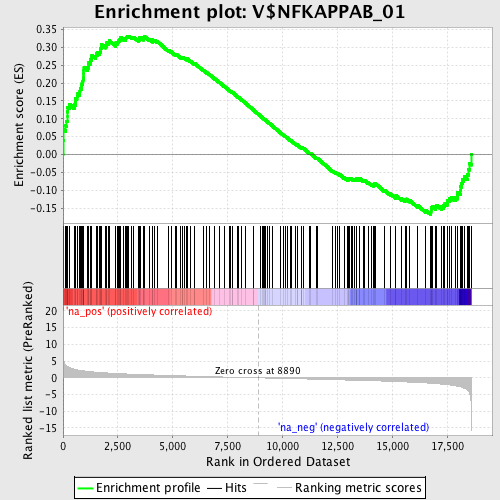
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | set04_DMproB_versus_LMproB |
| Phenotype | NoPhenotypeAvailable |
| Upregulated in class | na_pos |
| GeneSet | V$NFKAPPAB_01 |
| Enrichment Score (ES) | 0.33138192 |
| Normalized Enrichment Score (NES) | 1.5460604 |
| Nominal p-value | 0.0 |
| FDR q-value | 0.2512981 |
| FWER p-Value | 0.916 |

| PROBE | GENE SYMBOL | GENE_TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|---|
| 1 | EBI3 | 2 | 9.514 | 0.0391 | Yes | ||
| 2 | WNT10B | 11 | 7.646 | 0.0703 | Yes | ||
| 3 | VCAM1 | 121 | 3.849 | 0.0802 | Yes | ||
| 4 | RELB | 157 | 3.603 | 0.0932 | Yes | ||
| 5 | MADCAM1 | 187 | 3.449 | 0.1058 | Yes | ||
| 6 | BCL3 | 191 | 3.413 | 0.1198 | Yes | ||
| 7 | LTB | 209 | 3.330 | 0.1326 | Yes | ||
| 8 | CREB1 | 297 | 3.010 | 0.1403 | Yes | ||
| 9 | RPS6KA4 | 507 | 2.577 | 0.1396 | Yes | ||
| 10 | SLC6A12 | 554 | 2.509 | 0.1474 | Yes | ||
| 11 | MMP9 | 571 | 2.482 | 0.1568 | Yes | ||
| 12 | IL27RA | 633 | 2.410 | 0.1634 | Yes | ||
| 13 | NFKB2 | 672 | 2.356 | 0.1711 | Yes | ||
| 14 | ASH1L | 763 | 2.224 | 0.1754 | Yes | ||
| 15 | RRAS | 770 | 2.216 | 0.1842 | Yes | ||
| 16 | TNFRSF1B | 830 | 2.158 | 0.1899 | Yes | ||
| 17 | EDN2 | 855 | 2.140 | 0.1974 | Yes | ||
| 18 | CYLD | 888 | 2.101 | 0.2044 | Yes | ||
| 19 | NFAT5 | 907 | 2.083 | 0.2120 | Yes | ||
| 20 | IL27 | 937 | 2.061 | 0.2189 | Yes | ||
| 21 | HOXA11 | 940 | 2.059 | 0.2273 | Yes | ||
| 22 | IRF1 | 946 | 2.056 | 0.2355 | Yes | ||
| 23 | JAK3 | 951 | 2.053 | 0.2437 | Yes | ||
| 24 | PRX | 1111 | 1.921 | 0.2430 | Yes | ||
| 25 | TNIP1 | 1141 | 1.904 | 0.2493 | Yes | ||
| 26 | FGF1 | 1151 | 1.898 | 0.2567 | Yes | ||
| 27 | PPARBP | 1231 | 1.842 | 0.2600 | Yes | ||
| 28 | HIVEP1 | 1253 | 1.831 | 0.2664 | Yes | ||
| 29 | EGF | 1302 | 1.804 | 0.2712 | Yes | ||
| 30 | CXCL2 | 1311 | 1.800 | 0.2782 | Yes | ||
| 31 | CDC14A | 1498 | 1.677 | 0.2751 | Yes | ||
| 32 | DCAMKL1 | 1499 | 1.677 | 0.2820 | Yes | ||
| 33 | NTN1 | 1576 | 1.639 | 0.2846 | Yes | ||
| 34 | GATA4 | 1675 | 1.590 | 0.2859 | Yes | ||
| 35 | PLAU | 1699 | 1.579 | 0.2911 | Yes | ||
| 36 | SOX5 | 1703 | 1.576 | 0.2975 | Yes | ||
| 37 | IL4I1 | 1735 | 1.557 | 0.3022 | Yes | ||
| 38 | MAP3K8 | 1750 | 1.552 | 0.3079 | Yes | ||
| 39 | FOXJ2 | 1932 | 1.477 | 0.3041 | Yes | ||
| 40 | ARPC2 | 1993 | 1.455 | 0.3069 | Yes | ||
