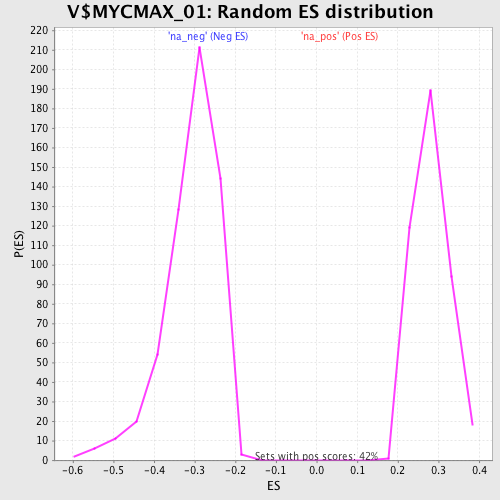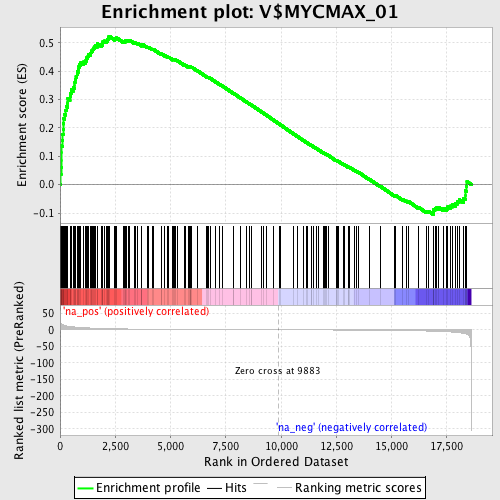
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | set04_DMpreB_versus_WTpreB |
| Phenotype | NoPhenotypeAvailable |
| Upregulated in class | na_pos |
| GeneSet | V$MYCMAX_01 |
| Enrichment Score (ES) | 0.5236071 |
| Normalized Enrichment Score (NES) | 1.8611715 |
| Nominal p-value | 0.0 |
| FDR q-value | 0.0115322545 |
| FWER p-Value | 0.027 |

| PROBE | GENE SYMBOL | GENE_TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|---|
| 1 | PUS1 | 15 | 26.459 | 0.0359 | Yes | ||
| 2 | RCL1 | 46 | 19.200 | 0.0609 | Yes | ||
| 3 | RANBP1 | 55 | 18.855 | 0.0867 | Yes | ||
| 4 | ACY1 | 57 | 18.355 | 0.1121 | Yes | ||
| 5 | TIMM8A | 60 | 18.225 | 0.1373 | Yes | ||
| 6 | FKBP11 | 97 | 15.642 | 0.1570 | Yes | ||
| 7 | PPAT | 110 | 15.096 | 0.1773 | Yes | ||
| 8 | CAD | 139 | 14.296 | 0.1957 | Yes | ||
| 9 | ATAD3A | 148 | 14.035 | 0.2147 | Yes | ||
| 10 | G6PC3 | 166 | 13.392 | 0.2324 | Yes | ||
| 11 | NUDC | 206 | 12.580 | 0.2477 | Yes | ||
| 12 | SUCLG2 | 253 | 11.870 | 0.2617 | Yes | ||
| 13 | OPRS1 | 289 | 11.463 | 0.2757 | Yes | ||
| 14 | QTRT1 | 342 | 10.813 | 0.2879 | Yes | ||
| 15 | LYAR | 351 | 10.704 | 0.3023 | Yes | ||
| 16 | IFRD2 | 449 | 9.828 | 0.3107 | Yes | ||
| 17 | TIMM10 | 474 | 9.631 | 0.3228 | Yes | ||
| 18 | PPRC1 | 502 | 9.457 | 0.3345 | Yes | ||
| 19 | MTHFD1 | 589 | 8.792 | 0.3420 | Yes | ||
| 20 | EIF4E | 663 | 8.322 | 0.3496 | Yes | ||
| 21 | PA2G4 | 666 | 8.313 | 0.3610 | Yes | ||
| 22 | SYNCRIP | 679 | 8.203 | 0.3718 | Yes | ||
| 23 | UBE1 | 718 | 7.942 | 0.3807 | Yes | ||
| 24 | PAICS | 764 | 7.678 | 0.3889 | Yes | ||
| 25 | AMPD2 | 789 | 7.536 | 0.3981 | Yes | ||
| 26 | NCL | 817 | 7.411 | 0.4069 | Yes | ||
| 27 | MRPL40 | 848 | 7.236 | 0.4153 | Yes | ||
| 28 | IVNS1ABP | 866 | 7.144 | 0.4243 | Yes | ||
| 29 | QTRTD1 | 939 | 6.783 | 0.4298 | Yes | ||
| 30 | CLN3 | 1052 | 6.281 | 0.4325 | Yes | ||
| 31 | GPS1 | 1136 | 5.929 | 0.4362 | Yes | ||
| 32 | PRDX4 | 1180 | 5.811 | 0.4420 | Yes | ||
| 33 | NOL5A | 1208 | 5.699 | 0.4484 | Yes | ||
| 34 | EEF1B2 | 1259 | 5.530 | 0.4534 | Yes | ||
| 35 | PPARGC1B | 1288 | 5.449 | 0.4594 | Yes | ||
