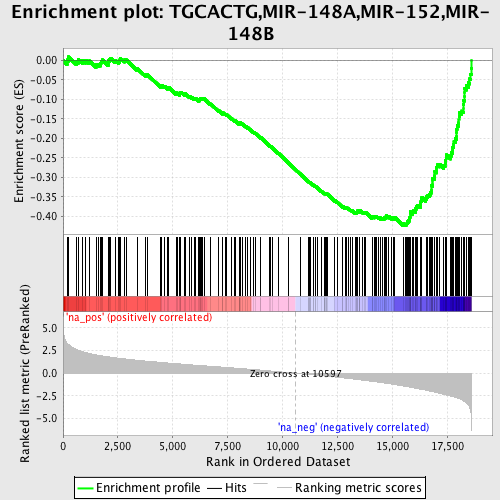
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | set03_truncNotch_versus_wtNotch |
| Phenotype | NoPhenotypeAvailable |
| Upregulated in class | na_neg |
| GeneSet | TGCACTG,MIR-148A,MIR-152,MIR-148B |
| Enrichment Score (ES) | -0.42448708 |
| Normalized Enrichment Score (NES) | -2.1756628 |
| Nominal p-value | 0.0 |
| FDR q-value | 0.0 |
| FWER p-Value | 0.0 |

| PROBE | GENE SYMBOL | GENE_TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|---|
| 1 | ETV5 | 182 | 3.305 | 0.0016 | No | ||
| 2 | HRB | 243 | 3.167 | 0.0094 | No | ||
| 3 | MNT | 624 | 2.589 | -0.0023 | No | ||
| 4 | ELMO1 | 706 | 2.505 | 0.0021 | No | ||
| 5 | SNPH | 879 | 2.371 | 0.0010 | No | ||
| 6 | ADCY2 | 1021 | 2.269 | 0.0012 | No | ||
| 7 | ALS2CR2 | 1201 | 2.166 | -0.0010 | No | ||
| 8 | KLHL18 | 1501 | 2.026 | -0.0102 | No | ||
| 9 | ITGA11 | 1607 | 1.980 | -0.0090 | No | ||
| 10 | MMP19 | 1703 | 1.943 | -0.0074 | No | ||
| 11 | FMR1 | 1729 | 1.931 | -0.0020 | No | ||
| 12 | PRKAG2 | 1785 | 1.912 | 0.0016 | No | ||
| 13 | POU3F2 | 2054 | 1.806 | -0.0066 | No | ||
| 14 | SLC2A1 | 2059 | 1.804 | -0.0006 | No | ||
| 15 | YPEL3 | 2108 | 1.785 | 0.0030 | No | ||
| 16 | FBXL19 | 2171 | 1.762 | 0.0058 | No | ||
| 17 | DEDD | 2366 | 1.696 | 0.0012 | No | ||
| 18 | PCDH10 | 2527 | 1.656 | -0.0018 | No | ||
| 19 | CANX | 2575 | 1.642 | 0.0014 | No | ||
| 20 | CHD7 | 2600 | 1.635 | 0.0058 | No | ||
| 21 | TRPM2 | 2780 | 1.583 | 0.0016 | No | ||
| 22 | ATXN1 | 2880 | 1.552 | 0.0016 | No | ||
| 23 | PHACTR2 | 3387 | 1.422 | -0.0210 | No | ||
| 24 | SEPT4 | 3741 | 1.342 | -0.0355 | No | ||
| 25 | PTPRM | 3848 | 1.319 | -0.0366 | No | ||
| 26 | ITPK1 | 4439 | 1.191 | -0.0645 | No | ||
| 27 | DYRK1B | 4493 | 1.179 | -0.0633 | No | ||
| 28 | DPP3 | 4621 | 1.151 | -0.0662 | No | ||
| 29 | ERBB3 | 4759 | 1.124 | -0.0697 | No | ||
| 30 | B4GALT5 | 4805 | 1.115 | -0.0683 | No | ||
| 31 | BTBD3 | 5145 | 1.053 | -0.0830 | No | ||
| 32 | RNF38 | 5218 | 1.039 | -0.0833 | No | ||
| 33 | MLLT6 | 5326 | 1.019 | -0.0856 | No | ||
| 34 | CHRD | 5327 | 1.019 | -0.0821 | No | ||
| 35 | ROBO1 | 5371 | 1.008 | -0.0809 | No | ||