| 41 | CXCL9 | 1995 | 1.453 | 0.3128 | Yes | ||
| 42 | ETV6 | 2086 | 1.415 | 0.3138 | Yes | ||
| 43 | TNFRSF9 | 2105 | 1.408 | 0.3186 | Yes | ||
| 44 | TCEA2 | 2406 | 1.305 | 0.3077 | Yes | ||
| 45 | GREM1 | 2408 | 1.304 | 0.3130 | Yes | ||
| 46 | RALGDS | 2481 | 1.279 | 0.3144 | Yes | ||
| 47 | UACA | 2513 | 1.270 | 0.3180 | Yes | ||
| 48 | KLK9 | 2555 | 1.256 | 0.3209 | Yes | ||
| 49 | PUNC | 2613 | 1.239 | 0.3229 | Yes | ||
| 50 | IL23A | 2622 | 1.235 | 0.3276 | Yes | ||
| 51 | BNC2 | 2759 | 1.193 | 0.3252 | Yes | ||
| 52 | HCST | 2857 | 1.165 | 0.3247 | Yes | ||
| 53 | HSD11B2 | 2869 | 1.161 | 0.3289 | Yes | ||
| 54 | SAMSN1 | 2915 | 1.148 | 0.3312 | Yes | ||
| 55 | TNF | 2999 | 1.123 | 0.3313 | Yes | ||
| 56 | HCFC1 | 3138 | 1.088 | 0.3283 | Yes | ||
| 57 | TRAF4 | 3217 | 1.066 | 0.3285 | Yes | ||
| 58 | SOX10 | 3429 | 1.011 | 0.3212 | Yes | ||
| 59 | TAL1 | 3464 | 0.999 | 0.3235 | Yes | ||
| 60 | TNFSF15 | 3475 | 0.997 | 0.3271 | Yes | ||
| 61 | KCNS3 | 3542 | 0.977 | 0.3275 | Yes | ||
| 62 | RND1 | 3675 | 0.945 | 0.3242 | Yes | ||
| 63 | MAP4K2 | 3677 | 0.945 | 0.3281 | Yes | ||
| 64 | C1QL1 | 3689 | 0.942 | 0.3314 | Yes | ||
| 65 | CXCL10 | 3936 | 0.888 | 0.3217 | No | ||
| 66 | HSD3B7 | 4068 | 0.858 | 0.3181 | No | ||
| 67 | HOXB9 | 4081 | 0.854 | 0.3210 | No | ||
| 68 | ARHGAP8 | 4185 | 0.831 | 0.3188 | No | ||
| 69 | SESN2 | 4295 | 0.799 | 0.3162 | No | ||
| 70 | IFNB1 | 4818 | 0.692 | 0.2908 | No | ||
| 71 | BIRC3 | 4919 | 0.675 | 0.2881 | No | ||
| 72 | SP6 | 5104 | 0.637 | 0.2808 | No | ||
| 73 | STAT6 | 5172 | 0.625 | 0.2797 | No | ||
| 74 | ARHGAP5 | 5357 | 0.592 | 0.2722 | No | ||
| 75 | UBE4B | 5435 | 0.577 | 0.2704 | No | ||
| 76 | SHOX2 | 5439 | 0.575 | 0.2726 | No | ||
| 77 | SLAMF8 | 5530 | 0.557 | 0.2700 | No | ||
| 78 | SEC14L2 | 5634 | 0.540 | 0.2666 | No | ||
| 79 | USP52 | 5651 | 0.539 | 0.2680 | No | ||
| 80 | TLX1 | 5808 | 0.507 | 0.2616 | No | ||
| 81 | LTA | 5983 | 0.476 | 0.2542 | No | ||
| 82 | GADD45B | 5988 | 0.476 | 0.2559 | No | ||
| 83 | SMPD3 | 6384 | 0.404 | 0.2361 | No | ||
| 84 | COL11A2 | 6544 | 0.377 | 0.2291 | No | ||
| 85 | RAP2C | 6651 | 0.363 | 0.2248 | No | ||
| 86 | FLRT1 | 6918 | 0.318 | 0.2117 | No | ||
| 87 | GNAO1 | 7105 | 0.281 | 0.2028 | No | ||
| 88 | SMOC1 | 7346 | 0.242 | 0.1908 | No | ||
| 89 | RBMS1 | 7561 | 0.211 | 0.1800 | No | ||
