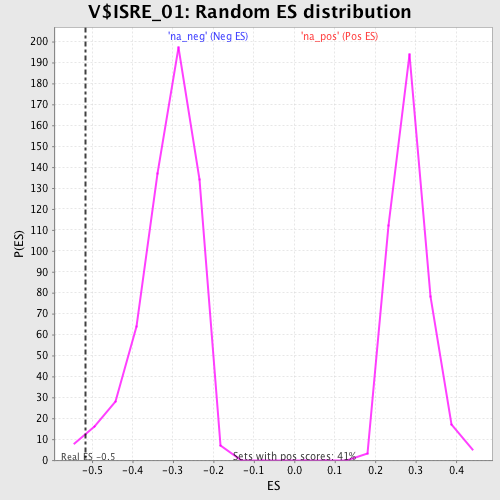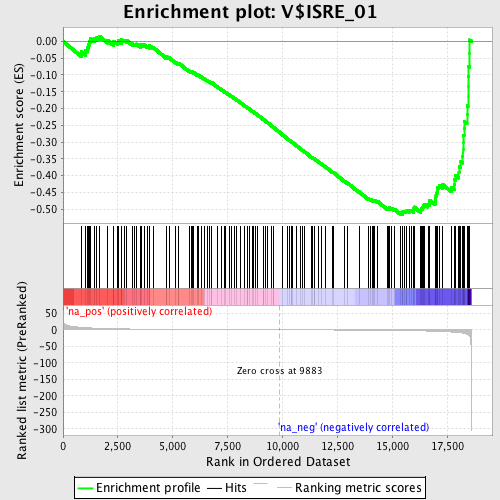
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | set04_DMpreB_versus_WTpreB |
| Phenotype | NoPhenotypeAvailable |
| Upregulated in class | na_neg |
| GeneSet | V$ISRE_01 |
| Enrichment Score (ES) | -0.5155496 |
| Normalized Enrichment Score (NES) | -1.6488416 |
| Nominal p-value | 0.015228426 |
| FDR q-value | 1.0 |
| FWER p-Value | 1.0 |

| PROBE | GENE SYMBOL | GENE_TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|---|
| 1 | ARHGEF6 | 838 | 7.291 | -0.0304 | No | ||
| 2 | BST2 | 1031 | 6.349 | -0.0278 | No | ||
| 3 | KCNN3 | 1102 | 6.045 | -0.0191 | No | ||
| 4 | SERHL | 1147 | 5.907 | -0.0094 | No | ||
| 5 | PGK1 | 1183 | 5.808 | 0.0007 | No | ||
| 6 | ANAPC1 | 1263 | 5.518 | 0.0077 | No | ||
| 7 | LY86 | 1418 | 4.993 | 0.0097 | No | ||
| 8 | HEPH | 1540 | 4.636 | 0.0126 | No | ||
| 9 | PIK3R3 | 1649 | 4.388 | 0.0158 | No | ||
| 10 | CRK | 2032 | 3.614 | 0.0025 | No | ||
| 11 | MOV10 | 2298 | 3.165 | -0.0053 | No | ||
| 12 | ADD3 | 2306 | 3.150 | 0.0008 | No | ||
| 13 | GPSN2 | 2478 | 2.926 | -0.0025 | No | ||
| 14 | ANGPTL4 | 2501 | 2.910 | 0.0023 | No | ||
| 15 | E2F3 | 2656 | 2.722 | -0.0004 | No | ||
| 16 | PSMA5 | 2658 | 2.718 | 0.0051 | No | ||
| 17 | TNFRSF17 | 2776 | 2.581 | 0.0041 | No | ||
| 18 | ESR1 | 2903 | 2.472 | 0.0023 | No | ||
| 19 | PRKACA | 3166 | 2.234 | -0.0073 | No | ||
| 20 | ADAM15 | 3258 | 2.177 | -0.0077 | No | ||
| 21 | TNFSF13B | 3338 | 2.118 | -0.0076 | No | ||
| 22 | KPNA3 | 3532 | 1.989 | -0.0140 | No | ||
| 23 | GPR65 | 3539 | 1.985 | -0.0103 | No | ||
| 24 | WNT8B | 3571 | 1.964 | -0.0079 | No | ||
| 25 | AMMECR1 | 3688 | 1.891 | -0.0103 | No | ||
| 26 | ERBB2 | 3838 | 1.816 | -0.0146 | No | ||
| 27 | PSMA3 | 3935 | 1.768 | -0.0162 | No | ||
| 28 | TNRC4 | 3938 | 1.767 | -0.0127 | No | ||
| 29 | FMR1 | 4115 | 1.684 | -0.0187 | No | ||
| 30 | CACNA2D2 | 4690 | 1.433 | -0.0469 | No | ||
| 31 | BDNF | 4697 | 1.430 | -0.0443 | No | ||
