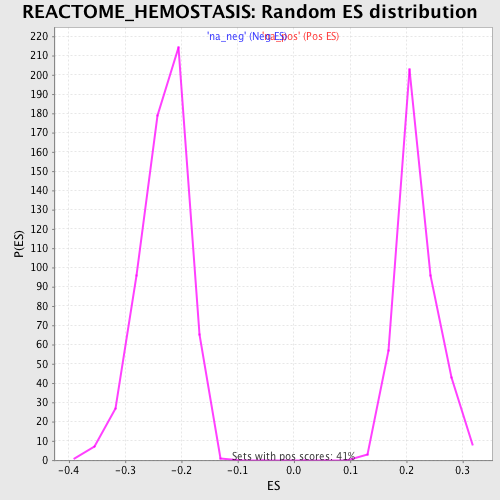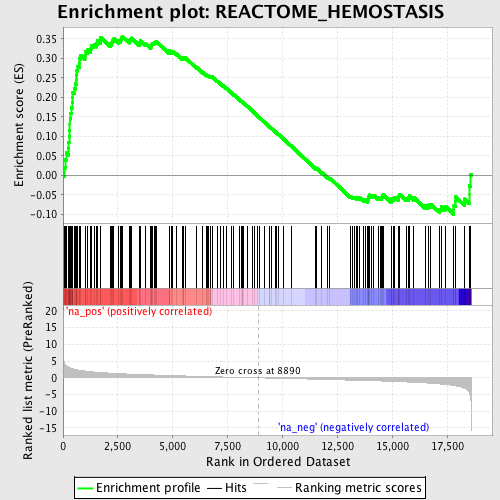
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | set04_DMproB_versus_LMproB |
| Phenotype | NoPhenotypeAvailable |
| Upregulated in class | na_pos |
| GeneSet | REACTOME_HEMOSTASIS |
| Enrichment Score (ES) | 0.35723034 |
| Normalized Enrichment Score (NES) | 1.6426725 |
| Nominal p-value | 0.0 |
| FDR q-value | 0.31064487 |
| FWER p-Value | 0.997 |

| PROBE | GENE SYMBOL | GENE_TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|---|
| 1 | ITGAL | 56 | 4.531 | 0.0211 | Yes | ||
| 2 | F10 | 102 | 3.994 | 0.0399 | Yes | ||
| 3 | CLU | 137 | 3.748 | 0.0581 | Yes | ||
| 4 | VCL | 249 | 3.154 | 0.0689 | Yes | ||
| 5 | LCP2 | 257 | 3.134 | 0.0852 | Yes | ||
| 6 | PLAUR | 272 | 3.074 | 0.1008 | Yes | ||
| 7 | GP5 | 287 | 3.026 | 0.1161 | Yes | ||
| 8 | MMRN1 | 311 | 2.984 | 0.1308 | Yes | ||
| 9 | CD63 | 313 | 2.969 | 0.1466 | Yes | ||
| 10 | APP | 352 | 2.860 | 0.1597 | Yes | ||
| 11 | FN1 | 371 | 2.825 | 0.1738 | Yes | ||
| 12 | TBXA2R | 408 | 2.752 | 0.1865 | Yes | ||
| 13 | FCER1G | 425 | 2.727 | 0.2002 | Yes | ||
| 14 | FLNA | 445 | 2.691 | 0.2135 | Yes | ||
| 15 | DOK2 | 523 | 2.555 | 0.2229 | Yes | ||
| 16 | SELL | 553 | 2.512 | 0.2347 | Yes | ||
| 17 | GP9 | 607 | 2.429 | 0.2448 | Yes | ||
| 18 | GP1BA | 612 | 2.427 | 0.2575 | Yes | ||
| 19 | PTPN6 | 628 | 2.412 | 0.2696 | Yes | ||
| 20 | PSAP | 675 | 2.347 | 0.2796 | Yes | ||
| 21 | SERPING1 | 737 | 2.257 | 0.2883 | Yes | ||
| 22 | F5 | 744 | 2.255 | 0.3000 | Yes | ||
| 23 | MAPK14 | 804 | 2.181 | 0.3084 | Yes | ||
| 24 | ALDOA | 998 | 2.006 | 0.3086 | Yes | ||
| 25 | ITGB2 | 1038 | 1.982 | 0.3171 | Yes | ||
| 26 | P2RY1 | 1130 | 1.913 | 0.3224 | Yes | ||
| 27 | PIK3CB | 1268 | 1.823 | 0.3246 | Yes | ||
| 28 | EGF | 1302 | 1.804 | 0.3325 | Yes | ||
| 29 | GNA13 | 1408 | 1.729 | 0.3360 | Yes | ||
| 30 | CD48 | 1534 | 1.658 | 0.3381 | Yes | ||
| 31 | PPIA | 1566 | 1.644 | 0.3451 | Yes | ||
| 32 | PLAU | 1699 | 1.579 | 0.3464 | Yes | ||
| 33 | CSK | 1719 | 1.565 | 0.3537 | Yes | ||
| 34 | GRB14 | 2147 | 1.397 | 0.3380 | Yes | ||
| 35 | WDR1 | 2226 | 1.363 | 0.3411 | Yes | ||
| 36 | P2RX1 | 2249 | 1.355 | 0.3471 | Yes | ||
| 37 | JAM2 | 2317 | 1.329 | 0.3506 | Yes | ||
| 38 | THBS1 | 2527 | 1.265 | 0.3460 | Yes | ||
| 39 | PLEK | 2624 | 1.235 | 0.3474 | Yes | ||
| 40 | FYB | 2640 | 1.227 | 0.3531 | Yes | ||
| 41 | PF4 | 2684 | 1.215 | 0.3572 | Yes | ||
| 42 | SPARC | 3013 | 1.119 | 0.3454 | No | ||
| 43 | PROC | 3059 | 1.107 | 0.3489 | No | ||
