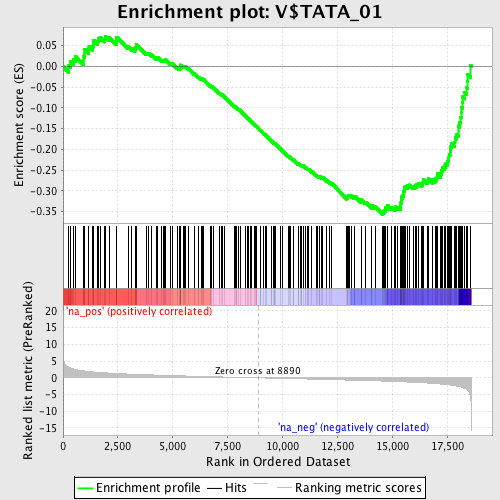
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | set04_DMproB_versus_LMproB |
| Phenotype | NoPhenotypeAvailable |
| Upregulated in class | na_neg |
| GeneSet | V$TATA_01 |
| Enrichment Score (ES) | -0.35697395 |
| Normalized Enrichment Score (NES) | -1.6069802 |
| Nominal p-value | 0.0 |
| FDR q-value | 0.65238315 |
| FWER p-Value | 0.857 |

| PROBE | GENE SYMBOL | GENE_TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|---|
| 1 | S100A4 | 251 | 3.153 | 0.0017 | No | ||
| 2 | GADD45G | 319 | 2.953 | 0.0124 | No | ||
| 3 | GCH1 | 484 | 2.616 | 0.0162 | No | ||
| 4 | PLAG1 | 560 | 2.500 | 0.0243 | No | ||
| 5 | HOXA11 | 940 | 2.059 | 0.0137 | No | ||
| 6 | WSB2 | 943 | 2.057 | 0.0236 | No | ||
| 7 | ITPR1 | 976 | 2.031 | 0.0317 | No | ||
| 8 | AQP1 | 984 | 2.022 | 0.0411 | No | ||
| 9 | EGFL7 | 1158 | 1.894 | 0.0409 | No | ||
| 10 | GSC | 1177 | 1.881 | 0.0491 | No | ||
| 11 | MAT2A | 1356 | 1.763 | 0.0480 | No | ||
| 12 | NOG | 1379 | 1.749 | 0.0553 | No | ||
| 13 | HOXD4 | 1388 | 1.744 | 0.0633 | No | ||
| 14 | GRIN2B | 1577 | 1.639 | 0.0611 | No | ||
| 15 | ANKMY2 | 1602 | 1.627 | 0.0677 | No | ||
| 16 | SOX5 | 1703 | 1.576 | 0.0699 | No | ||
| 17 | LHX6 | 1907 | 1.486 | 0.0661 | No | ||
| 18 | BRUNOL4 | 1912 | 1.485 | 0.0731 | No | ||
| 19 | AMPD1 | 2092 | 1.414 | 0.0703 | No | ||
| 20 | NNT | 2421 | 1.301 | 0.0588 | No | ||
| 21 | SFRS3 | 2439 | 1.298 | 0.0642 | No | ||
| 22 | RASGRP3 | 2443 | 1.296 | 0.0703 | No | ||
| 23 | TTC17 | 2961 | 1.136 | 0.0478 | No | ||
| 24 | EEF1A1 | 3129 | 1.090 | 0.0440 | No | ||
| 25 | HOXA10 | 3278 | 1.050 | 0.0410 | No | ||
| 26 | KBTBD5 | 3300 | 1.045 | 0.0450 | No | ||
| 27 | UCN | 3327 | 1.038 | 0.0486 | No | ||
| 28 | NANOS1 | 3352 | 1.031 | 0.0523 | No | ||
| 29 | RUNX2 | 3803 | 0.917 | 0.0323 | No | ||
| 30 | TCF7 | 3887 | 0.897 | 0.0322 | No | ||
| 31 | MID1 | 4032 | 0.866 | 0.0286 | No | ||
| 32 | DUOX2 | 4247 | 0.813 | 0.0209 | No | ||
| 33 | NR0B2 | 4309 | 0.795 | 0.0215 | No | ||
| 34 | SUPT4H1 | 4491 | 0.758 | 0.0153 | No | ||
| 35 | BHLHB5 | 4559 | 0.747 | 0.0153 | No | ||
| 36 | TMEPAI | 4641 | 0.728 | 0.0145 | No | ||
| 37 | FOXP3 | 4662 | 0.724 | 0.0169 | No | ||
| 38 | BACH1 | 4885 | 0.681 | 0.0081 | No | ||
| 39 | IL11 | 4972 | 0.664 | 0.0067 | No | ||
| 40 | NEFH | 5190 | 0.622 | -0.0021 | No | ||
| 41 | SCAMP3 | 5302 | 0.601 | -0.0052 | No | ||
| 42 | LIF | 5329 | 0.596 | -0.0037 | No | ||
| 43 | MITF | 5336 | 0.594 | -0.0011 | No | ||
