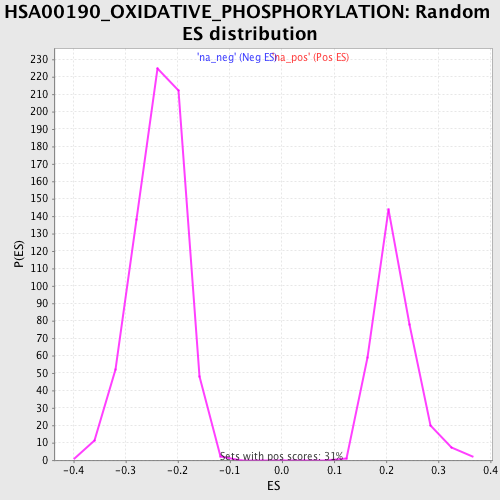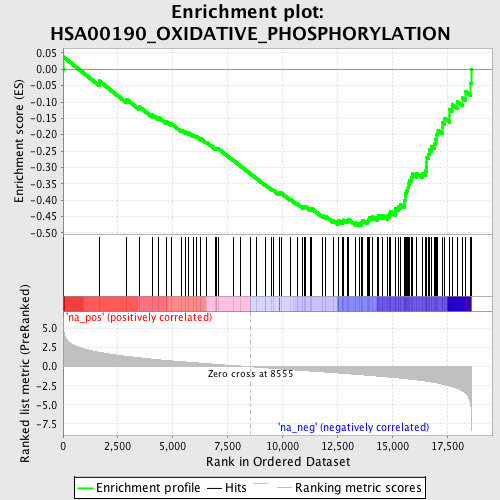
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | set04_transDMpreB_versus_DMpreB |
| Phenotype | NoPhenotypeAvailable |
| Upregulated in class | na_neg |
| GeneSet | HSA00190_OXIDATIVE_PHOSPHORYLATION |
| Enrichment Score (ES) | -0.47906923 |
| Normalized Enrichment Score (NES) | -2.0259712 |
| Nominal p-value | 0.0 |
| FDR q-value | 0.006737794 |
| FWER p-Value | 0.042 |

| PROBE | GENE SYMBOL | GENE_TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|---|
| 1 | COX6A2 | 15 | 5.054 | 0.0381 | No | ||
| 2 | TCIRG1 | 1644 | 1.887 | -0.0353 | No | ||
| 3 | NDUFB9 | 2873 | 1.363 | -0.0911 | No | ||
| 4 | COX10 | 3462 | 1.172 | -0.1138 | No | ||
| 5 | UQCRFS1 | 4079 | 0.982 | -0.1395 | No | ||
| 6 | ATP6V1D | 4335 | 0.917 | -0.1463 | No | ||
| 7 | COX8A | 4698 | 0.822 | -0.1595 | No | ||
| 8 | NDUFS1 | 4920 | 0.766 | -0.1655 | No | ||
| 9 | ATP5B | 5397 | 0.651 | -0.1862 | No | ||
| 10 | ATP6AP1 | 5584 | 0.614 | -0.1915 | No | ||
| 11 | NDUFA10 | 5711 | 0.590 | -0.1938 | No | ||
| 12 | COX4I2 | 5923 | 0.543 | -0.2010 | No | ||
| 13 | ATP5A1 | 6077 | 0.508 | -0.2053 | No | ||
| 14 | ATP6V1C2 | 6281 | 0.464 | -0.2127 | No | ||
| 15 | ATP6V1B1 | 6534 | 0.406 | -0.2232 | No | ||
| 16 | NDUFB10 | 6933 | 0.318 | -0.2423 | No | ||
| 17 | COX6B2 | 6971 | 0.309 | -0.2419 | No | ||
| 18 | ATP5D | 6997 | 0.304 | -0.2409 | No | ||
| 19 | ATP6V1G3 | 7072 | 0.287 | -0.2427 | No | ||
| 20 | SDHA | 7744 | 0.158 | -0.2777 | No | ||
| 21 | UQCRB | 8089 | 0.092 | -0.2956 | No | ||
| 22 | UQCRC1 | 8522 | 0.006 | -0.3189 | No | ||
| 23 | ATP4B | 8544 | 0.002 | -0.3200 | No | ||
| 24 | SDHD | 8824 | -0.054 | -0.3346 | No | ||
| 25 | NDUFA6 | 9214 | -0.139 | -0.3546 | No | ||
| 26 | NDUFS7 | 9216 | -0.140 | -0.3536 | No | ||
| 27 | COX8C | 9507 | -0.201 | -0.3677 | No | ||
| 28 | NDUFC2 | 9567 | -0.214 | -0.3692 | No | ||
| 29 | NDUFA13 | 9842 | -0.272 | -0.3819 | No | ||
| 30 | ATP5C1 | 9856 | -0.275 | -0.3805 | No | ||
| 31 | ATP6V0C | 9859 | -0.276 | -0.3785 | No | ||
| 32 | NDUFA3 | 9878 | -0.280 | -0.3773 | No | ||
| 33 | NDUFB5 | 9934 | -0.292 | -0.3780 | No | ||
| 34 | ATP6V1B2 | 10348 | -0.385 | -0.3974 | No | ||
| 35 | ATP6V0A4 | 10675 | -0.448 | -0.4115 | No | ||
| 36 | NDUFA5 | 10917 | -0.499 | -0.4207 | No | ||
| 37 | COX7B | 10983 | -0.515 | -0.4202 | No | ||
| 38 | ATP6V0D2 | 11051 | -0.528 | -0.4198 | No | ||
| 39 | ATP6V1H | 11291 | -0.573 | -0.4283 | No | ||
| 40 | ATP5I | 11310 | -0.576 | -0.4248 | No | ||
| 41 | ATP5O | 11826 | -0.676 | -0.4474 | No | ||
| 42 | ATP5G3 | 11977 | -0.704 | -0.4501 | No | ||
| 43 | ATP5G2 | 12328 | -0.777 | -0.4630 | No | ||
| 44 | ATP6V1G1 | 12529 | -0.818 | -0.4675 | No | ||
| 45 | NDUFAB1 | 12549 | -0.824 | -0.4622 | No | ||
| 46 | ATP6V0A2 | 12748 | -0.870 | -0.4662 | No | ||
