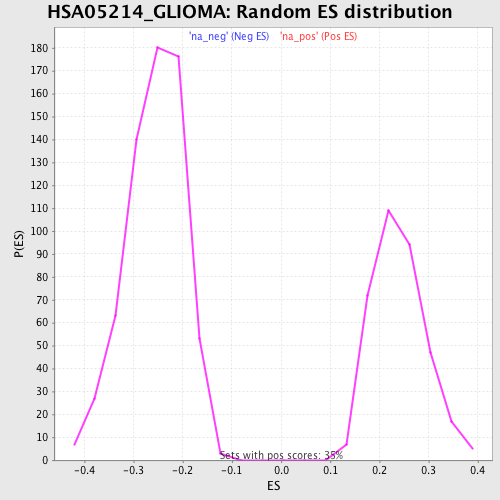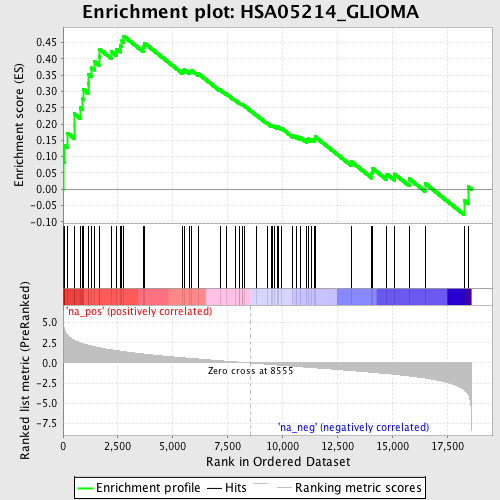
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | set04_transDMpreB_versus_DMpreB |
| Phenotype | NoPhenotypeAvailable |
| Upregulated in class | na_pos |
| GeneSet | HSA05214_GLIOMA |
| Enrichment Score (ES) | 0.46967265 |
| Normalized Enrichment Score (NES) | 1.9597737 |
| Nominal p-value | 0.0 |
| FDR q-value | 0.038638603 |
| FWER p-Value | 0.168 |

| PROBE | GENE SYMBOL | GENE_TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|---|
| 1 | CCND1 | 2 | 6.319 | 0.0840 | Yes | ||
| 2 | PRKCB1 | 67 | 4.060 | 0.1346 | Yes | ||
| 3 | CAMK2A | 208 | 3.401 | 0.1723 | Yes | ||
| 4 | TP53 | 501 | 2.825 | 0.1942 | Yes | ||
| 5 | MAP2K1 | 502 | 2.825 | 0.2318 | Yes | ||
| 6 | PLCG2 | 775 | 2.497 | 0.2504 | Yes | ||
| 7 | CAMK2G | 879 | 2.414 | 0.2770 | Yes | ||
| 8 | GRB2 | 924 | 2.371 | 0.3062 | Yes | ||
| 9 | PIK3CG | 1139 | 2.200 | 0.3239 | Yes | ||
| 10 | E2F1 | 1158 | 2.188 | 0.3521 | Yes | ||
| 11 | MAPK3 | 1303 | 2.079 | 0.3720 | Yes | ||
| 12 | E2F2 | 1423 | 2.010 | 0.3923 | Yes | ||
| 13 | MAP2K2 | 1638 | 1.892 | 0.4060 | Yes | ||
| 14 | PDGFRB | 1659 | 1.880 | 0.4299 | Yes | ||
| 15 | EGF | 2206 | 1.627 | 0.4222 | Yes | ||
| 16 | MDM2 | 2440 | 1.525 | 0.4299 | Yes | ||
| 17 | SOS2 | 2609 | 1.463 | 0.4403 | Yes | ||
| 18 | E2F3 | 2680 | 1.433 | 0.4556 | Yes | ||
| 19 | RAF1 | 2767 | 1.403 | 0.4697 | Yes | ||
| 20 | ARAF | 3649 | 1.111 | 0.4370 | No | ||
| 21 | PLCG1 | 3717 | 1.086 | 0.4478 | No | ||
| 22 | PDGFA | 5453 | 0.640 | 0.3628 | No | ||
| 23 | SHC3 | 5546 | 0.622 | 0.3662 | No | ||
| 24 | SOS1 | 5763 | 0.579 | 0.3622 | No | ||
| 25 | PIK3CD | 5865 | 0.555 | 0.3642 | No | ||
| 26 | PIK3R5 | 6156 | 0.490 | 0.3551 | No | ||
| 27 | CDK6 | 7151 | 0.272 | 0.3051 | No | ||
| 28 | PIK3R1 | 7425 | 0.214 | 0.2933 | No | ||
| 29 | PIK3R3 | 7845 | 0.138 | 0.2725 | No | ||
| 30 | CALML3 | 8046 | 0.101 | 0.2631 | No | ||
| 31 | PTEN | 8053 | 0.100 | 0.2641 | No | ||
| 32 | CALM2 | 8186 | 0.076 | 0.2580 | No | ||
| 33 | RB1 | 8189 | 0.075 | 0.2589 | No | ||
| 34 | CALM3 | 8250 | 0.066 | 0.2566 | No | ||
| 35 | CAMK2B | 8806 | -0.050 | 0.2273 | No | ||
| 36 | AKT1 | 9294 | -0.158 | 0.2032 | No | ||
| 37 | FRAP1 | 9494 | -0.199 | 0.1951 | No | ||
| 38 | PIK3R2 | 9527 | -0.203 | 0.1961 | No | ||
| 39 | NRAS | 9654 | -0.233 | 0.1924 | No | ||
| 40 | CALM1 | 9748 | -0.254 | 0.1907 | No | ||
| 41 | IGF1 | 9804 | -0.266 | 0.1913 | No | ||
| 42 | IGF1R | 9965 | -0.299 | 0.1867 | No | ||
| 43 | PRKCA | 10469 | -0.407 | 0.1650 | No | ||
| 44 | KRAS | 10637 | -0.440 | 0.1619 | No | ||
| 45 | TGFA | 10824 | -0.480 | 0.1582 | No | ||
| 46 | CAMK2D | 11082 | -0.534 | 0.1515 | No | ||
| 47 | PDGFRA | 11164 | -0.546 | 0.1544 | No | ||
| 48 | EGFR | 11316 | -0.578 | 0.1539 | No | ||
| 49 | CDKN1A | 11472 | -0.609 | 0.1537 | No | ||
| 50 | CDKN2A | 11488 | -0.612 | 0.1610 | No | ||
| 51 | HRAS | 13137 | -0.949 | 0.0849 | No | ||
| 52 | PIK3CB | 14076 | -1.164 | 0.0498 | No | ||
| 53 | PIK3CA | 14098 | -1.169 | 0.0642 | No | ||
| 54 | AKT2 | 14759 | -1.328 | 0.0463 | No | ||
| 55 | SHC1 | 15101 | -1.412 | 0.0468 | No | ||
| 56 | PDGFB | 15771 | -1.618 | 0.0322 | No | ||
| 57 | MAPK1 | 16496 | -1.873 | 0.0182 | No | ||
| 58 | AKT3 | 18272 | -3.301 | -0.0335 | No | ||
| 59 | CDK4 | 18466 | -3.907 | 0.0081 | No |