| 44 | KNG1 | 3103 | 1.097 | 0.3524 | No | ||
| 45 | THBD | 3477 | 0.996 | 0.3375 | No | ||
| 46 | STXBP3 | 3499 | 0.991 | 0.3417 | No | ||
| 47 | CFL1 | 3509 | 0.989 | 0.3464 | No | ||
| 48 | GP1BB | 3744 | 0.928 | 0.3387 | No | ||
| 49 | ITGA5 | 3995 | 0.876 | 0.3298 | No | ||
| 50 | OLR1 | 4037 | 0.865 | 0.3322 | No | ||
| 51 | GNAQ | 4047 | 0.863 | 0.3363 | No | ||
| 52 | COL1A2 | 4075 | 0.855 | 0.3394 | No | ||
| 53 | SELP | 4145 | 0.841 | 0.3402 | No | ||
| 54 | TFPI | 4190 | 0.830 | 0.3422 | No | ||
| 55 | PPBP | 4243 | 0.814 | 0.3437 | No | ||
| 56 | SOS1 | 4827 | 0.690 | 0.3158 | No | ||
| 57 | CD47 | 4828 | 0.690 | 0.3195 | No | ||
| 58 | PIK3CG | 4959 | 0.667 | 0.3160 | No | ||
| 59 | CD36 | 4978 | 0.663 | 0.3186 | No | ||
| 60 | ACTN4 | 5162 | 0.627 | 0.3120 | No | ||
| 61 | PLG | 5429 | 0.579 | 0.3007 | No | ||
| 62 | STX4 | 5501 | 0.564 | 0.2999 | No | ||
| 63 | PRKCZ | 5502 | 0.564 | 0.3029 | No | ||
| 64 | ITGAM | 5569 | 0.552 | 0.3022 | No | ||
| 65 | APBB1IP | 6090 | 0.458 | 0.2765 | No | ||
| 66 | F13B | 6369 | 0.406 | 0.2636 | No | ||
| 67 | PTK2 | 6518 | 0.380 | 0.2576 | No | ||
| 68 | HRG | 6602 | 0.369 | 0.2551 | No | ||
| 69 | PIK3R1 | 6623 | 0.366 | 0.2560 | No | ||
| 70 | C1QBP | 6696 | 0.356 | 0.2540 | No | ||
| 71 | COL1A1 | 6717 | 0.353 | 0.2548 | No | ||
| 72 | VWF | 6786 | 0.342 | 0.2529 | No | ||
| 73 | F12 | 7032 | 0.297 | 0.2412 | No | ||
| 74 | AKT1 | 7194 | 0.267 | 0.2339 | No | ||
| 75 | SLC16A3 | 7300 | 0.250 | 0.2296 | No | ||
| 76 | SYK | 7442 | 0.230 | 0.2232 | No | ||
| 77 | SERPINE1 | 7655 | 0.195 | 0.2127 | No | ||
| 78 | ITGB3 | 7783 | 0.175 | 0.2068 | No | ||
| 79 | PTPN1 | 8027 | 0.134 | 0.1944 | No | ||
| 80 | ITGAV | 8129 | 0.119 | 0.1895 | No | ||
| 81 | ADRA2A | 8167 | 0.114 | 0.1881 | No | ||
| 82 | A2M | 8173 | 0.113 | 0.1885 | No | ||
| 83 | GNG2 | 8236 | 0.104 | 0.1857 | No | ||
| 84 | CFD | 8394 | 0.081 | 0.1776 | No | ||
| 85 | LYN | 8625 | 0.044 | 0.1654 | No | ||
| 86 | F2RL2 | 8704 | 0.032 | 0.1613 | No | ||
| 87 | APOB | 8878 | 0.002 | 0.1520 | No | ||
| 88 | TEK | 8932 | -0.006 | 0.1491 | No | ||
| 89 | BCAR1 | 9181 | -0.043 | 0.1359 | No | ||
| 90 | PLCG1 | 9400 | -0.078 | 0.1246 | No | ||
| 91 | HGF | 9413 | -0.079 | 0.1243 | No | ||
| 92 | PLA2G4A | 9481 | -0.087 | 0.1212 | No | ||
| 93 | SERPINC1 | 9672 | -0.115 | 0.1115 | No | ||
| 94 | SOD1 | 9720 | -0.123 | 0.1096 | No | ||
| 95 | CD2 | 9806 | -0.136 | 0.1057 | No | ||
| 96 | F7 | 10041 | -0.174 | 0.0940 | No | ||
| 97 | ITGA2B | 10394 | -0.222 | 0.0761 | No | ||