| 90 | NR2F2 | 7629 | 0.200 | 0.1772 | No | ||
| 91 | KCNH3 | 7697 | 0.189 | 0.1743 | No | ||
| 92 | SIN3A | 7726 | 0.184 | 0.1736 | No | ||
| 93 | RIN2 | 7968 | 0.144 | 0.1611 | No | ||
| 94 | CXCL5 | 8008 | 0.137 | 0.1596 | No | ||
| 95 | MSC | 8142 | 0.117 | 0.1528 | No | ||
| 96 | ATP1B1 | 8321 | 0.092 | 0.1435 | No | ||
| 97 | CD83 | 8678 | 0.036 | 0.1244 | No | ||
| 98 | UBE2D3 | 8697 | 0.033 | 0.1235 | No | ||
| 99 | WNT10A | 9010 | -0.017 | 0.1067 | No | ||
| 100 | IL2RA | 9084 | -0.029 | 0.1028 | No | ||
| 101 | NFKBIB | 9110 | -0.033 | 0.1016 | No | ||
| 102 | STC2 | 9167 | -0.041 | 0.0988 | No | ||
| 103 | JARID2 | 9203 | -0.046 | 0.0971 | No | ||
| 104 | UBD | 9242 | -0.052 | 0.0952 | No | ||
| 105 | CXCL11 | 9327 | -0.066 | 0.0909 | No | ||
| 106 | ENO3 | 9385 | -0.075 | 0.0881 | No | ||
| 107 | ICAM1 | 9540 | -0.096 | 0.0802 | No | ||
| 108 | BCKDK | 9918 | -0.155 | 0.0604 | No | ||
| 109 | KRTAP13-1 | 10050 | -0.175 | 0.0540 | No | ||
| 110 | BMF | 10125 | -0.184 | 0.0507 | No | ||
| 111 | BFSP1 | 10215 | -0.197 | 0.0467 | No | ||
| 112 | CTGF | 10377 | -0.220 | 0.0389 | No | ||
| 113 | COL16A1 | 10424 | -0.227 | 0.0373 | No | ||
| 114 | IL1RN | 10598 | -0.259 | 0.0290 | No | ||
| 115 | BDNF | 10661 | -0.271 | 0.0268 | No | ||
| 116 | AP1S2 | 10671 | -0.272 | 0.0274 | No | ||
| 117 | SLC11A2 | 10849 | -0.300 | 0.0190 | No | ||
| 118 | CLCN1 | 10869 | -0.303 | 0.0193 | No | ||
| 119 | ATOH1 | 10881 | -0.305 | 0.0199 | No | ||
| 120 | TRIM47 | 10942 | -0.315 | 0.0180 | No | ||
| 121 | TNFSF18 | 11246 | -0.369 | 0.0030 | No | ||
| 122 | RIMS2 | 11271 | -0.373 | 0.0033 | No | ||
| 123 | FGF17 | 11558 | -0.417 | -0.0105 | No | ||
| 124 | MOBKL2C | 11583 | -0.421 | -0.0101 | No | ||
| 125 | LAMA1 | 12299 | -0.538 | -0.0466 | No | ||
| 126 | AGPAT1 | 12418 | -0.558 | -0.0507 | No | ||
| 127 | PCSK2 | 12491 | -0.575 | -0.0523 | No | ||
| 128 | SALL1 | 12605 | -0.597 | -0.0559 | No | ||
| 129 | CPD | 12809 | -0.630 | -0.0644 | No | ||
| 130 | IER5 | 12967 | -0.661 | -0.0701 | No | ||
| 131 | UBE2I | 12997 | -0.667 | -0.0690 | No | ||
| 132 | IL1A | 12999 | -0.667 | -0.0663 | No | ||
| 133 | SLC12A2 | 13074 | -0.680 | -0.0675 | No | ||
| 134 | FUT7 | 13141 | -0.690 | -0.0682 | No | ||
| 135 | MAG | 13208 | -0.698 | -0.0689 | No | ||
| 136 | ALG6 | 13283 | -0.710 | -0.0700 | No | ||
