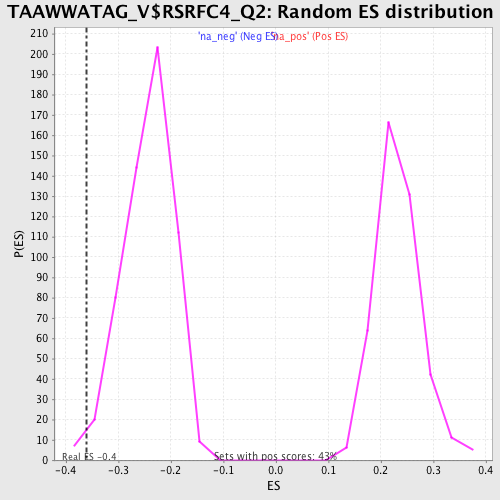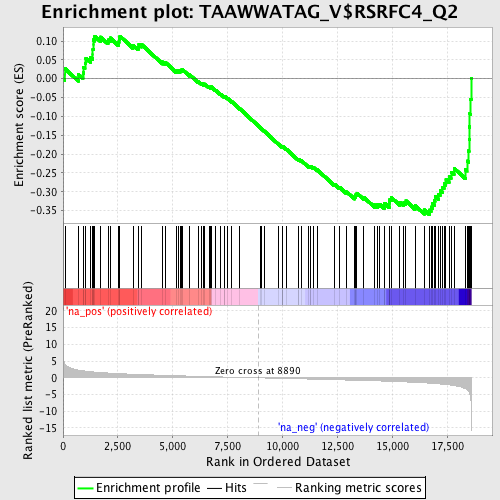
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | set04_DMproB_versus_LMproB |
| Phenotype | NoPhenotypeAvailable |
| Upregulated in class | na_neg |
| GeneSet | TAAWWATAG_V$RSRFC4_Q2 |
| Enrichment Score (ES) | -0.36019573 |
| Normalized Enrichment Score (NES) | -1.4810963 |
| Nominal p-value | 0.012173913 |
| FDR q-value | 0.4449073 |
| FWER p-Value | 0.996 |

| PROBE | GENE SYMBOL | GENE_TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|---|
| 1 | PRDX5 | 86 | 4.139 | 0.0266 | No | ||
| 2 | DNAJA4 | 698 | 2.306 | 0.0111 | No | ||
| 3 | NFAT5 | 907 | 2.083 | 0.0156 | No | ||
| 4 | TACSTD2 | 927 | 2.068 | 0.0302 | No | ||
| 5 | ALDOA | 998 | 2.006 | 0.0415 | No | ||
| 6 | CMYA1 | 1037 | 1.983 | 0.0545 | No | ||
| 7 | TNNC1 | 1239 | 1.840 | 0.0575 | No | ||
| 8 | MAT2A | 1356 | 1.763 | 0.0646 | No | ||
| 9 | SATB2 | 1358 | 1.762 | 0.0778 | No | ||
| 10 | NOG | 1379 | 1.749 | 0.0900 | No | ||
| 11 | NRP2 | 1381 | 1.748 | 0.1031 | No | ||
| 12 | ATP2A3 | 1443 | 1.711 | 0.1127 | No | ||
| 13 | SOX5 | 1703 | 1.576 | 0.1107 | No | ||
| 14 | TCEA3 | 2048 | 1.431 | 0.1029 | No | ||
| 15 | GRB14 | 2147 | 1.397 | 0.1082 | No | ||
| 16 | USP2 | 2524 | 1.266 | 0.0974 | No | ||
| 17 | EDEM1 | 2549 | 1.258 | 0.1056 | No | ||
| 18 | SMPX | 2588 | 1.246 | 0.1130 | No | ||
| 19 | SNAP25 | 3186 | 1.076 | 0.0889 | No | ||
| 20 | MYOCD | 3421 | 1.014 | 0.0839 | No | ||
| 21 | MUSK | 3453 | 1.005 | 0.0898 | No | ||
| 22 | BEX2 | 3561 | 0.972 | 0.0914 | No | ||
| 23 | FOXP1 | 4533 | 0.750 | 0.0446 | No | ||
| 24 | CDKN1A | 4661 | 0.724 | 0.0432 | No | ||
| 25 | USP13 | 5149 | 0.629 | 0.0217 | No | ||
| 26 | ARPC5L | 5238 | 0.613 | 0.0215 | No | ||
| 27 | LIF | 5329 | 0.596 | 0.0212 | No | ||
| 28 | DLC1 | 5373 | 0.588 | 0.0233 | No | ||
| 29 | SHOX2 | 5439 | 0.575 | 0.0241 | No | ||
| 30 | KCNA3 | 5765 | 0.516 | 0.0105 | No | ||
| 31 | YARS | 6155 | 0.446 | -0.0072 | No | ||
| 32 | MID2 | 6317 | 0.415 | -0.0127 | No | ||
| 33 | NEB | 6403 | 0.400 | -0.0143 | No | ||
| 34 | MYL3 | 6452 | 0.391 | -0.0139 | No | ||
| 35 | RAP2C | 6651 | 0.363 | -0.0219 | No | ||
| 36 | COL1A1 | 6717 | 0.353 | -0.0227 | No | ||
| 37 | TCF4 | 6744 | 0.349 | -0.0215 | No | ||
| 38 | TRPS1 | 6947 | 0.313 | -0.0300 | No | ||
| 39 | GNB4 | 7185 | 0.269 | -0.0408 | No | ||
| 40 | KCNJ9 | 7341 | 0.243 | -0.0473 | No | ||
| 41 | KCNN1 | 7370 | 0.238 | -0.0471 | No | ||
| 42 | STAG2 | 7498 | 0.220 | -0.0523 | No | ||
| 43 | PDGFRA | 7692 | 0.189 | -0.0613 | No | ||
| 44 | CENTD1 | 8057 | 0.129 | -0.0799 | No | ||
| 45 | SLC2A4 | 8997 | -0.015 | -0.1305 | No | ||
| 46 | CPNE7 | 9063 | -0.025 | -0.1339 | No | ||
| 47 | TNNI1 | 9158 | -0.040 | -0.1386 | No | ||
| 48 | PI15 | 9828 | -0.140 | -0.1737 | No | ||