| 44 | TRDN | 5343 | 0.593 | 0.0014 | No | ||
| 45 | KRTAP6-2 | 5359 | 0.590 | 0.0035 | No | ||
| 46 | MYF6 | 5492 | 0.566 | -0.0009 | No | ||
| 47 | HIST1H2AA | 5543 | 0.556 | -0.0009 | No | ||
| 48 | MMP10 | 5592 | 0.547 | -0.0009 | No | ||
| 49 | SPRR1B | 5703 | 0.527 | -0.0043 | No | ||
| 50 | LTA | 5983 | 0.476 | -0.0171 | No | ||
| 51 | GDF8 | 6180 | 0.439 | -0.0256 | No | ||
| 52 | SOSTDC1 | 6286 | 0.421 | -0.0293 | No | ||
| 53 | CKB | 6339 | 0.410 | -0.0301 | No | ||
| 54 | NPTX1 | 6419 | 0.397 | -0.0325 | No | ||
| 55 | COL1A1 | 6717 | 0.353 | -0.0469 | No | ||
| 56 | STC1 | 6780 | 0.343 | -0.0486 | No | ||
| 57 | MT4 | 6864 | 0.328 | -0.0515 | No | ||
| 58 | TGM3 | 7139 | 0.277 | -0.0650 | No | ||
| 59 | MAP2K5 | 7206 | 0.265 | -0.0673 | No | ||
| 60 | DDX3X | 7250 | 0.258 | -0.0684 | No | ||
| 61 | H3F3B | 7356 | 0.240 | -0.0729 | No | ||
| 62 | KLF2 | 7801 | 0.172 | -0.0962 | No | ||
| 63 | NPPA | 7834 | 0.168 | -0.0971 | No | ||
| 64 | PLUNC | 7891 | 0.159 | -0.0994 | No | ||
| 65 | MYH3 | 7922 | 0.153 | -0.1003 | No | ||
| 66 | SLC34A1 | 7998 | 0.138 | -0.1037 | No | ||
| 67 | HOXC5 | 8083 | 0.125 | -0.1076 | No | ||
| 68 | TTR | 8331 | 0.090 | -0.1206 | No | ||
| 69 | FSHB | 8406 | 0.079 | -0.1242 | No | ||
| 70 | CXCR4 | 8439 | 0.072 | -0.1256 | No | ||
| 71 | HIST1H1B | 8557 | 0.054 | -0.1317 | No | ||
| 72 | NEUROG2 | 8570 | 0.052 | -0.1321 | No | ||
| 73 | STAC | 8733 | 0.028 | -0.1407 | No | ||
| 74 | MYOZ2 | 8759 | 0.023 | -0.1420 | No | ||
| 75 | CRABP2 | 8798 | 0.017 | -0.1440 | No | ||
| 76 | DES | 8834 | 0.011 | -0.1458 | No | ||
| 77 | SLC2A4 | 8997 | -0.015 | -0.1545 | No | ||
| 78 | SMYD1 | 9014 | -0.018 | -0.1553 | No | ||
| 79 | ECE2 | 9126 | -0.035 | -0.1612 | No | ||
| 80 | JARID2 | 9203 | -0.046 | -0.1651 | No | ||
| 81 | PDLIM7 | 9259 | -0.055 | -0.1678 | No | ||
| 82 | CNN1 | 9490 | -0.088 | -0.1798 | No | ||
| 83 | H2AFZ | 9497 | -0.089 | -0.1797 | No | ||
| 84 | TNFRSF17 | 9575 | -0.101 | -0.1834 | No | ||
| 85 | CDX1 | 9616 | -0.106 | -0.1851 | No | ||
| 86 | MIP | 9639 | -0.110 | -0.1857 | No | ||
| 87 | MYCT1 | 9662 | -0.113 | -0.1864 | No | ||
| 88 | COL10A1 | 9895 | -0.150 | -0.1983 | No | ||
| 89 | TNNC2 | 9986 | -0.164 | -0.2023 | No | ||
| 90 | ADAMTS15 | 10269 | -0.206 | -0.2166 | No | ||
| 91 | CSH1 | 10304 | -0.210 | -0.2175 | No | ||
| 92 | HNRPA0 | 10381 | -0.221 | -0.2205 | No | ||
| 93 | EDN1 | 10501 | -0.241 | -0.2258 | No | ||
| 94 | RBP2 | 10514 | -0.244 | -0.2253 | No | ||
| 95 | CYP1A2 | 10719 | -0.280 | -0.2350 | No | ||
