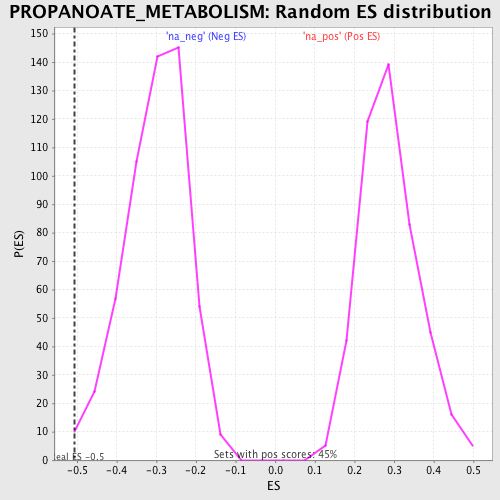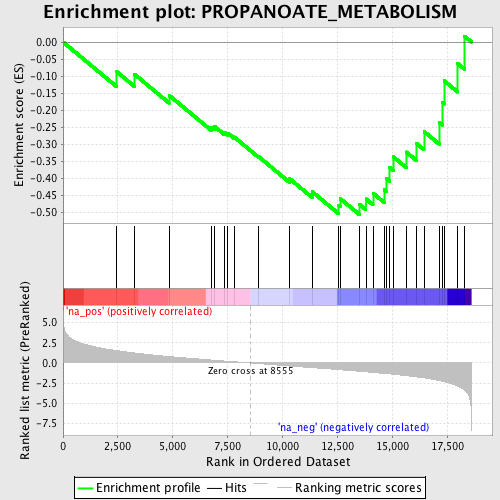
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | set04_transDMpreB_versus_DMpreB |
| Phenotype | NoPhenotypeAvailable |
| Upregulated in class | na_neg |
| GeneSet | PROPANOATE_METABOLISM |
| Enrichment Score (ES) | -0.5072165 |
| Normalized Enrichment Score (NES) | -1.6813139 |
| Nominal p-value | 0.0054945056 |
| FDR q-value | 0.09017781 |
| FWER p-Value | 0.929 |

| PROBE | GENE SYMBOL | GENE_TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|---|
| 1 | ALDH2 | 2451 | 1.521 | -0.0876 | No | ||
| 2 | HADHA | 3240 | 1.241 | -0.0940 | No | ||
| 3 | SUCLA2 | 4830 | 0.789 | -0.1565 | No | ||
| 4 | ALDH1A2 | 6767 | 0.355 | -0.2503 | No | ||
| 5 | ALDH3A1 | 6906 | 0.323 | -0.2484 | No | ||
| 6 | PCCB | 7338 | 0.235 | -0.2647 | No | ||
| 7 | ALDH1A1 | 7505 | 0.199 | -0.2679 | No | ||
| 8 | MUT | 7797 | 0.148 | -0.2792 | No | ||
| 9 | ALDH9A1 | 8910 | -0.072 | -0.3369 | No | ||
| 10 | ALDH1A3 | 10315 | -0.378 | -0.4015 | No | ||
| 11 | ACADM | 11357 | -0.585 | -0.4405 | No | ||
| 12 | ALDH6A1 | 12545 | -0.823 | -0.4804 | Yes | ||
| 13 | LDHC | 12625 | -0.842 | -0.4602 | Yes | ||
| 14 | ACAT1 | 13500 | -1.030 | -0.4773 | Yes | ||
| 15 | EHHADH | 13804 | -1.102 | -0.4615 | Yes | ||
| 16 | ACADL | 14125 | -1.177 | -0.4446 | Yes | ||
| 17 | LDHA | 14647 | -1.302 | -0.4348 | Yes | ||
| 18 | MLYCD | 14745 | -1.323 | -0.4015 | Yes | ||
| 19 | ACAT2 | 14887 | -1.358 | -0.3696 | Yes | ||
| 20 | MCEE | 15049 | -1.400 | -0.3376 | Yes | ||
| 21 | SUCLG2 | 15636 | -1.575 | -0.3234 | Yes | ||
| 22 | ACADSB | 16090 | -1.731 | -0.2974 | Yes | ||
| 23 | PCCA | 16449 | -1.849 | -0.2629 | Yes | ||
| 24 | SUCLG1 | 17156 | -2.196 | -0.2371 | Yes | ||
| 25 | ALDH1B1 | 17291 | -2.282 | -0.1780 | Yes | ||
| 26 | SDS | 17373 | -2.346 | -0.1141 | Yes | ||
| 27 | ECHS1 | 17985 | -2.883 | -0.0632 | Yes | ||
| 28 | ACACA | 18291 | -3.340 | 0.0175 | Yes |