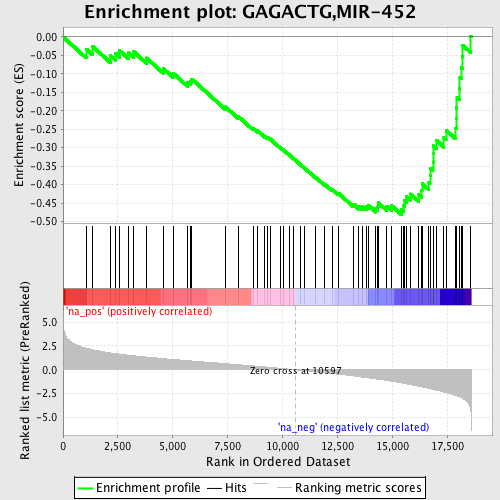
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | set03_truncNotch_versus_wtNotch |
| Phenotype | NoPhenotypeAvailable |
| Upregulated in class | na_neg |
| GeneSet | GAGACTG,MIR-452 |
| Enrichment Score (ES) | -0.4811004 |
| Normalized Enrichment Score (NES) | -2.0850134 |
| Nominal p-value | 0.0 |
| FDR q-value | 6.460215E-4 |
| FWER p-Value | 0.0080 |

| PROBE | GENE SYMBOL | GENE_TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|---|
| 1 | SH3GLB1 | 1052 | 2.253 | -0.0326 | No | ||
| 2 | EEF2 | 1329 | 2.099 | -0.0249 | No | ||
| 3 | LPHN1 | 2164 | 1.765 | -0.0510 | No | ||
| 4 | RBM14 | 2381 | 1.692 | -0.0445 | No | ||
| 5 | FLRT3 | 2579 | 1.642 | -0.0375 | No | ||
| 6 | DCAMKL1 | 2988 | 1.524 | -0.0431 | No | ||
| 7 | CTBP1 | 3221 | 1.462 | -0.0400 | No | ||
| 8 | TLN1 | 3800 | 1.329 | -0.0569 | No | ||
| 9 | NXPH1 | 4561 | 1.165 | -0.0854 | No | ||
| 10 | PLXNB1 | 5015 | 1.079 | -0.0982 | No | ||
| 11 | MEIS1 | 5679 | 0.952 | -0.1238 | No | ||
| 12 | GGA3 | 5791 | 0.930 | -0.1198 | No | ||
| 13 | HOXC8 | 5865 | 0.918 | -0.1139 | No | ||
| 14 | EN1 | 7399 | 0.645 | -0.1896 | No | ||
| 15 | ACVR2A | 7995 | 0.531 | -0.2160 | No | ||
| 16 | EPHB3 | 8658 | 0.406 | -0.2473 | No | ||
| 17 | CILP | 8857 | 0.372 | -0.2540 | No | ||
| 18 | MTMR4 | 9191 | 0.300 | -0.2687 | No | ||
| 19 | AXIN2 | 9321 | 0.276 | -0.2727 | No | ||
| 20 | PPM1G | 9438 | 0.246 | -0.2764 | No | ||
| 21 | ZIC1 | 9924 | 0.143 | -0.3010 | No | ||
| 22 | PPM2C | 10028 | 0.123 | -0.3052 | No | ||
| 23 | ITPKA | 10300 | 0.065 | -0.3191 | No | ||
| 24 | DYNLL1 | 10492 | 0.022 | -0.3292 | No | ||
| 25 | PRICKLE2 | 10800 | -0.051 | -0.3452 | No | ||
| 26 | SLC25A27 | 11012 | -0.097 | -0.3555 | No | ||
| 27 | TAF4 | 11511 | -0.211 | -0.3801 | No | ||
| 28 | LZTS1 | 11934 | -0.308 | -0.3996 | No | ||
| 29 | NPEPPS | 12259 | -0.384 | -0.4129 | No | ||
| 30 | CBX1 | 12541 | -0.458 | -0.4231 | No | ||
| 31 | DYRK1A | 13236 | -0.637 | -0.4537 | No | ||
| 32 | SLC6A9 | 13481 | -0.712 | -0.4592 | No | ||
| 33 | GPR126 | 13641 | -0.764 | -0.4596 | No | ||
| 34 | MOBKL2B | 13808 | -0.813 | -0.4598 | No | ||
| 35 | DPP10 | 13906 | -0.836 | -0.4561 | No | ||
| 36 | ABCC5 | 14247 | -0.948 | -0.4643 | No | ||
| 37 | RORC | 14348 | -0.980 | -0.4592 | No | ||
| 38 | TNRC15 | 14363 | -0.986 | -0.4493 | No | ||
| 39 | ZDHHC5 | 14758 | -1.119 | -0.4586 | No | ||
| 40 | TCF4 | 14973 | -1.198 | -0.4573 | No | ||
| 41 | LDB1 | 15416 | -1.370 | -0.4664 | Yes | ||
| 42 | BCL2L2 | 15529 | -1.416 | -0.4573 | Yes | ||
| 43 | BCAS3 | 15567 | -1.432 | -0.4439 | Yes | ||
| 44 | PRRX1 | 15649 | -1.465 | -0.4326 | Yes | ||
| 45 | P4HA1 | 15827 | -1.545 | -0.4255 | Yes | ||
| 46 | CDC42EP3 | 16215 | -1.721 | -0.4279 | Yes | ||
| 47 | PAIP2 | 16330 | -1.778 | -0.4150 | Yes | ||
| 48 | RSBN1 | 16360 | -1.791 | -0.3974 | Yes | ||
| 49 | PLA2G4A | 16673 | -1.943 | -0.3934 | Yes | ||
| 50 | TCF12 | 16722 | -1.970 | -0.3748 | Yes | ||
| 51 | RAB8B | 16757 | -1.998 | -0.3553 | Yes | ||
| 52 | MAPK6 | 16859 | -2.048 | -0.3387 | Yes | ||
| 53 | HIP2 | 16863 | -2.053 | -0.3169 | Yes | ||
| 54 | TCF20 | 16867 | -2.054 | -0.2950 | Yes | ||
| 55 | PPP1R12A | 17014 | -2.128 | -0.2801 | Yes | ||
| 56 | RAB35 | 17355 | -2.324 | -0.2735 | Yes | ||
| 57 | SYNCRIP | 17454 | -2.383 | -0.2532 | Yes | ||
| 58 | CUGBP1 | 17880 | -2.670 | -0.2475 | Yes | ||
| 59 | GTF2E1 | 17912 | -2.692 | -0.2203 | Yes | ||
| 60 | ADIPOR2 | 17918 | -2.696 | -0.1917 | Yes | ||
| 61 | OGT | 17951 | -2.718 | -0.1643 | Yes | ||
| 62 | CNOT7 | 18067 | -2.828 | -0.1401 | Yes | ||
| 63 | EIF4E | 18085 | -2.842 | -0.1106 | Yes | ||
| 64 | SP1 | 18146 | -2.912 | -0.0826 | Yes | ||
| 65 | RHOT1 | 18188 | -2.961 | -0.0531 | Yes | ||
| 66 | PPP2CA | 18211 | -2.984 | -0.0222 | Yes | ||
| 67 | CALU | 18572 | -4.106 | 0.0024 | Yes |