| 32 | KCNJ16 | 4829 | 1.384 | -0.0485 | No | ||
| 33 | KCNE4 | 5126 | 1.279 | -0.0619 | No | ||
| 34 | NT5C3 | 5262 | 1.236 | -0.0667 | No | ||
| 35 | IL4 | 5273 | 1.232 | -0.0647 | No | ||
| 36 | IL27 | 5746 | 1.079 | -0.0880 | No | ||
| 37 | PCDH7 | 5843 | 1.048 | -0.0911 | No | ||
| 38 | COL12A1 | 5876 | 1.041 | -0.0907 | No | ||
| 39 | CDK6 | 5959 | 1.019 | -0.0930 | No | ||
| 40 | THBS1 | 6139 | 0.962 | -0.1008 | No | ||
| 41 | PROK2 | 6161 | 0.956 | -0.0999 | No | ||
| 42 | CBX4 | 6299 | 0.912 | -0.1055 | No | ||
| 43 | OTX1 | 6446 | 0.869 | -0.1116 | No | ||
| 44 | GATA6 | 6593 | 0.827 | -0.1178 | No | ||
| 45 | CRY1 | 6670 | 0.807 | -0.1203 | No | ||
| 46 | UNC5C | 6746 | 0.788 | -0.1227 | No | ||
| 47 | PAX2 | 6782 | 0.778 | -0.1230 | No | ||
| 48 | LGP2 | 7028 | 0.717 | -0.1348 | No | ||
| 49 | IFIT2 | 7225 | 0.666 | -0.1440 | No | ||
| 50 | RANGAP1 | 7372 | 0.623 | -0.1507 | No | ||
| 51 | MMP25 | 7423 | 0.610 | -0.1521 | No | ||
| 52 | CHKA | 7564 | 0.573 | -0.1585 | No | ||
| 53 | IFNB1 | 7668 | 0.547 | -0.1630 | No | ||
| 54 | FIGN | 7813 | 0.509 | -0.1697 | No | ||
| 55 | TCF15 | 7920 | 0.481 | -0.1745 | No | ||
| 56 | CXXC5 | 8063 | 0.443 | -0.1813 | No | ||
| 57 | MET | 8268 | 0.394 | -0.1915 | No | ||
| 58 | LMO1 | 8392 | 0.364 | -0.1974 | No | ||
| 59 | LRP2 | 8421 | 0.356 | -0.1982 | No | ||
| 60 | AIF1 | 8502 | 0.336 | -0.2018 | No | ||
| 61 | NKX2-2 | 8634 | 0.305 | -0.2083 | No | ||
| 62 | PITX3 | 8662 | 0.300 | -0.2092 | No | ||
| 63 | MSX1 | 8774 | 0.270 | -0.2146 | No | ||
| 64 | SEMA6D | 8844 | 0.254 | -0.2178 | No | ||
| 65 | ERG | 9143 | 0.185 | -0.2336 | No | ||
| 66 | TLR7 | 9146 | 0.184 | -0.2333 | No | ||
| 67 | EPN1 | 9206 | 0.169 | -0.2362 | No | ||
| 68 | CXCL10 | 9329 | 0.138 | -0.2425 | No | ||
| 69 | ZCCHC5 | 9490 | 0.094 | -0.2510 | No | ||
| 70 | NPR3 | 9570 | 0.076 | -0.2551 | No | ||
| 71 | MIA2 | 9603 | 0.069 | -0.2567 | No | ||
| 72 | ETV6 | 10015 | -0.037 | -0.2788 | No | ||
| 73 | CXCL11 | 10204 | -0.079 | -0.2889 | No | ||
| 74 | PURA | 10326 | -0.109 | -0.2952 | No | ||