| 96 | NR4A1 | 10721 | -0.280 | -0.2337 | No | ||
| 97 | GPRC5D | 10823 | -0.295 | -0.2377 | No | ||
| 98 | CLCN1 | 10869 | -0.303 | -0.2387 | No | ||
| 99 | ATOH1 | 10881 | -0.305 | -0.2378 | No | ||
| 100 | EGR2 | 10935 | -0.314 | -0.2392 | No | ||
| 101 | NDRG2 | 11062 | -0.337 | -0.2444 | No | ||
| 102 | OLIG2 | 11159 | -0.352 | -0.2479 | No | ||
| 103 | PTP4A1 | 11189 | -0.357 | -0.2477 | No | ||
| 104 | PCDH8 | 11333 | -0.382 | -0.2536 | No | ||
| 105 | POU3F4 | 11527 | -0.411 | -0.2621 | No | ||
| 106 | HIST1H1A | 11607 | -0.424 | -0.2643 | No | ||
| 107 | HIST1H2BA | 11689 | -0.438 | -0.2666 | No | ||
| 108 | RGS4 | 11699 | -0.439 | -0.2649 | No | ||
| 109 | MYH4 | 11785 | -0.453 | -0.2674 | No | ||
| 110 | MMP20 | 11835 | -0.460 | -0.2678 | No | ||
| 111 | ASB4 | 12001 | -0.486 | -0.2744 | No | ||
| 112 | BNIP3 | 12140 | -0.511 | -0.2794 | No | ||
| 113 | GRID2 | 12239 | -0.529 | -0.2821 | No | ||
| 114 | SCN8A | 12936 | -0.655 | -0.3167 | No | ||
| 115 | ADAM11 | 12970 | -0.662 | -0.3153 | No | ||
| 116 | DKK2 | 12978 | -0.664 | -0.3125 | No | ||
| 117 | CSH2 | 13019 | -0.671 | -0.3114 | No | ||
| 118 | SEPP1 | 13063 | -0.679 | -0.3104 | No | ||
| 119 | IL22 | 13156 | -0.692 | -0.3121 | No | ||
| 120 | HIST1H1C | 13273 | -0.708 | -0.3149 | No | ||
| 121 | ELAVL3 | 13298 | -0.712 | -0.3128 | No | ||
| 122 | PANK1 | 13581 | -0.766 | -0.3243 | No | ||
| 123 | RPS3 | 13597 | -0.768 | -0.3214 | No | ||
| 124 | FGF4 | 13782 | -0.801 | -0.3275 | No | ||
| 125 | RHOBTB1 | 14057 | -0.854 | -0.3382 | No | ||
| 126 | COL13A1 | 14070 | -0.856 | -0.3347 | No | ||
| 127 | CX3CL1 | 14219 | -0.886 | -0.3385 | No | ||
| 128 | PDYN | 14561 | -0.960 | -0.3523 | Yes | ||
| 129 | HSPB2 | 14617 | -0.971 | -0.3506 | Yes | ||
| 130 | GTF2A1 | 14644 | -0.978 | -0.3472 | Yes | ||
| 131 | SLC25A4 | 14691 | -0.987 | -0.3450 | Yes | ||
| 132 | RHOA | 14709 | -0.991 | -0.3411 | Yes | ||
| 133 | OLIG3 | 14769 | -1.006 | -0.3394 | Yes | ||
| 134 | ABCA1 | 14786 | -1.012 | -0.3353 | Yes | ||
| 135 | CKM | 14977 | -1.053 | -0.3405 | Yes | ||
| 136 | ABLIM1 | 15096 | -1.081 | -0.3417 | Yes | ||
| 137 | PPARG | 15132 | -1.090 | -0.3383 | Yes | ||
| 138 | FGF20 | 15247 | -1.124 | -0.3390 | Yes | ||
| 139 | CILP | 15370 | -1.154 | -0.3400 | Yes | ||
| 140 | SIM1 | 15374 | -1.154 | -0.3346 | Yes | ||
| 141 | GBE1 | 15378 | -1.157 | -0.3291 | Yes | ||
| 142 | CRYBA1 | 15408 | -1.164 | -0.3251 | Yes | ||
| 143 | KIRREL2 | 15424 | -1.167 | -0.3202 | Yes | ||
| 144 | ZFP36L2 | 15427 | -1.168 | -0.3146 | Yes | ||