| 36 | PSME3 | 1362 | 5.209 | 0.4627 | Yes | ||
| 37 | PRPS1 | 1398 | 5.067 | 0.4678 | Yes | ||
| 38 | MPP3 | 1438 | 4.947 | 0.4726 | Yes | ||
| 39 | RAMP2 | 1485 | 4.790 | 0.4767 | Yes | ||
| 40 | RPS19 | 1514 | 4.699 | 0.4817 | Yes | ||
| 41 | DIABLO | 1562 | 4.601 | 0.4856 | Yes | ||
| 42 | PABPC1 | 1592 | 4.530 | 0.4903 | Yes | ||
| 43 | ENO3 | 1689 | 4.322 | 0.4911 | Yes | ||
| 44 | ARL3 | 1694 | 4.316 | 0.4969 | Yes | ||
| 45 | SFXN2 | 1872 | 3.898 | 0.4927 | Yes | ||
| 46 | HNRPA1 | 1915 | 3.826 | 0.4957 | Yes | ||
| 47 | MAT2A | 1931 | 3.792 | 0.5002 | Yes | ||
| 48 | HIRA | 1938 | 3.779 | 0.5051 | Yes | ||
| 49 | HNRPDL | 2005 | 3.662 | 0.5066 | Yes | ||
| 50 | RHEBL1 | 2100 | 3.496 | 0.5063 | Yes | ||
| 51 | GPRC5C | 2145 | 3.415 | 0.5087 | Yes | ||
| 52 | PTGES2 | 2149 | 3.406 | 0.5133 | Yes | ||
| 53 | SYT3 | 2181 | 3.355 | 0.5162 | Yes | ||
| 54 | KCNN4 | 2186 | 3.349 | 0.5207 | Yes | ||
| 55 | HMGA1 | 2217 | 3.288 | 0.5236 | Yes | ||
| 56 | DIRC2 | 2480 | 2.924 | 0.5135 | No | ||
| 57 | HTF9C | 2494 | 2.917 | 0.5168 | No | ||
| 58 | TGFB2 | 2556 | 2.830 | 0.5174 | No | ||
| 59 | ATOH8 | 2862 | 2.510 | 0.5044 | No | ||
| 60 | BCL7C | 2929 | 2.453 | 0.5042 | No | ||
| 61 | TFRC | 2957 | 2.430 | 0.5061 | No | ||
| 62 | ILF3 | 2971 | 2.413 | 0.5088 | No | ||
| 63 | SRP72 | 3020 | 2.369 | 0.5094 | No | ||
| 64 | TRIB1 | 3107 | 2.284 | 0.5080 | No | ||
| 65 | GPM6B | 3148 | 2.255 | 0.5089 | No | ||
| 66 | TIMM9 | 3377 | 2.087 | 0.4995 | No | ||
| 67 | NPM1 | 3426 | 2.056 | 0.4997 | No | ||
| 68 | CRMP1 | 3504 | 2.007 | 0.4983 | No | ||
| 69 | NR6A1 | 3687 | 1.892 | 0.4911 | No | ||
| 70 | RTN4RL2 | 3694 | 1.890 | 0.4934 | No | ||
| 71 | U2AF2 | 3699 | 1.888 | 0.4958 | No | ||
| 72 | STMN1 | 3958 | 1.757 | 0.4842 | No | ||
| 73 | RAB3IL1 | 4004 | 1.738 | 0.4842 | No | ||
| 74 | FLT3 | 4175 | 1.654 | 0.4773 | No | ||
| 75 | LZTS2 | 4205 | 1.638 | 0.4780 | No | ||
| 76 | POGK | 4576 | 1.479 | 0.4600 | No | ||
| 77 | LRP8 | 4596 | 1.469 | 0.4610 | No | ||
| 78 | BOK | 4741 | 1.414 | 0.4551 | No | ||
| 79 | RFX5 | 4864 | 1.370 | 0.4504 | No | ||
| 80 | KCTD15 | 4913 | 1.350 | 0.4497 | No | ||
| 81 | TCOF1 | 5108 | 1.286 | 0.4410 | No | ||
| 82 | POU3F2 | 5133 | 1.276 | 0.4414 | No | ||
