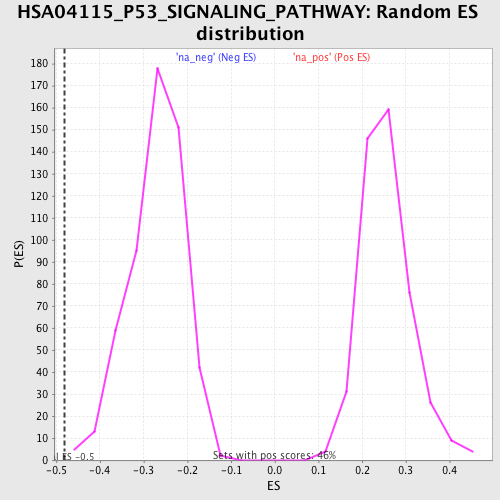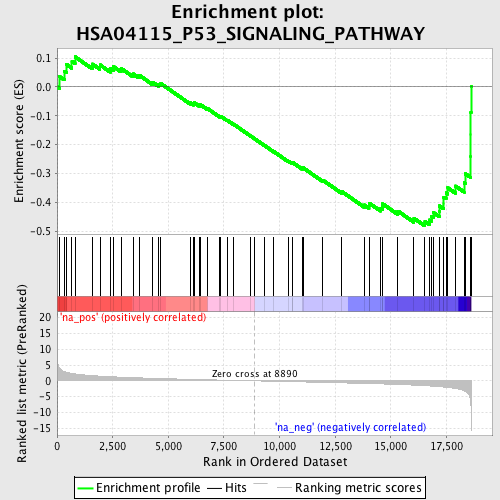
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | set04_DMproB_versus_LMproB |
| Phenotype | NoPhenotypeAvailable |
| Upregulated in class | na_neg |
| GeneSet | HSA04115_P53_SIGNALING_PATHWAY |
| Enrichment Score (ES) | -0.48204514 |
| Normalized Enrichment Score (NES) | -1.7861943 |
| Nominal p-value | 0.0018348624 |
| FDR q-value | 0.26696214 |
| FWER p-Value | 0.666 |

| PROBE | GENE SYMBOL | GENE_TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|---|
| 1 | CD82 | 103 | 3.993 | 0.0358 | No | ||
| 2 | GADD45G | 319 | 2.953 | 0.0548 | No | ||
| 3 | SCOTIN | 415 | 2.742 | 0.0781 | No | ||
| 4 | CDK2 | 668 | 2.362 | 0.0890 | No | ||
| 5 | CCND3 | 808 | 2.176 | 0.1040 | No | ||
| 6 | SFN | 1582 | 1.636 | 0.0793 | No | ||
| 7 | BID | 1927 | 1.480 | 0.0761 | No | ||
| 8 | ATM | 2389 | 1.310 | 0.0648 | No | ||
| 9 | THBS1 | 2527 | 1.265 | 0.0705 | No | ||
| 10 | TNFRSF10B | 2876 | 1.159 | 0.0638 | No | ||
| 11 | CCNB2 | 3422 | 1.014 | 0.0449 | No | ||
| 12 | MDM2 | 3684 | 0.943 | 0.0406 | No | ||
| 13 | SESN2 | 4295 | 0.799 | 0.0160 | No | ||
| 14 | CCNB1 | 4563 | 0.746 | 0.0094 | No | ||
| 15 | CDKN1A | 4661 | 0.724 | 0.0116 | No | ||
| 16 | GADD45B | 5988 | 0.476 | -0.0549 | No | ||
| 17 | MDM4 | 6141 | 0.449 | -0.0584 | No | ||
| 18 | CDKN2A | 6152 | 0.447 | -0.0543 | No | ||
| 19 | CCNE1 | 6421 | 0.397 | -0.0647 | No | ||
| 20 | ATR | 6423 | 0.397 | -0.0606 | No | ||
| 21 | IGFBP3 | 6767 | 0.345 | -0.0755 | No | ||
| 22 | CCND1 | 7311 | 0.248 | -0.1022 | No | ||
| 23 | GTSE1 | 7362 | 0.240 | -0.1024 | No | ||
| 24 | SERPINE1 | 7655 | 0.195 | -0.1161 | No | ||
| 25 | RCHY1 | 7917 | 0.154 | -0.1286 | No | ||
| 26 | PTEN | 8691 | 0.035 | -0.1699 | No | ||
| 27 | LRDD | 8887 | 0.000 | -0.1804 | No | ||
| 28 | RPRM | 9302 | -0.062 | -0.2021 | No | ||
| 29 | CCNB3 | 9745 | -0.126 | -0.2246 | No | ||
| 30 | CCNG2 | 10409 | -0.224 | -0.2580 | No | ||
| 31 | GADD45A | 10581 | -0.255 | -0.2646 | No | ||
| 32 | SERPINB5 | 10601 | -0.259 | -0.2629 | No | ||
| 33 | CHEK1 | 11033 | -0.331 | -0.2827 | No | ||
| 34 | CHEK2 | 11054 | -0.334 | -0.2803 | No | ||
| 35 | PERP | 11937 | -0.476 | -0.3229 | No | ||
| 36 | BAI1 | 12784 | -0.625 | -0.3621 | No | ||
| 37 | PPM1D | 13816 | -0.807 | -0.4093 | No | ||
| 38 | BBC3 | 14020 | -0.847 | -0.4114 | No | ||
| 39 | TSC2 | 14028 | -0.849 | -0.4030 | No | ||
| 40 | RRM2B | 14535 | -0.953 | -0.4204 | No | ||
| 41 | CYCS | 14621 | -0.973 | -0.4149 | No | ||
| 42 | SESN1 | 14636 | -0.976 | -0.4056 | No | ||
| 43 | CASP9 | 15313 | -1.138 | -0.4302 | No | ||
| 44 | APAF1 | 16038 | -1.338 | -0.4554 | No | ||
| 45 | RRM2 | 16534 | -1.516 | -0.4663 | Yes | ||
| 46 | CDK6 | 16749 | -1.603 | -0.4613 | Yes | ||
| 47 | EI24 | 16824 | -1.639 | -0.4483 | Yes | ||
| 48 | DDB2 | 16929 | -1.685 | -0.4364 | Yes | ||
| 49 | CASP8 | 17177 | -1.818 | -0.4309 | Yes | ||
| 50 | CCNE2 | 17187 | -1.827 | -0.4125 | Yes | ||
| 51 | SESN3 | 17381 | -1.940 | -0.4028 | Yes | ||
| 52 | CCND2 | 17382 | -1.941 | -0.3827 | Yes | ||
| 53 | FAS | 17486 | -2.013 | -0.3674 | Yes | ||
| 54 | PMAIP1 | 17564 | -2.073 | -0.3501 | Yes | ||
| 55 | CASP3 | 17902 | -2.386 | -0.3435 | Yes | ||
| 56 | IGF1 | 18296 | -3.101 | -0.3326 | Yes | ||
| 57 | CDK4 | 18355 | -3.310 | -0.3014 | Yes | ||
| 58 | CCNG1 | 18592 | -6.932 | -0.2423 | Yes | ||
| 59 | TP53 | 18597 | -7.392 | -0.1660 | Yes | ||
| 60 | BAX | 18598 | -7.485 | -0.0884 | Yes | ||
| 61 | ZMAT3 | 18608 | -8.626 | 0.0004 | Yes |