| 36 | CUL5 | 5535 | 0.976 | -0.0863 | No | ||
| 37 | GDF6 | 5558 | 0.973 | -0.0842 | No | ||
| 38 | ZFPM2 | 5771 | 0.934 | -0.0924 | No | ||
| 39 | HOXC8 | 5865 | 0.918 | -0.0943 | No | ||
| 40 | GADD45A | 5991 | 0.894 | -0.0980 | No | ||
| 41 | ANK2 | 6021 | 0.888 | -0.0964 | No | ||
| 42 | ROBO2 | 6179 | 0.859 | -0.1020 | No | ||
| 43 | BACH2 | 6215 | 0.851 | -0.1009 | No | ||
| 44 | BTBD11 | 6239 | 0.848 | -0.0992 | No | ||
| 45 | GPM6A | 6271 | 0.842 | -0.0980 | No | ||
| 46 | ABCA1 | 6317 | 0.834 | -0.0975 | No | ||
| 47 | PRKAA1 | 6369 | 0.826 | -0.0974 | No | ||
| 48 | MUM1L1 | 6428 | 0.814 | -0.0977 | No | ||
| 49 | TMSB10 | 6729 | 0.758 | -0.1114 | No | ||
| 50 | SLC24A3 | 7098 | 0.699 | -0.1289 | No | ||
| 51 | MAP3K9 | 7261 | 0.670 | -0.1354 | No | ||
| 52 | LDLR | 7285 | 0.666 | -0.1343 | No | ||
| 53 | ACVR1 | 7384 | 0.648 | -0.1374 | No | ||
| 54 | SYT1 | 7444 | 0.637 | -0.1384 | No | ||
| 55 | TRPS1 | 7667 | 0.593 | -0.1484 | No | ||
| 56 | MAF1 | 7802 | 0.567 | -0.1537 | No | ||
| 57 | SNF1LK | 7841 | 0.559 | -0.1538 | No | ||
| 58 | TEAD1 | 8022 | 0.526 | -0.1618 | No | ||
| 59 | NDST1 | 8029 | 0.525 | -0.1603 | No | ||
| 60 | RORB | 8056 | 0.520 | -0.1599 | No | ||
| 61 | EPS15 | 8074 | 0.516 | -0.1590 | No | ||
| 62 | NPTX1 | 8175 | 0.496 | -0.1627 | No | ||
| 63 | ARHGEF12 | 8323 | 0.469 | -0.1690 | No | ||
| 64 | FBN1 | 8415 | 0.453 | -0.1724 | No | ||
| 65 | JPH3 | 8521 | 0.432 | -0.1766 | No | ||
| 66 | HMGB3 | 8693 | 0.400 | -0.1845 | No | ||
| 67 | PLEKHH1 | 8763 | 0.387 | -0.1869 | No | ||
| 68 | BZRAP1 | 8991 | 0.346 | -0.1980 | No | ||
| 69 | TGIF2 | 9008 | 0.342 | -0.1977 | No | ||
| 70 | BCL11A | 9418 | 0.251 | -0.2190 | No | ||
| 71 | RGMA | 9443 | 0.245 | -0.2195 | No | ||
| 72 | ESR1 | 9562 | 0.220 | -0.2251 | No | ||
| 73 | ESRRG | 9810 | 0.163 | -0.2380 | No | ||
| 74 | ZCCHC2 | 10282 | 0.069 | -0.2633 | No | ||
| 75 | EDG1 | 10290 | 0.068 | -0.2635 | No | ||
| 76 | PRICKLE2 | 10800 | -0.051 | -0.2909 | No | ||
| 77 | MTF1 | 11168 | -0.133 | -0.3104 | No | ||
| 78 | CSF1 | 11177 | -0.135 | -0.3104 | No | ||
| 79 | GPRC5B | 11224 | -0.144 | -0.3124 | No | ||