| 145 | SCHIP1 | 15484 | -1.185 | -0.3119 | Yes | ||
| 146 | BMI1 | 15504 | -1.189 | -0.3072 | Yes | ||
| 147 | FGFRL1 | 15512 | -1.191 | -0.3018 | Yes | ||
| 148 | DKK1 | 15538 | -1.200 | -0.2973 | Yes | ||
| 149 | TPM1 | 15553 | -1.203 | -0.2922 | Yes | ||
| 150 | HOXD9 | 15600 | -1.214 | -0.2888 | Yes | ||
| 151 | TSHB | 15696 | -1.240 | -0.2880 | Yes | ||
| 152 | ARHGEF2 | 15769 | -1.259 | -0.2858 | Yes | ||
| 153 | CCL7 | 15949 | -1.311 | -0.2891 | Yes | ||
| 154 | PURA | 16040 | -1.338 | -0.2875 | Yes | ||
| 155 | ASB16 | 16118 | -1.363 | -0.2851 | Yes | ||
| 156 | DDX5 | 16177 | -1.384 | -0.2815 | Yes | ||
| 157 | PPP1R10 | 16336 | -1.441 | -0.2831 | Yes | ||
| 158 | HIST1H1T | 16398 | -1.463 | -0.2793 | Yes | ||
| 159 | SOX3 | 16406 | -1.467 | -0.2725 | Yes | ||
| 160 | FGF16 | 16598 | -1.541 | -0.2754 | Yes | ||
| 161 | EIF4A2 | 16655 | -1.564 | -0.2709 | Yes | ||
| 162 | TCTA | 16850 | -1.650 | -0.2734 | Yes | ||
| 163 | PABPC4 | 16951 | -1.695 | -0.2706 | Yes | ||
| 164 | SERPINH1 | 17038 | -1.745 | -0.2668 | Yes | ||
| 165 | EEF2 | 17044 | -1.747 | -0.2585 | Yes | ||
| 166 | ESRRG | 17214 | -1.841 | -0.2588 | Yes | ||
| 167 | RPL10 | 17249 | -1.866 | -0.2516 | Yes | ||
| 168 | TSC1 | 17282 | -1.889 | -0.2441 | Yes | ||
| 169 | KRTAP21-1 | 17377 | -1.939 | -0.2398 | Yes | ||
| 170 | ADAMTS1 | 17448 | -1.991 | -0.2339 | Yes | ||
| 171 | ACTA1 | 17524 | -2.040 | -0.2281 | Yes | ||
| 172 | MT3 | 17559 | -2.069 | -0.2199 | Yes | ||
| 173 | MRPS18B | 17587 | -2.099 | -0.2112 | Yes | ||
| 174 | AGT | 17635 | -2.132 | -0.2034 | Yes | ||
| 175 | RLN3 | 17657 | -2.144 | -0.1941 | Yes | ||
| 176 | DIO2 | 17705 | -2.181 | -0.1861 | Yes | ||
| 177 | NASP | 17847 | -2.317 | -0.1825 | Yes | ||
| 178 | CFL2 | 17862 | -2.341 | -0.1718 | Yes | ||
| 179 | PRDX1 | 17924 | -2.412 | -0.1634 | Yes | ||
| 180 | HDLBP | 18009 | -2.536 | -0.1557 | Yes | ||
| 181 | BZW2 | 18011 | -2.539 | -0.1434 | Yes | ||
| 182 | IL10 | 18075 | -2.641 | -0.1340 | Yes | ||
| 183 | CRYAB | 18125 | -2.725 | -0.1234 | Yes | ||
| 184 | CSNK1E | 18143 | -2.747 | -0.1110 | Yes | ||
| 185 | CCL5 | 18148 | -2.750 | -0.0979 | Yes | ||
| 186 | HOXC4 | 18197 | -2.860 | -0.0866 | Yes | ||
| 187 | RGS1 | 18213 | -2.890 | -0.0734 | Yes | ||
| 188 | CDC25B | 18309 | -3.141 | -0.0633 | Yes | ||
| 189 | DIRAS1 | 18385 | -3.415 | -0.0508 | Yes | ||
| 190 | CA2 | 18421 | -3.585 | -0.0353 | Yes | ||
| 191 | TMPO | 18435 | -3.645 | -0.0183 | Yes | ||
| 192 | TNFSF10 | 18575 | -5.771 | 0.0022 | Yes |