| 83 | KRTCAP2 | 5160 | 1.268 | 0.4418 | No | ||
| 84 | ADAMTS19 | 5215 | 1.251 | 0.4406 | No | ||
| 85 | MEOX2 | 5306 | 1.219 | 0.4374 | No | ||
| 86 | CHD4 | 5615 | 1.122 | 0.4223 | No | ||
| 87 | KIAA0427 | 5680 | 1.098 | 0.4203 | No | ||
| 88 | NDUFS1 | 5802 | 1.062 | 0.4152 | No | ||
| 89 | OPRD1 | 5854 | 1.047 | 0.4139 | No | ||
| 90 | IPO13 | 5879 | 1.039 | 0.4141 | No | ||
| 91 | ELAVL3 | 5893 | 1.035 | 0.4148 | No | ||
| 92 | ADAMTS17 | 5901 | 1.033 | 0.4159 | No | ||
| 93 | HSPBAP1 | 5952 | 1.021 | 0.4146 | No | ||
| 94 | RUNX2 | 6201 | 0.942 | 0.4024 | No | ||
| 95 | DAZL | 6639 | 0.814 | 0.3799 | No | ||
| 96 | SNCAIP | 6672 | 0.806 | 0.3793 | No | ||
| 97 | RAI14 | 6699 | 0.798 | 0.3790 | No | ||
| 98 | TCERG1 | 6793 | 0.776 | 0.3750 | No | ||
| 99 | ODC1 | 7047 | 0.711 | 0.3623 | No | ||
| 100 | DAZAP1 | 7216 | 0.669 | 0.3541 | No | ||
| 101 | HOXB5 | 7359 | 0.628 | 0.3473 | No | ||
| 102 | SLC12A5 | 7839 | 0.499 | 0.3220 | No | ||
| 103 | PCDHA10 | 7868 | 0.492 | 0.3211 | No | ||
| 104 | RFX4 | 8158 | 0.419 | 0.3061 | No | ||
| 105 | GPC3 | 8430 | 0.353 | 0.2919 | No | ||
| 106 | RORC | 8567 | 0.320 | 0.2849 | No | ||
| 107 | CEBPA | 8656 | 0.301 | 0.2806 | No | ||
| 108 | PITX3 | 8662 | 0.300 | 0.2807 | No | ||
| 109 | TESK2 | 9115 | 0.191 | 0.2565 | No | ||
| 110 | NRAS | 9203 | 0.170 | 0.2520 | No | ||
| 111 | SMAD2 | 9357 | 0.129 | 0.2439 | No | ||
| 112 | MNT | 9650 | 0.062 | 0.2282 | No | ||
| 113 | CHRM1 | 9946 | -0.018 | 0.2122 | No | ||
| 114 | NEUROD1 | 9994 | -0.031 | 0.2097 | No | ||
| 115 | APOA5 | 10575 | -0.174 | 0.1785 | No | ||
| 116 | PTMA | 10754 | -0.223 | 0.1691 | No | ||
| 117 | COL2A1 | 11018 | -0.293 | 0.1553 | No | ||
| 118 | CNNM1 | 11142 | -0.329 | 0.1491 | No | ||
| 119 | IPO7 | 11203 | -0.344 | 0.1463 | No | ||
| 120 | ZBTB10 | 11380 | -0.391 | 0.1373 | No | ||
| 121 | CGREF1 | 11386 | -0.394 | 0.1376 | No | ||
| 122 | LIN28 | 11478 | -0.419 | 0.1332 | No | ||
| 123 | HOXB7 | 11617 | -0.461 | 0.1264 | No | ||
| 124 | PER1 | 11715 | -0.484 | 0.1218 | No | ||
| 125 | HHIP | 11900 | -0.536 | 0.1126 | No | ||
| 126 | HOXA11 | 11964 | -0.549 | 0.1099 | No | ||
| 127 | FOXD3 | 12009 | -0.560 | 0.1083 | No | ||