| 80 | MAFB | 11225 | -0.145 | -0.3119 | No | ||
| 81 | AKAP7 | 11264 | -0.152 | -0.3134 | No | ||
| 82 | ITGA5 | 11275 | -0.154 | -0.3134 | No | ||
| 83 | SOX11 | 11402 | -0.185 | -0.3196 | No | ||
| 84 | UBE2D1 | 11408 | -0.187 | -0.3192 | No | ||
| 85 | PBXIP1 | 11505 | -0.210 | -0.3237 | No | ||
| 86 | TAF4 | 11511 | -0.211 | -0.3233 | No | ||
| 87 | ADAMTS18 | 11584 | -0.226 | -0.3264 | No | ||
| 88 | DNER | 11770 | -0.268 | -0.3355 | No | ||
| 89 | ATP7A | 11898 | -0.299 | -0.3413 | No | ||
| 90 | MGAT4A | 11922 | -0.304 | -0.3415 | No | ||
| 91 | BRPF1 | 11930 | -0.306 | -0.3409 | No | ||
| 92 | NOG | 11937 | -0.308 | -0.3401 | No | ||
| 93 | SSR1 | 12008 | -0.325 | -0.3428 | No | ||
| 94 | ZNRF1 | 12013 | -0.326 | -0.3419 | No | ||
| 95 | LIN28 | 12044 | -0.335 | -0.3423 | No | ||
| 96 | RSBN1L | 12389 | -0.415 | -0.3596 | No | ||
| 97 | E2F7 | 12496 | -0.447 | -0.3638 | No | ||
| 98 | EFNB2 | 12732 | -0.503 | -0.3748 | No | ||
| 99 | BAZ2A | 12744 | -0.508 | -0.3736 | No | ||
| 100 | PSCD3 | 12860 | -0.536 | -0.3780 | No | ||
| 101 | SLC25A3 | 12904 | -0.546 | -0.3784 | No | ||
| 102 | E2F3 | 12935 | -0.554 | -0.3781 | No | ||
| 103 | SIX4 | 13005 | -0.576 | -0.3799 | No | ||
| 104 | WNT10B | 13116 | -0.607 | -0.3837 | No | ||
| 105 | CLOCK | 13167 | -0.620 | -0.3843 | No | ||
| 106 | FBXO33 | 13320 | -0.667 | -0.3902 | No | ||
| 107 | MITF | 13381 | -0.686 | -0.3911 | No | ||
| 108 | FOSB | 13389 | -0.687 | -0.3891 | No | ||
| 109 | UCP3 | 13400 | -0.691 | -0.3872 | No | ||
| 110 | RXRA | 13420 | -0.696 | -0.3858 | No | ||
| 111 | DICER1 | 13505 | -0.720 | -0.3879 | No | ||
| 112 | COL2A1 | 13516 | -0.723 | -0.3859 | No | ||
| 113 | INHBB | 13519 | -0.724 | -0.3835 | No | ||
| 114 | AKAP1 | 13665 | -0.770 | -0.3887 | No | ||
| 115 | EIF2C4 | 13719 | -0.789 | -0.3888 | No | ||
| 116 | STARD13 | 13801 | -0.811 | -0.3904 | No | ||
| 117 | PPARGC1A | 14078 | -0.893 | -0.4023 | No | ||
| 118 | CAMK2A | 14111 | -0.907 | -0.4009 | No | ||
| 119 | GAP43 | 14170 | -0.923 | -0.4008 | No | ||
| 120 | SLC7A11 | 14220 | -0.939 | -0.4002 | No | ||
| 121 | SIRT7 | 14261 | -0.951 | -0.3991 | No | ||
