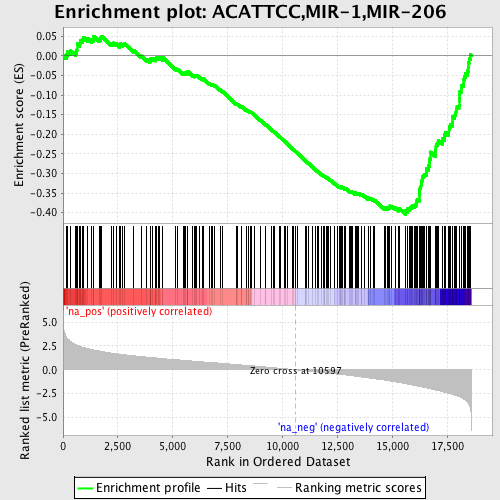
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | set03_truncNotch_versus_wtNotch |
| Phenotype | NoPhenotypeAvailable |
| Upregulated in class | na_neg |
| GeneSet | ACATTCC,MIR-1,MIR-206 |
| Enrichment Score (ES) | -0.40479943 |
| Normalized Enrichment Score (NES) | -2.1056468 |
| Nominal p-value | 0.0 |
| FDR q-value | 3.8949715E-4 |
| FWER p-Value | 0.0040 |

| PROBE | GENE SYMBOL | GENE_TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|---|
| 1 | SMARCB1 | 139 | 3.434 | 0.0037 | No | ||
| 2 | RKHD2 | 216 | 3.218 | 0.0102 | No | ||
| 3 | CELSR3 | 350 | 2.945 | 0.0126 | No | ||
| 4 | GRK6 | 578 | 2.636 | 0.0089 | No | ||
| 5 | MNT | 624 | 2.589 | 0.0150 | No | ||
| 6 | PHC2 | 635 | 2.579 | 0.0229 | No | ||
| 7 | TAGLN2 | 636 | 2.579 | 0.0314 | No | ||
| 8 | PIAS3 | 761 | 2.463 | 0.0327 | No | ||
| 9 | NR1H3 | 801 | 2.434 | 0.0386 | No | ||
| 10 | ACACA | 886 | 2.366 | 0.0418 | No | ||
| 11 | TRIM2 | 943 | 2.323 | 0.0464 | No | ||
| 12 | DGKZ | 1131 | 2.208 | 0.0435 | No | ||
| 13 | GCH1 | 1294 | 2.113 | 0.0416 | No | ||
| 14 | TSPAN4 | 1369 | 2.082 | 0.0444 | No | ||
| 15 | KTN1 | 1386 | 2.074 | 0.0504 | No | ||
| 16 | SEMA6D | 1641 | 1.966 | 0.0430 | No | ||
| 17 | SH3BGRL3 | 1709 | 1.940 | 0.0458 | No | ||
| 18 | RNF13 | 1757 | 1.920 | 0.0495 | No | ||
| 19 | BICD1 | 2188 | 1.754 | 0.0319 | No | ||
| 20 | CREB5 | 2274 | 1.724 | 0.0330 | No | ||
| 21 | TRAPPC3 | 2416 | 1.683 | 0.0308 | No | ||
| 22 | RSPO3 | 2589 | 1.639 | 0.0269 | No | ||
| 23 | ANXA4 | 2609 | 1.633 | 0.0312 | No | ||
| 24 | RIT2 | 2725 | 1.597 | 0.0302 | No | ||
| 25 | ADAR | 2781 | 1.582 | 0.0324 | No | ||
| 26 | SRGAP2 | 3213 | 1.466 | 0.0138 | No | ||
| 27 | MAP1A | 3556 | 1.386 | -0.0003 | No | ||
| 28 | ETS1 | 3817 | 1.325 | -0.0100 | No | ||
| 29 | SLC37A3 | 3961 | 1.295 | -0.0136 | No | ||
| 30 | YPEL2 | 3974 | 1.293 | -0.0100 | No | ||
| 31 | HIVEP2 | 4003 | 1.287 | -0.0072 | No | ||
| 32 | SYNJ2 | 4068 | 1.275 | -0.0065 | No | ||
| 33 | FZD7 | 4190 | 1.249 | -0.0090 | No | ||
| 34 | TBC1D15 | 4228 | 1.240 | -0.0070 | No | ||
| 35 | DAAM1 | 4252 | 1.235 | -0.0041 | No | ||
| 36 | NR4A2 | 4328 | 1.217 | -0.0042 | No | ||
| 37 | PTPRG | 4408 | 1.198 | -0.0046 | No | ||
| 38 | MYLK | 4541 | 1.166 | -0.0079 | No | ||
| 39 | HS3ST3B1 | 4544 | 1.166 | -0.0042 | No | ||
