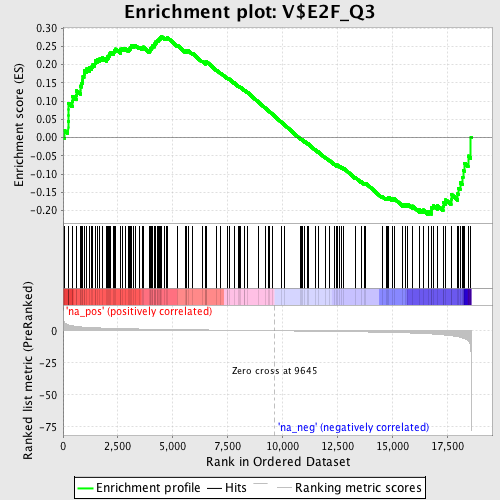
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | set03_wtNotch_versus_normalThy |
| Phenotype | NoPhenotypeAvailable |
| Upregulated in class | na_pos |
| GeneSet | V$E2F_Q3 |
| Enrichment Score (ES) | 0.27825278 |
| Normalized Enrichment Score (NES) | 1.2103553 |
| Nominal p-value | 0.07777778 |
| FDR q-value | 0.6104003 |
| FWER p-Value | 1.0 |

| PROBE | GENE SYMBOL | GENE_TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|---|
| 1 | HNRPA1 | 66 | 6.164 | 0.0192 | Yes | ||
| 2 | DNAJC11 | 222 | 4.614 | 0.0279 | Yes | ||
| 3 | IMPDH2 | 223 | 4.602 | 0.0449 | Yes | ||
| 4 | SLC25A3 | 234 | 4.571 | 0.0613 | Yes | ||
| 5 | TCP1 | 237 | 4.557 | 0.0781 | Yes | ||
| 6 | PCSK4 | 239 | 4.551 | 0.0948 | Yes | ||
| 7 | RBBP7 | 416 | 3.867 | 0.0996 | Yes | ||
| 8 | SHMT1 | 434 | 3.820 | 0.1128 | Yes | ||
| 9 | GCH1 | 598 | 3.411 | 0.1166 | Yes | ||
| 10 | ABCF2 | 615 | 3.369 | 0.1282 | Yes | ||
| 11 | EPHB2 | 786 | 3.039 | 0.1303 | Yes | ||
| 12 | SLC38A1 | 787 | 3.037 | 0.1415 | Yes | ||
| 13 | ING3 | 842 | 2.947 | 0.1495 | Yes | ||
| 14 | UNG | 881 | 2.882 | 0.1581 | Yes | ||
| 15 | NCL | 903 | 2.857 | 0.1675 | Yes | ||
| 16 | USP1 | 954 | 2.794 | 0.1751 | Yes | ||
| 17 | TFRC | 975 | 2.772 | 0.1843 | Yes | ||
| 18 | RFC1 | 1084 | 2.666 | 0.1883 | Yes | ||
| 19 | CACNA1G | 1202 | 2.554 | 0.1914 | Yes | ||
| 20 | PVRL1 | 1313 | 2.437 | 0.1945 | Yes | ||
| 21 | ERBB2IP | 1358 | 2.397 | 0.2010 | Yes | ||
| 22 | UXT | 1453 | 2.319 | 0.2045 | Yes | ||
| 23 | ATAD2 | 1479 | 2.298 | 0.2116 | Yes | ||
| 24 | RAD51 | 1567 | 2.241 | 0.2152 | Yes | ||
| 25 | MCM4 | 1678 | 2.171 | 0.2173 | Yes | ||
| 26 | PPP1R8 | 1786 | 2.110 | 0.2193 | Yes | ||
| 27 | SFRS2 | 1972 | 2.002 | 0.2167 | Yes | ||
| 28 | CNOT3 | 2018 | 1.975 | 0.2215 | Yes | ||
| 29 | HMGN2 | 2089 | 1.941 | 0.2249 | Yes | ||
| 30 | MCM6 | 2131 | 1.924 | 0.2298 | Yes | ||
| 31 | ATRX | 2174 | 1.898 | 0.2346 | Yes | ||
| 32 | LIG1 | 2301 | 1.838 | 0.2346 | Yes | ||
| 33 | HS6ST3 | 2327 | 1.826 | 0.2400 | Yes | ||
| 34 | PHF15 | 2394 | 1.798 | 0.2430 | Yes | ||
| 35 | PAPOLG | 2624 | 1.684 | 0.2369 | Yes | ||
| 36 | BAT3 | 2627 | 1.683 | 0.2430 | Yes | ||
| 37 | GRIK2 | 2709 | 1.644 | 0.2447 | Yes | ||
| 38 | AP4M1 | 2838 | 1.597 | 0.2437 | Yes | ||
| 39 | HTF9C | 2998 | 1.539 | 0.2407 | Yes | ||
| 40 | PRPS1 | 3042 | 1.526 | 0.2441 | Yes | ||
| 41 | KCNA1 | 3071 | 1.514 | 0.2481 | Yes | ||
| 42 | OLFML3 | 3101 | 1.505 | 0.2521 | Yes | ||
| 43 | STAG2 | 3198 | 1.469 | 0.2524 | Yes | ||