| 122 | CDK5R1 | 14394 | -0.995 | -0.4028 | No | ||
| 123 | KLC2 | 14477 | -1.020 | -0.4037 | No | ||
| 124 | MAMDC1 | 14563 | -1.044 | -0.4047 | No | ||
| 125 | RUNX2 | 14630 | -1.071 | -0.4045 | No | ||
| 126 | MEOX2 | 14655 | -1.079 | -0.4021 | No | ||
| 127 | RAB34 | 14714 | -1.103 | -0.4014 | No | ||
| 128 | NME7 | 14727 | -1.107 | -0.3982 | No | ||
| 129 | B4GALT2 | 14837 | -1.144 | -0.4001 | No | ||
| 130 | EPN2 | 14964 | -1.193 | -0.4028 | No | ||
| 131 | CHD9 | 15073 | -1.235 | -0.4044 | No | ||
| 132 | ARHGEF17 | 15112 | -1.254 | -0.4021 | No | ||
| 133 | TRAK2 | 15497 | -1.405 | -0.4181 | No | ||
| 134 | SULF1 | 15616 | -1.452 | -0.4194 | Yes | ||
| 135 | LMTK2 | 15668 | -1.476 | -0.4171 | Yes | ||
| 136 | SMS | 15696 | -1.489 | -0.4134 | Yes | ||
| 137 | CABP7 | 15730 | -1.503 | -0.4099 | Yes | ||
| 138 | PICALM | 15769 | -1.519 | -0.4067 | Yes | ||
| 139 | ARFGEF1 | 15799 | -1.532 | -0.4029 | Yes | ||
| 140 | CPD | 15815 | -1.539 | -0.3984 | Yes | ||
| 141 | QKI | 15817 | -1.540 | -0.3931 | Yes | ||
| 142 | PDK4 | 15824 | -1.544 | -0.3881 | Yes | ||
| 143 | EIF2C1 | 15942 | -1.597 | -0.3889 | Yes | ||
| 144 | CNTN4 | 15976 | -1.613 | -0.3850 | Yes | ||
| 145 | KLF4 | 16058 | -1.654 | -0.3837 | Yes | ||
| 146 | BCL7A | 16072 | -1.659 | -0.3786 | Yes | ||
| 147 | ARRDC3 | 16120 | -1.680 | -0.3753 | Yes | ||
| 148 | MMD | 16151 | -1.693 | -0.3711 | Yes | ||
| 149 | MAX | 16283 | -1.753 | -0.3721 | Yes | ||
| 150 | DDX6 | 16290 | -1.756 | -0.3663 | Yes | ||
| 151 | HAS3 | 16307 | -1.766 | -0.3610 | Yes | ||
| 152 | BMP1 | 16333 | -1.779 | -0.3562 | Yes | ||
| 153 | USP33 | 16351 | -1.789 | -0.3509 | Yes | ||
| 154 | GMFB | 16539 | -1.876 | -0.3545 | Yes | ||
| 155 | USP48 | 16573 | -1.893 | -0.3497 | Yes | ||
| 156 | OTX1 | 16622 | -1.922 | -0.3456 | Yes | ||
| 157 | WDR20 | 16698 | -1.958 | -0.3429 | Yes | ||
| 158 | TNRC6A | 16743 | -1.984 | -0.3384 | Yes | ||
| 159 | MLLT10 | 16774 | -2.004 | -0.3331 | Yes | ||
| 160 | CFL2 | 16783 | -2.009 | -0.3265 | Yes | ||
| 161 | LTBP1 | 16784 | -2.009 | -0.3195 | Yes | ||
| 162 | NCOA1 | 16840 | -2.036 | -0.3154 | Yes | ||
| 163 | PAPD4 | 16842 | -2.038 | -0.3084 | Yes | ||