| 40 | GPR125 | 5134 | 1.054 | -0.0328 | No | ||
| 41 | RNF38 | 5218 | 1.039 | -0.0339 | No | ||
| 42 | SNAP25 | 5493 | 0.983 | -0.0455 | No | ||
| 43 | CPLX2 | 5519 | 0.980 | -0.0437 | No | ||
| 44 | GDF6 | 5558 | 0.973 | -0.0425 | No | ||
| 45 | ZBTB4 | 5664 | 0.955 | -0.0451 | No | ||
| 46 | BET1 | 5671 | 0.953 | -0.0423 | No | ||
| 47 | MEIS1 | 5679 | 0.952 | -0.0396 | No | ||
| 48 | TMOD2 | 5917 | 0.905 | -0.0495 | No | ||
| 49 | SLC38A3 | 6001 | 0.893 | -0.0510 | No | ||
| 50 | CDK6 | 6023 | 0.888 | -0.0493 | No | ||
| 51 | RARB | 6070 | 0.879 | -0.0489 | No | ||
| 52 | BACH2 | 6215 | 0.851 | -0.0539 | No | ||
| 53 | CITED2 | 6359 | 0.828 | -0.0590 | No | ||
| 54 | SUHW4 | 6382 | 0.823 | -0.0575 | No | ||
| 55 | HIC2 | 6656 | 0.773 | -0.0698 | No | ||
| 56 | SLC7A2 | 6761 | 0.753 | -0.0729 | No | ||
| 57 | VGLL4 | 6824 | 0.738 | -0.0739 | No | ||
| 58 | TNPO2 | 6889 | 0.729 | -0.0750 | No | ||
| 59 | CREM | 7150 | 0.689 | -0.0868 | No | ||
| 60 | LRRC8A | 7260 | 0.670 | -0.0906 | No | ||
| 61 | STC2 | 7892 | 0.549 | -0.1230 | No | ||
| 62 | SFRP1 | 7906 | 0.547 | -0.1220 | No | ||
| 63 | MAB21L1 | 7967 | 0.537 | -0.1235 | No | ||
| 64 | RIMS4 | 8116 | 0.508 | -0.1298 | No | ||
| 65 | NAP1L5 | 8117 | 0.508 | -0.1282 | No | ||
| 66 | MYO1E | 8342 | 0.466 | -0.1388 | No | ||
| 67 | WDR40B | 8352 | 0.464 | -0.1378 | No | ||
| 68 | MET | 8434 | 0.448 | -0.1407 | No | ||
| 69 | TKT | 8471 | 0.441 | -0.1412 | No | ||
| 70 | CEBPZ | 8530 | 0.430 | -0.1429 | No | ||
| 71 | GAS2L1 | 8565 | 0.423 | -0.1434 | No | ||
| 72 | VAMP2 | 8735 | 0.393 | -0.1513 | No | ||
| 73 | TGIF2 | 9008 | 0.342 | -0.1650 | No | ||
| 74 | WDR6 | 9204 | 0.296 | -0.1746 | No | ||
| 75 | ANKRD29 | 9243 | 0.290 | -0.1757 | No | ||
| 76 | UBN1 | 9494 | 0.234 | -0.1885 | No | ||
| 77 | EYA4 | 9595 | 0.213 | -0.1933 | No | ||
| 78 | POGK | 9637 | 0.202 | -0.1948 | No | ||
| 79 | NGFR | 9847 | 0.158 | -0.2057 | No | ||
| 80 | MIPOL1 | 9895 | 0.148 | -0.2077 | No | ||
| 81 | TFEC | 10081 | 0.113 | -0.2174 | No | ||
| 82 | KCTD13 | 10123 | 0.104 | -0.2193 | No | ||
| 83 | GJA1 | 10216 | 0.082 | -0.2240 | No | ||
| 84 | CDK9 | 10456 | 0.030 | -0.2369 | No | ||
| 85 | SLC25A25 | 10493 | 0.022 | -0.2388 | No | ||
| 86 | MAP4K3 | 10573 | 0.003 | -0.2431 | No | ||
| 87 | UBE4A | 10610 | -0.003 | -0.2450 | No | ||
