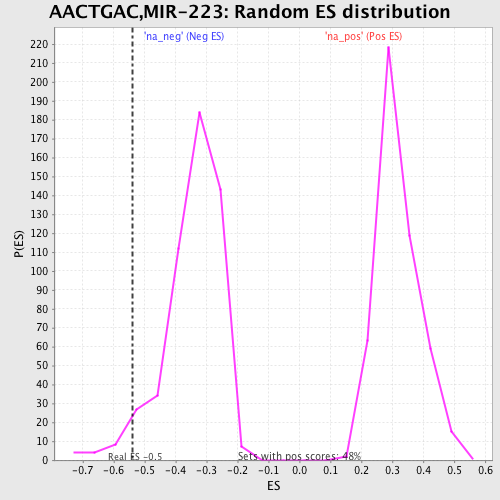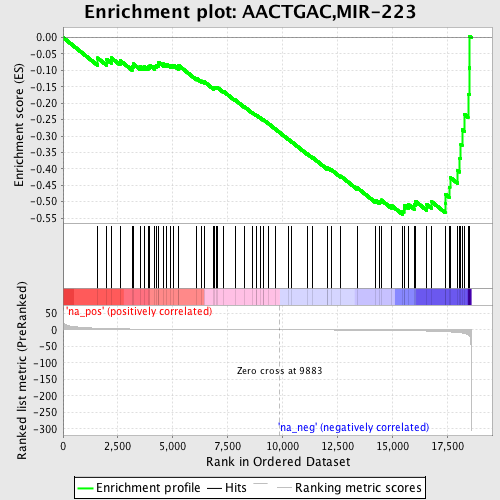
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | set04_DMpreB_versus_WTpreB |
| Phenotype | NoPhenotypeAvailable |
| Upregulated in class | na_neg |
| GeneSet | AACTGAC,MIR-223 |
| Enrichment Score (ES) | -0.5390813 |
| Normalized Enrichment Score (NES) | -1.5649824 |
| Nominal p-value | 0.045889102 |
| FDR q-value | 1.0 |
| FWER p-Value | 1.0 |

| PROBE | GENE SYMBOL | GENE_TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|---|
| 1 | SLC8A1 | 1566 | 4.588 | -0.0622 | No | ||
| 2 | APRIN | 1985 | 3.694 | -0.0668 | No | ||
| 3 | RASSF4 | 2196 | 3.333 | -0.0619 | No | ||
| 4 | RPS6KB1 | 2599 | 2.783 | -0.0701 | No | ||
| 5 | PLEKHH1 | 3144 | 2.259 | -0.0884 | No | ||
| 6 | SEPT6 | 3193 | 2.216 | -0.0802 | No | ||
| 7 | KPNA3 | 3532 | 1.989 | -0.0888 | No | ||
| 8 | QKI | 3701 | 1.887 | -0.0887 | No | ||
| 9 | RNF4 | 3873 | 1.796 | -0.0892 | No | ||
| 10 | STMN1 | 3958 | 1.757 | -0.0852 | No | ||
| 11 | FBXO8 | 4176 | 1.653 | -0.0889 | No | ||
| 12 | ANKRD17 | 4249 | 1.611 | -0.0849 | No | ||
| 13 | TSPAN7 | 4334 | 1.578 | -0.0818 | No | ||
| 14 | OLFM1 | 4357 | 1.563 | -0.0754 | No | ||
| 15 | IGF1R | 4582 | 1.477 | -0.0803 | No | ||
| 16 | NFIB | 4720 | 1.421 | -0.0808 | No | ||
| 17 | EFNA1 | 4915 | 1.349 | -0.0847 | No | ||
| 18 | SCN3A | 5052 | 1.301 | -0.0857 | No | ||
| 19 | F3 | 5268 | 1.233 | -0.0913 | No | ||
| 20 | STK39 | 5271 | 1.233 | -0.0854 | No | ||
| 21 | SLC26A7 | 6088 | 0.977 | -0.1247 | No | ||
| 22 | PTBP2 | 6313 | 0.909 | -0.1323 | No | ||
| 23 | IL28RA | 6436 | 0.871 | -0.1347 | No | ||
| 24 | NFIA | 6866 | 0.757 | -0.1541 | No | ||
| 25 | CRIM1 | 6904 | 0.750 | -0.1525 | No | ||
| 26 | PRDM1 | 6981 | 0.728 | -0.1531 | No | ||
| 27 | IL6ST | 7017 | 0.719 | -0.1514 | No | ||
| 28 | KIF4A | 7298 | 0.644 | -0.1634 | No | ||
| 29 | FBXW7 | 7836 | 0.500 | -0.1899 | No | ||
| 30 | ZCCHC14 | 8258 | 0.396 | -0.2107 | No | ||
| 31 | PCTK2 | 8642 | 0.303 | -0.2299 | No | ||