| 49 | TNNC2 | 9986 | -0.164 | -0.1810 | No | ||
| 50 | GPR30 | 9998 | -0.166 | -0.1803 | No | ||
| 51 | KCNA7 | 10021 | -0.171 | -0.1802 | No | ||
| 52 | CAPN3 | 10181 | -0.193 | -0.1873 | No | ||
| 53 | NR4A1 | 10721 | -0.280 | -0.2143 | No | ||
| 54 | HOXA4 | 10744 | -0.283 | -0.2134 | No | ||
| 55 | COL8A1 | 10844 | -0.299 | -0.2165 | No | ||
| 56 | ZHX2 | 11202 | -0.361 | -0.2330 | No | ||
| 57 | SV2A | 11260 | -0.371 | -0.2333 | No | ||
| 58 | SLC12A8 | 11293 | -0.375 | -0.2322 | No | ||
| 59 | CPT1B | 11410 | -0.395 | -0.2355 | No | ||
| 60 | SEMA6A | 11585 | -0.421 | -0.2417 | No | ||
| 61 | LRRC20 | 12376 | -0.552 | -0.2802 | No | ||
| 62 | SALL1 | 12605 | -0.597 | -0.2880 | No | ||
| 63 | HFE2 | 12924 | -0.652 | -0.3002 | No | ||
| 64 | SDHC | 13276 | -0.709 | -0.3138 | No | ||
| 65 | MYOG | 13314 | -0.715 | -0.3104 | No | ||
| 66 | WBP4 | 13346 | -0.721 | -0.3067 | No | ||
| 67 | NRXN3 | 13391 | -0.728 | -0.3035 | No | ||
| 68 | PACS1 | 13694 | -0.784 | -0.3139 | No | ||
| 69 | RAP2B | 14201 | -0.882 | -0.3346 | No | ||
| 70 | ADAM23 | 14326 | -0.909 | -0.3344 | No | ||
| 71 | HOXA9 | 14426 | -0.928 | -0.3328 | No | ||
| 72 | CKMT2 | 14633 | -0.975 | -0.3365 | No | ||
| 73 | SLC8A3 | 14645 | -0.978 | -0.3297 | No | ||
| 74 | PPARGC1A | 14863 | -1.028 | -0.3337 | No | ||
| 75 | SLN | 14866 | -1.029 | -0.3260 | No | ||
| 76 | PER1 | 14893 | -1.035 | -0.3196 | No | ||
| 77 | TPP2 | 14966 | -1.049 | -0.3156 | No | ||
| 78 | KTN1 | 15352 | -1.149 | -0.3277 | No | ||
| 79 | MYL1 | 15532 | -1.198 | -0.3283 | No | ||
| 80 | CLDN14 | 15627 | -1.218 | -0.3242 | No | ||
| 81 | PURA | 16040 | -1.338 | -0.3363 | No | ||
| 82 | ELAVL4 | 16464 | -1.492 | -0.3479 | No | ||
| 83 | TYRO3 | 16693 | -1.579 | -0.3483 | Yes | ||
| 84 | NLK | 16771 | -1.613 | -0.3402 | Yes | ||
| 85 | FGF13 | 16847 | -1.650 | -0.3318 | Yes | ||
| 86 | PCDH9 | 16924 | -1.682 | -0.3232 | Yes | ||
| 87 | PABPC4 | 16951 | -1.695 | -0.3118 | Yes | ||
| 88 | RASGEF1B | 17107 | -1.782 | -0.3067 | Yes | ||
| 89 | ESR1 | 17182 | -1.821 | -0.2970 | Yes | ||
| 90 | ALS2CR2 | 17289 | -1.893 | -0.2884 | Yes | ||
| 91 | CRTAP | 17364 | -1.932 | -0.2778 | Yes | ||
| 92 | PDLIM1 | 17450 | -1.992 | -0.2673 | Yes | ||
| 93 | CUTL1 | 17590 | -2.100 | -0.2590 | Yes | ||
| 94 | DSPP | 17682 | -2.162 | -0.2475 | Yes | ||
| 95 | LECT1 | 17818 | -2.287 | -0.2376 | Yes | ||
| 96 | JUN | 18337 | -3.236 | -0.2411 | Yes | ||
| 97 | HIBADH | 18409 | -3.532 | -0.2182 | Yes | ||
| 98 | EIF5A | 18481 | -4.050 | -0.1915 | Yes | ||
| 99 | GRIA3 | 18502 | -4.254 | -0.1604 | Yes | ||
| 100 | INPPL1 | 18516 | -4.435 | -0.1276 | Yes | ||
| 101 | RALY | 18536 | -4.695 | -0.0931 | Yes | ||
| 102 | LZTS2 | 18559 | -5.300 | -0.0543 | Yes | ||
| 103 | GPR23 | 18599 | -7.586 | 0.0009 | Yes |