| 164 | TOMM70A | 16846 | -2.039 | -0.3015 | Yes | ||
| 165 | CDC14A | 16915 | -2.072 | -0.2979 | Yes | ||
| 166 | WNT1 | 16941 | -2.088 | -0.2920 | Yes | ||
| 167 | RPS6KA5 | 16943 | -2.089 | -0.2848 | Yes | ||
| 168 | PPP1R12A | 17014 | -2.128 | -0.2812 | Yes | ||
| 169 | ELF5 | 17022 | -2.134 | -0.2742 | Yes | ||
| 170 | ARL8B | 17040 | -2.142 | -0.2676 | Yes | ||
| 171 | YWHAB | 17144 | -2.202 | -0.2656 | Yes | ||
| 172 | RAB35 | 17355 | -2.324 | -0.2689 | Yes | ||
| 173 | ST8SIA3 | 17409 | -2.354 | -0.2636 | Yes | ||
| 174 | ABCB7 | 17430 | -2.367 | -0.2564 | Yes | ||
| 175 | RALBP1 | 17451 | -2.381 | -0.2492 | Yes | ||
| 176 | CNOT4 | 17473 | -2.390 | -0.2421 | Yes | ||
| 177 | PPP1CB | 17671 | -2.520 | -0.2440 | Yes | ||
| 178 | ARFIP1 | 17696 | -2.534 | -0.2365 | Yes | ||
| 179 | OSBPL11 | 17740 | -2.567 | -0.2299 | Yes | ||
| 180 | ZBED4 | 17759 | -2.576 | -0.2219 | Yes | ||
| 181 | PDIA3 | 17798 | -2.602 | -0.2149 | Yes | ||
| 182 | ZFYVE26 | 17814 | -2.612 | -0.2066 | Yes | ||
| 183 | NLK | 17875 | -2.665 | -0.2006 | Yes | ||
| 184 | UBR1 | 17929 | -2.703 | -0.1941 | Yes | ||
| 185 | SFRS2IP | 17936 | -2.710 | -0.1850 | Yes | ||
| 186 | PLAA | 17943 | -2.715 | -0.1759 | Yes | ||
| 187 | RNF44 | 17956 | -2.727 | -0.1670 | Yes | ||
| 188 | MTMR12 | 18007 | -2.771 | -0.1601 | Yes | ||
| 189 | ITSN2 | 18042 | -2.802 | -0.1522 | Yes | ||
| 190 | C1GALT1 | 18044 | -2.805 | -0.1425 | Yes | ||
| 191 | CYB5R4 | 18073 | -2.832 | -0.1342 | Yes | ||
| 192 | RNF12 | 18176 | -2.944 | -0.1295 | Yes | ||
| 193 | LBR | 18241 | -3.037 | -0.1224 | Yes | ||
| 194 | NARG1 | 18243 | -3.040 | -0.1119 | Yes | ||
| 195 | ARPP-19 | 18261 | -3.075 | -0.1021 | Yes | ||
| 196 | MOSPD1 | 18298 | -3.130 | -0.0932 | Yes | ||
| 197 | DNMT1 | 18303 | -3.135 | -0.0825 | Yes | ||
| 198 | SKP1A | 18307 | -3.141 | -0.0717 | Yes | ||
| 199 | XPO4 | 18383 | -3.265 | -0.0644 | Yes | ||
| 200 | ATP2A2 | 18465 | -3.491 | -0.0567 | Yes | ||
| 201 | POU4F2 | 18534 | -3.788 | -0.0472 | Yes | ||
| 202 | TGFA | 18567 | -4.065 | -0.0348 | Yes | ||
| 203 | PIK3R3 | 18604 | -4.890 | -0.0197 | Yes | ||
| 204 | ARHGAP21 | 18614 | -5.846 | 0.0001 | Yes |