| 32 | WDR26 | 8812 | 0.260 | -0.2378 | No | ||
| 33 | RCN2 | 8819 | 0.259 | -0.2368 | No | ||
| 34 | ACVR2A | 8986 | 0.218 | -0.2447 | No | ||
| 35 | SPRED1 | 9122 | 0.189 | -0.2511 | No | ||
| 36 | SOX11 | 9133 | 0.186 | -0.2507 | No | ||
| 37 | INPP5B | 9356 | 0.130 | -0.2620 | No | ||
| 38 | UBTF | 9695 | 0.050 | -0.2800 | No | ||
| 39 | SLC23A2 | 10255 | -0.092 | -0.3097 | No | ||
| 40 | SP3 | 10431 | -0.137 | -0.3185 | No | ||
| 41 | MMP16 | 11121 | -0.325 | -0.3541 | No | ||
| 42 | ZBTB10 | 11380 | -0.391 | -0.3661 | No | ||
| 43 | CALML4 | 12037 | -0.568 | -0.3987 | No | ||
| 44 | ATP7A | 12058 | -0.574 | -0.3970 | No | ||
| 45 | ANKRD16 | 12220 | -0.628 | -0.4026 | No | ||
| 46 | VNN1 | 12645 | -0.763 | -0.4218 | No | ||
| 47 | PLAGL2 | 13402 | -1.019 | -0.4576 | No | ||
| 48 | SON | 14230 | -1.381 | -0.4955 | No | ||
| 49 | CSNK1G1 | 14418 | -1.481 | -0.4984 | No | ||
| 50 | SLC35F1 | 14491 | -1.525 | -0.4948 | No | ||
| 51 | RASA1 | 14954 | -1.781 | -0.5111 | No | ||
| 52 | ALCAM | 15474 | -2.178 | -0.5285 | Yes | ||
| 53 | RBM16 | 15537 | -2.248 | -0.5209 | Yes | ||
| 54 | SMARCD1 | 15565 | -2.267 | -0.5114 | Yes | ||
| 55 | PDE4D | 15754 | -2.452 | -0.5096 | Yes | ||
| 56 | RHOB | 16003 | -2.745 | -0.5096 | Yes | ||
| 57 | SLC39A1 | 16061 | -2.816 | -0.4990 | Yes | ||
| 58 | CBFB | 16574 | -3.709 | -0.5086 | Yes | ||
| 59 | RAB8B | 16801 | -4.206 | -0.5004 | Yes | ||
| 60 | MTPN | 17413 | -5.809 | -0.5051 | Yes | ||
| 61 | STIM1 | 17422 | -5.840 | -0.4772 | Yes | ||
| 62 | PURB | 17595 | -6.397 | -0.4554 | Yes | ||
| 63 | UBE2A | 17634 | -6.549 | -0.4256 | Yes | ||
| 64 | ATP1B1 | 17985 | -8.207 | -0.4046 | Yes | ||
| 65 | PKNOX1 | 18082 | -8.811 | -0.3670 | Yes | ||
| 66 | PDPK1 | 18128 | -9.213 | -0.3247 | Yes | ||
| 67 | RNF34 | 18186 | -9.788 | -0.2802 | Yes | ||
| 68 | HHEX | 18291 | -10.672 | -0.2340 | Yes | ||
| 69 | MEF2C | 18457 | -14.583 | -0.1721 | Yes | ||
| 70 | LMO2 | 18513 | -17.031 | -0.0923 | Yes | ||
| 71 | DUSP10 | 18536 | -20.147 | 0.0043 | Yes |