| 75 | SLC12A7 | 10389 | -0.126 | -0.2983 | No | ||
| 76 | OSBP | 10417 | -0.135 | -0.2995 | No | ||
| 77 | TNFSF15 | 10433 | -0.138 | -0.3000 | No | ||
| 78 | EYA4 | 10632 | -0.189 | -0.3103 | No | ||
| 79 | FYCO1 | 10825 | -0.242 | -0.3202 | No | ||
| 80 | PLXNC1 | 10905 | -0.262 | -0.3240 | No | ||
| 81 | ZBP1 | 10997 | -0.287 | -0.3283 | No | ||
| 82 | HTR2C | 11329 | -0.379 | -0.3455 | No | ||
| 83 | SLC24A1 | 11375 | -0.389 | -0.3471 | No | ||
| 84 | OVOL1 | 11446 | -0.409 | -0.3501 | No | ||
| 85 | TBX21 | 11456 | -0.414 | -0.3497 | No | ||
| 86 | SDPR | 11648 | -0.469 | -0.3591 | No | ||
| 87 | EGFL6 | 11797 | -0.506 | -0.3660 | No | ||
| 88 | IL22RA1 | 11940 | -0.543 | -0.3726 | No | ||
| 89 | EREG | 12297 | -0.649 | -0.3906 | No | ||
| 90 | RRBP1 | 12302 | -0.649 | -0.3895 | No | ||
| 91 | CDH3 | 12826 | -0.819 | -0.4161 | No | ||
| 92 | PIGR | 12961 | -0.858 | -0.4216 | No | ||
| 93 | PRDM16 | 13485 | -1.061 | -0.4477 | No | ||
| 94 | PLAG1 | 13919 | -1.229 | -0.4687 | No | ||
| 95 | TOP1 | 14010 | -1.275 | -0.4709 | No | ||
| 96 | JARID2 | 14105 | -1.321 | -0.4733 | No | ||
| 97 | SIX1 | 14168 | -1.354 | -0.4739 | No | ||
| 98 | ZFPM1 | 14203 | -1.367 | -0.4729 | No | ||
| 99 | FGF9 | 14329 | -1.436 | -0.4767 | No | ||
| 100 | PPARGC1A | 14798 | -1.685 | -0.4986 | No | ||
| 101 | ZNF385 | 14845 | -1.714 | -0.4976 | No | ||
| 102 | SLC25A28 | 14858 | -1.720 | -0.4947 | No | ||
| 103 | HPCAL1 | 14980 | -1.801 | -0.4975 | No | ||
| 104 | ASPA | 15085 | -1.872 | -0.4993 | No | ||
| 105 | HOMER1 | 15379 | -2.090 | -0.5109 | No | ||
| 106 | USF1 | 15466 | -2.173 | -0.5111 | Yes | ||
| 107 | B2M | 15485 | -2.188 | -0.5076 | Yes | ||
| 108 | YWHAG | 15545 | -2.257 | -0.5061 | Yes | ||
| 109 | RIPK2 | 15631 | -2.337 | -0.5059 | Yes | ||
| 110 | DGKA | 15671 | -2.375 | -0.5032 | Yes | ||
| 111 | SLC11A2 | 15773 | -2.475 | -0.5035 | Yes | ||
| 112 | OPTN | 15858 | -2.566 | -0.5028 | Yes | ||
| 113 | USP18 | 15974 | -2.713 | -0.5035 | Yes | ||