| 44 | APRIN | 3276 | 1.441 | 0.2535 | Yes | ||
| 45 | POLA2 | 3468 | 1.373 | 0.2483 | Yes | ||
| 46 | RHD | 3604 | 1.335 | 0.2459 | Yes | ||
| 47 | CPNE5 | 3646 | 1.317 | 0.2486 | Yes | ||
| 48 | PLK4 | 3925 | 1.235 | 0.2381 | Yes | ||
| 49 | MCM7 | 3963 | 1.224 | 0.2406 | Yes | ||
| 50 | KLF4 | 3967 | 1.222 | 0.2450 | Yes | ||
| 51 | NRP2 | 4032 | 1.205 | 0.2460 | Yes | ||
| 52 | MCM3 | 4053 | 1.199 | 0.2493 | Yes | ||
| 53 | CDC45L | 4089 | 1.188 | 0.2518 | Yes | ||
| 54 | CLTA | 4154 | 1.172 | 0.2527 | Yes | ||
| 55 | AP1S2 | 4157 | 1.171 | 0.2569 | Yes | ||
| 56 | RBBP4 | 4202 | 1.157 | 0.2588 | Yes | ||
| 57 | PIAS1 | 4218 | 1.153 | 0.2623 | Yes | ||
| 58 | RANBP1 | 4279 | 1.139 | 0.2632 | Yes | ||
| 59 | GMNN | 4284 | 1.138 | 0.2672 | Yes | ||
| 60 | LASP1 | 4348 | 1.125 | 0.2680 | Yes | ||
| 61 | KCND2 | 4374 | 1.118 | 0.2708 | Yes | ||
| 62 | ADAMTS2 | 4422 | 1.106 | 0.2723 | Yes | ||
| 63 | NASP | 4443 | 1.100 | 0.2753 | Yes | ||
| 64 | FBXO5 | 4464 | 1.095 | 0.2783 | Yes | ||
| 65 | E2F1 | 4642 | 1.052 | 0.2726 | No | ||
| 66 | SEMA5A | 4715 | 1.034 | 0.2725 | No | ||
| 67 | HIST1H2BK | 4741 | 1.027 | 0.2749 | No | ||
| 68 | EZH2 | 5215 | 0.913 | 0.2527 | No | ||
| 69 | ARHGAP6 | 5558 | 0.831 | 0.2373 | No | ||
| 70 | STK35 | 5620 | 0.816 | 0.2370 | No | ||
| 71 | E2F3 | 5630 | 0.814 | 0.2395 | No | ||
| 72 | POLE2 | 5700 | 0.799 | 0.2387 | No | ||
| 73 | SALL1 | 5900 | 0.755 | 0.2307 | No | ||
| 74 | SNPH | 6334 | 0.659 | 0.2097 | No | ||
| 75 | ARID4A | 6489 | 0.631 | 0.2037 | No | ||
| 76 | DLST | 6498 | 0.629 | 0.2056 | No | ||
| 77 | KPNB1 | 6534 | 0.622 | 0.2060 | No | ||
| 78 | CDC6 | 6535 | 0.622 | 0.2083 | No | ||
| 79 | HNRPD | 6999 | 0.530 | 0.1852 | No | ||
| 80 | STMN1 | 7185 | 0.491 | 0.1770 | No | ||
| 81 | PAQR4 | 7474 | 0.430 | 0.1630 | No | ||
| 82 | NDUFA11 | 7499 | 0.425 | 0.1633 | No | ||
| 83 | DNMT1 | 7574 | 0.412 | 0.1608 | No | ||
| 84 | MCM2 | 7792 | 0.369 | 0.1504 | No | ||
| 85 | TOPBP1 | 7998 | 0.326 | 0.1405 | No | ||
| 86 | HIST1H1D | 8038 | 0.318 | 0.1396 | No | ||
| 87 | CDC25A | 8094 | 0.305 | 0.1377 | No | ||
| 88 | NUP153 | 8272 | 0.267 | 0.1292 | No | ||
| 89 | PODN | 8277 | 0.265 | 0.1299 | No | ||
| 90 | NPR3 | 8396 | 0.247 | 0.1244 | No | ||
| 91 | CDCA7 | 8907 | 0.143 | 0.0974 | No | ||
| 92 | TREX2 | 9242 | 0.081 | 0.0796 | No | ||
| 93 | MRPL18 | 9378 | 0.052 | 0.0725 | No | ||
| 94 | ONECUT1 | 9386 | 0.051 | 0.0723 | No | ||
| 95 | UFD1L | 9544 | 0.017 | 0.0638 | No | ||
| 96 | TBX3 | 9958 | -0.062 | 0.0417 | No | ||
| 97 | BARHL1 | 10097 | -0.090 | 0.0346 | No | ||