| 98 | PLAT | 11506 | -0.408 | 0.0181 | No | ||
| 99 | FGB | 11533 | -0.413 | 0.0189 | No | ||
| 100 | CD9 | 11796 | -0.454 | 0.0071 | No | ||
| 101 | HSPA5 | 12061 | -0.497 | -0.0045 | No | ||
| 102 | TMSB4X | 12159 | -0.515 | -0.0070 | No | ||
| 103 | MERTK | 13098 | -0.683 | -0.0542 | No | ||
| 104 | MAG | 13208 | -0.698 | -0.0563 | No | ||
| 105 | F2R | 13282 | -0.710 | -0.0565 | No | ||
| 106 | F2 | 13369 | -0.725 | -0.0573 | No | ||
| 107 | LAMP2 | 13429 | -0.733 | -0.0566 | No | ||
| 108 | SPN | 13527 | -0.756 | -0.0578 | No | ||
| 109 | APOA1 | 13680 | -0.782 | -0.0619 | No | ||
| 110 | ITGAX | 13769 | -0.800 | -0.0624 | No | ||
| 111 | PLCG2 | 13893 | -0.822 | -0.0647 | No | ||
| 112 | PFN1 | 13903 | -0.822 | -0.0608 | No | ||
| 113 | KLKB1 | 13934 | -0.829 | -0.0580 | No | ||
| 114 | PPIL2 | 13935 | -0.829 | -0.0536 | No | ||
| 115 | CXADR | 13957 | -0.833 | -0.0503 | No | ||
| 116 | F9 | 14069 | -0.856 | -0.0517 | No | ||
| 117 | RAP1A | 14135 | -0.869 | -0.0506 | No | ||
| 118 | TUBA4A | 14357 | -0.915 | -0.0577 | No | ||
| 119 | CAV1 | 14452 | -0.935 | -0.0578 | No | ||
| 120 | F3 | 14525 | -0.951 | -0.0566 | No | ||
| 121 | ITGA4 | 14550 | -0.957 | -0.0528 | No | ||
| 122 | SERPINB2 | 14582 | -0.963 | -0.0494 | No | ||
| 123 | SERPINF2 | 14949 | -1.045 | -0.0636 | No | ||
| 124 | FGA | 14969 | -1.050 | -0.0591 | No | ||
| 125 | ANGPT2 | 15063 | -1.073 | -0.0584 | No | ||
| 126 | SHC1 | 15121 | -1.088 | -0.0557 | No | ||
| 127 | CALM1 | 15269 | -1.128 | -0.0576 | No | ||
| 128 | TIMP1 | 15305 | -1.137 | -0.0535 | No | ||
| 129 | LEFTY2 | 15325 | -1.141 | -0.0484 | No | ||
| 130 | KRAS | 15643 | -1.222 | -0.0591 | No | ||
| 131 | F11 | 15747 | -1.250 | -0.0580 | No | ||
| 132 | PDPK1 | 15766 | -1.257 | -0.0523 | No | ||
| 133 | BSG | 15986 | -1.322 | -0.0571 | No | ||
| 134 | GRB7 | 16520 | -1.511 | -0.0779 | No | ||
| 135 | CAP1 | 16637 | -1.558 | -0.0759 | No | ||
| 136 | ITGB1 | 16759 | -1.607 | -0.0738 | No | ||
| 137 | FGG | 17173 | -1.815 | -0.0865 | No | ||
| 138 | CALU | 17247 | -1.864 | -0.0806 | No | ||
| 139 | FYN | 17408 | -1.958 | -0.0788 | No | ||
| 140 | ANGPT1 | 17808 | -2.275 | -0.0883 | No | ||
| 141 | PTPN11 | 17811 | -2.276 | -0.0763 | No | ||
| 142 | GNB1 | 17881 | -2.364 | -0.0674 | No | ||
| 143 | VAV1 | 17890 | -2.372 | -0.0552 | No | ||
| 144 | IGF1 | 18296 | -3.101 | -0.0606 | No | ||
| 145 | F8 | 18508 | -4.299 | -0.0491 | No | ||
| 146 | GAS6 | 18510 | -4.358 | -0.0260 | No | ||
| 147 | PROS1 | 18581 | -5.944 | 0.0019 | No |