| 114 | TCIRG1 | 15977 | -2.718 | -0.4980 | Yes | ||
| 115 | BBX | 15994 | -2.735 | -0.4932 | Yes | ||
| 116 | KYNU | 16284 | -3.161 | -0.5024 | Yes | ||
| 117 | GPX1 | 16313 | -3.203 | -0.4973 | Yes | ||
| 118 | SH3BGRL | 16366 | -3.277 | -0.4934 | Yes | ||
| 119 | TAPBP | 16417 | -3.381 | -0.4892 | Yes | ||
| 120 | GNB4 | 16483 | -3.509 | -0.4855 | Yes | ||
| 121 | MYB | 16630 | -3.818 | -0.4855 | Yes | ||
| 122 | TCF7L2 | 16696 | -3.946 | -0.4810 | Yes | ||
| 123 | GPR18 | 16709 | -3.973 | -0.4734 | Yes | ||
| 124 | MEIS1 | 16952 | -4.525 | -0.4772 | Yes | ||
| 125 | GSDMDC1 | 16977 | -4.558 | -0.4692 | Yes | ||
| 126 | MAP2K6 | 16991 | -4.589 | -0.4604 | Yes | ||
| 127 | PPP2R5C | 17001 | -4.612 | -0.4515 | Yes | ||
| 128 | HIST1H1C | 17045 | -4.690 | -0.4441 | Yes | ||
| 129 | BLNK | 17078 | -4.805 | -0.4360 | Yes | ||
| 130 | LSP1 | 17132 | -4.978 | -0.4286 | Yes | ||
| 131 | STAT6 | 17308 | -5.487 | -0.4269 | Yes | ||
| 132 | BCL11A | 17700 | -6.753 | -0.4341 | Yes | ||
| 133 | UBE1L | 17831 | -7.260 | -0.4263 | Yes | ||
| 134 | IRF2 | 17840 | -7.315 | -0.4117 | Yes | ||
| 135 | SSBP2 | 17874 | -7.539 | -0.3980 | Yes | ||
| 136 | THRAP1 | 18042 | -8.516 | -0.3895 | Yes | ||
| 137 | CDKN2C | 18047 | -8.548 | -0.3722 | Yes | ||
| 138 | CXCR4 | 18129 | -9.221 | -0.3576 | Yes | ||
| 139 | MAP3K11 | 18217 | -10.093 | -0.3416 | Yes | ||
| 140 | DDX58 | 18225 | -10.159 | -0.3211 | Yes | ||
| 141 | DAPP1 | 18229 | -10.221 | -0.3002 | Yes | ||
| 142 | PML | 18233 | -10.280 | -0.2793 | Yes | ||
| 143 | EHD4 | 18277 | -10.570 | -0.2599 | Yes | ||
| 144 | CD151 | 18288 | -10.663 | -0.2385 | Yes | ||
| 145 | MS4A1 | 18409 | -12.978 | -0.2183 | Yes | ||
| 146 | ISGF3G | 18414 | -13.083 | -0.1917 | Yes | ||
| 147 | MEF2C | 18457 | -14.583 | -0.1640 | Yes | ||
| 148 | EGR2 | 18458 | -14.584 | -0.1340 | Yes | ||
| 149 | SLA | 18460 | -14.650 | -0.1040 | Yes | ||
| 150 | JMJD2A | 18474 | -15.105 | -0.0736 | Yes | ||
| 151 | ETV5 | 18533 | -19.367 | -0.0370 | Yes | ||
| 152 | DUSP10 | 18536 | -20.147 | 0.0043 | Yes |