2 | 8885 | -0.067 | 0.1205 | No | ||
| 122 | BHLHB5 | 8981 | -0.086 | 0.1156 | No | ||
| 123 | ESRRG | 9060 | -0.103 | 0.1118 | No | ||
| 124 | OTUB1 | 9132 | -0.121 | 0.1083 | No | ||
| 125 | EIF2C1 | 9211 | -0.138 | 0.1046 | No | ||
| 126 | ZIC1 | 9310 | -0.161 | 0.0998 | No | ||
| 127 | NLGN1 | 9644 | -0.230 | 0.0825 | No | ||
| 128 | SP4 | 9686 | -0.238 | 0.0811 | No | ||
| 129 | HIPK1 | 9732 | -0.249 | 0.0795 | No | ||
| 130 | SPRED2 | 9760 | -0.258 | 0.0789 | No | ||
| 131 | DLL1 | 10122 | -0.335 | 0.0605 | No | ||
| 132 | ZBTB11 | 10156 | -0.340 | 0.0599 | No | ||
| 133 | GAB1 | 10210 | -0.353 | 0.0582 | No | ||
| 134 | NDST1 | 10276 | -0.370 | 0.0560 | No | ||
| 135 | LRRTM3 | 10283 | -0.371 | 0.0569 | No | ||
| 136 | SEMA4C | 10410 | -0.396 | 0.0515 | No | ||
| 137 | EYA1 | 10412 | -0.397 | 0.0528 | No | ||
| 138 | EFNA3 | 10508 | -0.413 | 0.0491 | No | ||
| 139 | SH2D3C | 10512 | -0.414 | 0.0504 | No | ||
| 140 | NRXN3 | 10596 | -0.433 | 0.0473 | No | ||
| 141 | DCUN1D1 | 10651 | -0.443 | 0.0460 | No | ||
| 142 | ADAM10 | 10666 | -0.446 | 0.0467 | No | ||
| 143 | PIAS1 | 10846 | -0.483 | 0.0387 | No | ||
| 144 | APLP2 | 10912 | -0.497 | 0.0369 | No | ||
| 145 | SPIRE2 | 10961 | -0.508 | 0.0361 | No | ||
| 146 | GTF2IRD1 | 11010 | -0.521 | 0.0353 | No | ||
| 147 | TRAM2 | 11135 | -0.541 | 0.0304 | No | ||
| 148 | GPC4 | 11252 | -0.565 | 0.0261 | No | ||
| 149 | FMNL2 | 11279 | -0.570 | 0.0266 | No | ||
| 150 | ZIC2 | 11309 | -0.576 | 0.0271 | No | ||
| 151 | CTDSPL | 11332 | -0.581 | 0.0279 | No | ||
| 152 | SS18 | 11411 | -0.596 | 0.0257 | No | ||
| 153 | AFF3 | 11453 | -0.606 | 0.0256 | No | ||
| 154 | KPNA4 | 11754 | -0.661 | 0.0116 | No | ||
| 155 | JUN | 12005 | -0.707 | 0.0005 | No | ||
| 156 | HS3ST3B1 | 12087 | -0.721 | -0.0014 | No | ||
| 157 | GNB1 | 12227 | -0.753 | -0.0064 | No | ||
| 158 | PPP3CB | 12239 | -0.757 | -0.0043 | No | ||
| 159 | GABRG2 | 12334 | -0.779 | -0.0067 | No | ||
| 160 | IL1A | 12481 | -0.806 | -0.0119 | No | ||
| 161 | OVOL1 | 12583 | -0.834 | -0.0145 | No | ||
| 162 | ENAH | 12584 | -0.834 | -0.0115 | No | ||
| 163 | MLLT10 | 12640 | -0.846 | -0.0116 | No | ||
| 164 | CDH11 | 12655 | -0.848 | -0.0094 | No | ||
| 165 | SP3 | 12743 | -0.869 | -0.0111 | No | ||
| 166 | CSNK1A1 | 12887 | -0.897 | -0.0158 | No | ||
| 167 | RALGPS1 | 12921 | -0.903 | -0.0144 | No | ||
| 168 | PPP2R5C | 12959 | -0.911 | -0.0133 | No | ||
| 169 | MAST1 | 12973 | -0.916 | -0.0108 | No | ||
| 170 | FIGN | 12980 | -0.918 | -0.0079 | No | ||
| 171 | FHOD3 | 12990 | -0.920 | -0.0052 | No | ||