| 128 | UBE2B | 12066 | -0.577 | 0.1061 | No | ||
| 129 | ABCA1 | 12124 | -0.596 | 0.1038 | No | ||
| 130 | FGF14 | 12487 | -0.707 | 0.0852 | No | ||
| 131 | SFRS14 | 12513 | -0.715 | 0.0848 | No | ||
| 132 | SATB2 | 12568 | -0.734 | 0.0829 | No | ||
| 133 | HOXA3 | 12604 | -0.746 | 0.0820 | No | ||
| 134 | PFN1 | 12820 | -0.816 | 0.0715 | No | ||
| 135 | SNX5 | 12856 | -0.826 | 0.0708 | No | ||
| 136 | UBXD3 | 13053 | -0.896 | 0.0614 | No | ||
| 137 | ARMET | 13057 | -0.897 | 0.0625 | No | ||
| 138 | NRIP3 | 13089 | -0.907 | 0.0620 | No | ||
| 139 | LAP3 | 13317 | -0.988 | 0.0511 | No | ||
| 140 | CBX5 | 13421 | -1.026 | 0.0470 | No | ||
| 141 | SLC1A7 | 13492 | -1.064 | 0.0446 | No | ||
| 142 | TOP1 | 14010 | -1.275 | 0.0184 | No | ||
| 143 | FKBP10 | 14494 | -1.526 | -0.0057 | No | ||
| 144 | SLC6A15 | 15146 | -1.910 | -0.0383 | No | ||
| 145 | KCMF1 | 15163 | -1.924 | -0.0365 | No | ||
| 146 | EFNB1 | 15500 | -2.206 | -0.0517 | No | ||
| 147 | ZADH2 | 15669 | -2.373 | -0.0575 | No | ||
| 148 | TRIM46 | 15781 | -2.482 | -0.0601 | No | ||
| 149 | WDR23 | 16224 | -3.048 | -0.0798 | No | ||
| 150 | USP15 | 16587 | -3.735 | -0.0942 | No | ||
| 151 | GADD45G | 16689 | -3.925 | -0.0943 | No | ||
| 152 | ZHX2 | 16890 | -4.381 | -0.0990 | No | ||
| 153 | CDC14A | 16908 | -4.437 | -0.0938 | No | ||
| 154 | EIF2C2 | 16911 | -4.445 | -0.0877 | No | ||
| 155 | MXD4 | 16968 | -4.544 | -0.0845 | No | ||
| 156 | DNAJB5 | 17042 | -4.686 | -0.0819 | No | ||
| 157 | H3F3A | 17131 | -4.977 | -0.0798 | No | ||
| 158 | BCAS3 | 17364 | -5.682 | -0.0845 | No | ||
| 159 | ETV4 | 17497 | -6.025 | -0.0832 | No | ||
| 160 | SDC1 | 17551 | -6.254 | -0.0774 | No | ||
| 161 | RPS6KA5 | 17680 | -6.679 | -0.0751 | No | ||
| 162 | STXBP2 | 17749 | -6.908 | -0.0692 | No | ||
| 163 | ESRRA | 17890 | -7.595 | -0.0663 | No | ||
| 164 | PPP1R9B | 17968 | -8.059 | -0.0592 | No | ||
| 165 | ETV1 | 18059 | -8.622 | -0.0522 | No | ||
| 166 | ATF7IP | 18257 | -10.399 | -0.0484 | No | ||
| 167 | BMP4 | 18339 | -11.411 | -0.0369 | No | ||
| 168 | CSK | 18344 | -11.494 | -0.0212 | No | ||
| 169 | GJA1 | 18398 | -12.744 | -0.0064 | No | ||
| 170 | ISGF3G | 18414 | -13.083 | 0.0110 | No |