| 98 | H2AFZ | 10801 | -0.236 | -0.0026 | No | ||
| 99 | EPHB1 | 10857 | -0.246 | -0.0047 | No | ||
| 100 | IPO7 | 10891 | -0.255 | -0.0055 | No | ||
| 101 | SEZ6 | 11023 | -0.282 | -0.0116 | No | ||
| 102 | PCSK1 | 11120 | -0.303 | -0.0157 | No | ||
| 103 | SFRS1 | 11161 | -0.311 | -0.0167 | No | ||
| 104 | SMAD6 | 11501 | -0.385 | -0.0336 | No | ||
| 105 | PRKDC | 11625 | -0.410 | -0.0387 | No | ||
| 106 | WEE1 | 11945 | -0.478 | -0.0542 | No | ||
| 107 | CASP8AP2 | 12131 | -0.519 | -0.0623 | No | ||
| 108 | FMO4 | 12366 | -0.570 | -0.0729 | No | ||
| 109 | CTCF | 12473 | -0.599 | -0.0764 | No | ||
| 110 | ORC1L | 12488 | -0.604 | -0.0750 | No | ||
| 111 | ASCL1 | 12600 | -0.629 | -0.0786 | No | ||
| 112 | NOL4 | 12698 | -0.652 | -0.0815 | No | ||
| 113 | ADCY8 | 12797 | -0.673 | -0.0843 | No | ||
| 114 | TRIM47 | 13312 | -0.803 | -0.1091 | No | ||
| 115 | PEG3 | 13585 | -0.869 | -0.1207 | No | ||
| 116 | MAT2A | 13726 | -0.905 | -0.1249 | No | ||
| 117 | JPH1 | 13790 | -0.924 | -0.1249 | No | ||
| 118 | FLI1 | 14542 | -1.147 | -0.1613 | No | ||
| 119 | DMD | 14721 | -1.204 | -0.1665 | No | ||
| 120 | SSBP3 | 14795 | -1.235 | -0.1659 | No | ||
| 121 | LHX5 | 14832 | -1.249 | -0.1632 | No | ||
| 122 | SLC6A4 | 15001 | -1.317 | -0.1674 | No | ||
| 123 | PCNA | 15099 | -1.356 | -0.1677 | No | ||
| 124 | HIST1H2AH | 15486 | -1.548 | -0.1828 | No | ||
| 125 | CTDSP1 | 15610 | -1.614 | -0.1835 | No | ||
| 126 | MDGA1 | 15716 | -1.672 | -0.1830 | No | ||
| 127 | MCM8 | 15922 | -1.787 | -0.1875 | No | ||
| 128 | MEIS1 | 16251 | -2.021 | -0.1978 | No | ||
| 129 | MXD3 | 16408 | -2.146 | -0.1983 | No | ||
| 130 | SLC25A14 | 16641 | -2.336 | -0.2022 | No | ||
| 131 | MAZ | 16783 | -2.485 | -0.2007 | No | ||
| 132 | PPP1CC | 16790 | -2.489 | -0.1918 | No | ||
| 133 | PHF13 | 16858 | -2.565 | -0.1859 | No | ||
| 134 | CBX5 | 17049 | -2.784 | -0.1859 | No | ||
| 135 | ZCCHC8 | 17318 | -3.172 | -0.1887 | No | ||
| 136 | INSM1 | 17344 | -3.223 | -0.1781 | No | ||
| 137 | ATE1 | 17443 | -3.402 | -0.1708 | No | ||
| 138 | DDB2 | 17682 | -3.864 | -0.1694 | No | ||
| 139 | KLF5 | 17688 | -3.882 | -0.1553 | No | ||
| 140 | PHF12 | 17991 | -4.670 | -0.1544 | No | ||
| 141 | DPYSL2 | 18025 | -4.769 | -0.1385 | No | ||
| 142 | PIM1 | 18097 | -5.045 | -0.1237 | No | ||
| 143 | E2F7 | 18197 | -5.528 | -0.1086 | No | ||
| 144 | SYT11 | 18229 | -5.673 | -0.0893 | No | ||
| 145 | RAB11B | 18285 | -6.081 | -0.0698 | No | ||
| 146 | DCK | 18459 | -7.779 | -0.0504 | No | ||
| 147 | TBX6 | 18589 | -15.903 | 0.0015 | No |