| 172 | PCGF2 | 13314 | -0.990 | -0.0193 | No | ||
| 173 | SRPK2 | 13395 | -1.008 | -0.0201 | No | ||
| 174 | GRM5 | 13426 | -1.015 | -0.0182 | No | ||
| 175 | XPO7 | 13436 | -1.016 | -0.0152 | No | ||
| 176 | CDC73 | 13488 | -1.028 | -0.0144 | No | ||
| 177 | WDFY3 | 13778 | -1.093 | -0.0263 | No | ||
| 178 | NUDT4 | 13949 | -1.137 | -0.0316 | No | ||
| 179 | LRIG1 | 13993 | -1.146 | -0.0299 | No | ||
| 180 | CLPX | 14112 | -1.172 | -0.0323 | No | ||
| 181 | MADD | 14631 | -1.298 | -0.0559 | No | ||
| 182 | IRX3 | 14731 | -1.321 | -0.0567 | No | ||
| 183 | TBC1D4 | 14859 | -1.351 | -0.0589 | No | ||
| 184 | ETF1 | 14934 | -1.370 | -0.0582 | No | ||
| 185 | RHOT1 | 15040 | -1.397 | -0.0590 | No | ||
| 186 | SPAG8 | 15052 | -1.400 | -0.0547 | No | ||
| 187 | RPS6KA5 | 15107 | -1.413 | -0.0527 | No | ||
| 188 | JPH1 | 15124 | -1.418 | -0.0487 | No | ||
| 189 | UTX | 15186 | -1.435 | -0.0470 | No | ||
| 190 | ARHGAP5 | 15242 | -1.448 | -0.0449 | No | ||
| 191 | PPP2R5D | 15363 | -1.488 | -0.0463 | No | ||
| 192 | SSH2 | 15380 | -1.495 | -0.0420 | No | ||
| 193 | FGF13 | 15436 | -1.513 | -0.0397 | No | ||
| 194 | CAPN3 | 15568 | -1.553 | -0.0414 | No | ||
| 195 | CITED2 | 15609 | -1.567 | -0.0381 | No | ||
| 196 | DUSP16 | 15974 | -1.688 | -0.0520 | No | ||
| 197 | ANKRD50 | 15995 | -1.694 | -0.0472 | No | ||
| 198 | CDH2 | 16010 | -1.700 | -0.0421 | No | ||
| 199 | PBX3 | 16043 | -1.713 | -0.0378 | No | ||
| 200 | NLGN3 | 16062 | -1.720 | -0.0328 | No | ||
| 201 | SRPX | 16124 | -1.740 | -0.0301 | No | ||
| 202 | TMEM16D | 16129 | -1.741 | -0.0243 | No | ||
| 203 | PUM2 | 16163 | -1.755 | -0.0199 | No | ||
| 204 | CRSP6 | 16232 | -1.774 | -0.0175 | No | ||
| 205 | TTN | 16405 | -1.833 | -0.0204 | No | ||
| 206 | MARK1 | 16453 | -1.851 | -0.0165 | No | ||
| 207 | MAPK1 | 16496 | -1.873 | -0.0123 | No | ||
| 208 | CDH13 | 16562 | -1.899 | -0.0092 | No | ||
| 209 | KBTBD2 | 16880 | -2.050 | -0.0193 | No | ||
| 210 | UBE2G1 | 17068 | -2.144 | -0.0220 | No | ||
| 211 | SNAI1 | 17371 | -2.345 | -0.0303 | No | ||
| 212 | ZFX | 17376 | -2.347 | -0.0223 | No | ||
| 213 | HIVEP2 | 17465 | -2.405 | -0.0188 | No | ||
| 214 | VGLL4 | 17627 | -2.532 | -0.0187 | No | ||
| 215 | UNC5A | 17692 | -2.587 | -0.0132 | No | ||
| 216 | LRP12 | 17718 | -2.615 | -0.0054 | No | ||
| 217 | TRPS1 | 17732 | -2.624 | 0.0030 | No | ||
| 218 | AR | 18189 | -3.160 | -0.0108 | No | ||
| 219 | SFRS3 | 18223 | -3.218 | -0.0014 | No | ||
| 220 | SLC25A26 | 18232 | -3.228 | 0.0094 | No | ||
| 221 | AP3S1 | 18254 | -3.277 | 0.0197 | No |