| 47 | ATP4A | 12773 | -0.873 | -0.4608 | No | ||
| 48 | ATP12A | 12943 | -0.909 | -0.4629 | No | ||
| 49 | ATP5E | 12996 | -0.921 | -0.4586 | No | ||
| 50 | CYC1 | 13347 | -0.997 | -0.4698 | No | ||
| 51 | ATP6V1F | 13519 | -1.034 | -0.4711 | Yes | ||
| 52 | COX5A | 13617 | -1.060 | -0.4682 | Yes | ||
| 53 | NDUFB7 | 13636 | -1.062 | -0.4610 | Yes | ||
| 54 | ATP6V0A1 | 13859 | -1.115 | -0.4644 | Yes | ||
| 55 | ATP6V1E1 | 13912 | -1.128 | -0.4585 | Yes | ||
| 56 | COX7A2 | 13976 | -1.142 | -0.4531 | Yes | ||
| 57 | NDUFB2 | 14095 | -1.168 | -0.4505 | Yes | ||
| 58 | NDUFS8 | 14334 | -1.229 | -0.4539 | Yes | ||
| 59 | UQCR | 14369 | -1.237 | -0.4462 | Yes | ||
| 60 | ATP5G1 | 14549 | -1.282 | -0.4460 | Yes | ||
| 61 | COX7C | 14783 | -1.333 | -0.4483 | Yes | ||
| 62 | ATP5F1 | 14860 | -1.351 | -0.4420 | Yes | ||
| 63 | UQCRH | 14927 | -1.368 | -0.4350 | Yes | ||
| 64 | NDUFB11 | 15137 | -1.422 | -0.4354 | Yes | ||
| 65 | ATP6V0D1 | 15167 | -1.430 | -0.4259 | Yes | ||
| 66 | COX5B | 15284 | -1.461 | -0.4209 | Yes | ||
| 67 | COX6A1 | 15368 | -1.490 | -0.4139 | Yes | ||
| 68 | NDUFA1 | 15551 | -1.547 | -0.4119 | Yes | ||
| 69 | NDUFC1 | 15556 | -1.548 | -0.4001 | Yes | ||
| 70 | ATP6V1E2 | 15588 | -1.559 | -0.3898 | Yes | ||
| 71 | COX15 | 15601 | -1.563 | -0.3784 | Yes | ||
| 72 | COX6C | 15654 | -1.581 | -0.3691 | Yes | ||
| 73 | ATP6V0B | 15718 | -1.601 | -0.3601 | Yes | ||
| 74 | NDUFS5 | 15751 | -1.610 | -0.3495 | Yes | ||
| 75 | NDUFB8 | 15795 | -1.624 | -0.3393 | Yes | ||
| 76 | ATP6V1G2 | 15890 | -1.658 | -0.3316 | Yes | ||
| 77 | COX6B1 | 15905 | -1.664 | -0.3195 | Yes | ||
| 78 | NDUFB4 | 16118 | -1.738 | -0.3176 | Yes | ||
| 79 | ATP5J2 | 16386 | -1.826 | -0.3180 | Yes | ||
| 80 | NDUFV1 | 16516 | -1.880 | -0.3104 | Yes | ||
| 81 | ATP6V1A | 16556 | -1.896 | -0.2979 | Yes | ||
| 82 | NDUFA11 | 16577 | -1.905 | -0.2844 | Yes | ||
| 83 | ATP6V0E1 | 16583 | -1.910 | -0.2699 | Yes | ||
| 84 | ATP5H | 16663 | -1.947 | -0.2592 | Yes | ||
| 85 | SDHC | 16695 | -1.962 | -0.2458 | Yes | ||
| 86 | NDUFA7 | 16802 | -2.007 | -0.2360 | Yes | ||
| 87 | NDUFS3 | 16932 | -2.073 | -0.2270 | Yes | ||
| 88 | COX7A1 | 16987 | -2.101 | -0.2138 | Yes | ||
| 89 | NDUFA4 | 17027 | -2.120 | -0.1995 | Yes | ||
| 90 | NDUFS2 | 17085 | -2.153 | -0.1860 | Yes | ||
| 91 | NDUFA2 | 17274 | -2.274 | -0.1787 | Yes | ||
| 92 | COX17 | 17278 | -2.276 | -0.1613 | Yes | ||
| 93 | NDUFA8 | 17394 | -2.364 | -0.1493 | Yes | ||
| 94 | ATP5J | 17623 | -2.531 | -0.1421 | Yes | ||
| 95 | NDUFS4 | 17625 | -2.531 | -0.1227 | Yes | ||
| 96 | NDUFB3 | 17725 | -2.621 | -0.1079 | Yes | ||
| 97 | NDUFA9 | 17956 | -2.841 | -0.0984 | Yes | ||
| 98 | COX4I1 | 18211 | -3.196 | -0.0875 | Yes | ||
| 99 | UQCRC2 | 18342 | -3.482 | -0.0677 | Yes | ||
| 100 | ATP6V1C1 | 18570 | -4.835 | -0.0427 | Yes | ||
| 101 | SDHB | 18604 | -5.865 | 0.0006 | Yes |