| 137 | FTHL17 | 13351 | -0.721 | -0.0707 | No | ||
| 138 | IL13 | 13353 | -0.721 | -0.0677 | No | ||
| 139 | EHF | 13386 | -0.727 | -0.0665 | No | ||
| 140 | FKHL18 | 13510 | -0.753 | -0.0700 | No | ||
| 141 | BMP2K | 13521 | -0.755 | -0.0675 | No | ||
| 142 | ACTN3 | 13692 | -0.784 | -0.0735 | No | ||
| 143 | TAZ | 13726 | -0.792 | -0.0720 | No | ||
| 144 | PFN1 | 13903 | -0.822 | -0.0781 | No | ||
| 145 | GABRB1 | 14065 | -0.855 | -0.0833 | No | ||
| 146 | TRIB2 | 14161 | -0.874 | -0.0849 | No | ||
| 147 | MSX1 | 14169 | -0.876 | -0.0817 | No | ||
| 148 | SDC4 | 14245 | -0.891 | -0.0820 | No | ||
| 149 | ASCL3 | 14668 | -0.982 | -0.1009 | No | ||
| 150 | DDR1 | 14915 | -1.040 | -0.1099 | No | ||
| 151 | NOL4 | 15162 | -1.099 | -0.1187 | No | ||
| 152 | MSN | 15168 | -1.102 | -0.1145 | No | ||
| 153 | ZFP36L2 | 15427 | -1.168 | -0.1237 | No | ||
| 154 | BCL6B | 15586 | -1.211 | -0.1272 | No | ||
| 155 | BAI2 | 15630 | -1.219 | -0.1245 | No | ||
| 156 | GRIN2D | 15788 | -1.264 | -0.1278 | No | ||
| 157 | ZBTB11 | 16152 | -1.373 | -0.1419 | No | ||
| 158 | CHD4 | 16518 | -1.510 | -0.1554 | No | ||
| 159 | CDK6 | 16749 | -1.603 | -0.1613 | No | ||
| 160 | KCNN3 | 16768 | -1.611 | -0.1556 | No | ||
| 161 | NLK | 16771 | -1.613 | -0.1491 | No | ||
| 162 | GRK5 | 16830 | -1.642 | -0.1455 | No | ||
| 163 | DAP3 | 16983 | -1.711 | -0.1466 | No | ||
| 164 | P2RY10 | 17025 | -1.736 | -0.1417 | No | ||
| 165 | CHX10 | 17251 | -1.867 | -0.1462 | No | ||
| 166 | ERN1 | 17327 | -1.912 | -0.1424 | No | ||
| 167 | ZBTB5 | 17402 | -1.949 | -0.1384 | No | ||
| 168 | CXCL16 | 17500 | -2.023 | -0.1353 | No | ||
| 169 | PCBP4 | 17517 | -2.033 | -0.1278 | No | ||
| 170 | LIG1 | 17601 | -2.111 | -0.1236 | No | ||
| 171 | UBE2H | 17683 | -2.162 | -0.1190 | No | ||
| 172 | GNB1 | 17881 | -2.364 | -0.1200 | No | ||
| 173 | FBXL12 | 17968 | -2.477 | -0.1144 | No | ||
| 174 | ITGB4 | 17987 | -2.505 | -0.1051 | No | ||
| 175 | BLR1 | 18105 | -2.688 | -0.1003 | No | ||
| 176 | TSLP | 18123 | -2.724 | -0.0900 | No | ||
| 177 | CCL5 | 18148 | -2.750 | -0.0800 | No | ||
| 178 | GPHN | 18196 | -2.860 | -0.0707 | No | ||
| 179 | EDG5 | 18271 | -3.047 | -0.0622 | No | ||
| 180 | ABHD8 | 18439 | -3.667 | -0.0561 | No | ||
| 181 | EIF5A | 18481 | -4.050 | -0.0416 | No | ||
| 182 | GNG4 | 18518 | -4.441 | -0.0252 | No | ||
| 183 | TP53 | 18597 | -7.392 | 0.0010 | No |