| 88 | SPRED1 | 10692 | -0.025 | -0.2494 | No | ||
| 89 | GDAP1L1 | 11039 | -0.102 | -0.2678 | No | ||
| 90 | TLOC1 | 11067 | -0.109 | -0.2689 | No | ||
| 91 | PPIB | 11095 | -0.113 | -0.2700 | No | ||
| 92 | CLTC | 11166 | -0.132 | -0.2734 | No | ||
| 93 | SFRS9 | 11352 | -0.172 | -0.2829 | No | ||
| 94 | DSCR1L1 | 11489 | -0.207 | -0.2896 | No | ||
| 95 | OLFML2A | 11581 | -0.224 | -0.2938 | No | ||
| 96 | PAX3 | 11631 | -0.235 | -0.2957 | No | ||
| 97 | CORO1C | 11788 | -0.273 | -0.3033 | No | ||
| 98 | COL4A3 | 11850 | -0.288 | -0.3057 | No | ||
| 99 | ATP7A | 11898 | -0.299 | -0.3072 | No | ||
| 100 | MAL2 | 11917 | -0.303 | -0.3072 | No | ||
| 101 | CBL | 12024 | -0.331 | -0.3119 | No | ||
| 102 | SP2 | 12034 | -0.332 | -0.3113 | No | ||
| 103 | SLC25A22 | 12097 | -0.346 | -0.3135 | No | ||
| 104 | BSCL2 | 12182 | -0.367 | -0.3169 | No | ||
| 105 | RSBN1L | 12389 | -0.415 | -0.3267 | No | ||
| 106 | NCOA3 | 12515 | -0.452 | -0.3320 | No | ||
| 107 | EDN1 | 12601 | -0.473 | -0.3351 | No | ||
| 108 | FOXP1 | 12643 | -0.483 | -0.3357 | No | ||
| 109 | SLC8A2 | 12658 | -0.486 | -0.3349 | No | ||
| 110 | MKL1 | 12673 | -0.490 | -0.3340 | No | ||
| 111 | EFNB2 | 12732 | -0.503 | -0.3355 | No | ||
| 112 | ADPGK | 12823 | -0.527 | -0.3387 | No | ||
| 113 | YWHAQ | 12839 | -0.530 | -0.3378 | No | ||
| 114 | SETBP1 | 12882 | -0.541 | -0.3383 | No | ||
| 115 | TGFBR3 | 13065 | -0.593 | -0.3462 | No | ||
| 116 | FN1 | 13081 | -0.597 | -0.3451 | No | ||
| 117 | MAPKBP1 | 13142 | -0.615 | -0.3463 | No | ||
| 118 | SLC35F1 | 13201 | -0.628 | -0.3474 | No | ||
| 119 | CPEB1 | 13307 | -0.664 | -0.3509 | No | ||
| 120 | FBXO33 | 13320 | -0.667 | -0.3494 | No | ||
| 121 | FOSB | 13389 | -0.687 | -0.3508 | No | ||
| 122 | ARHGEF18 | 13433 | -0.699 | -0.3509 | No | ||
| 123 | ANXA2 | 13462 | -0.707 | -0.3501 | No | ||
| 124 | FGF14 | 13588 | -0.748 | -0.3544 | No | ||
| 125 | AKAP11 | 13601 | -0.752 | -0.3526 | No | ||
| 126 | ZFP36L1 | 13742 | -0.797 | -0.3576 | No | ||
| 127 | BSN | 13899 | -0.834 | -0.3633 | No | ||
| 128 | IGF1 | 13921 | -0.840 | -0.3617 | No | ||
| 129 | BDNF | 14026 | -0.876 | -0.3645 | No | ||
| 130 | DGKG | 14124 | -0.910 | -0.3668 | No | ||
| 131 | E2F5 | 14208 | -0.936 | -0.3682 | No | ||
| 132 | MEOX2 | 14655 | -1.079 | -0.3889 | No | ||
| 133 | BLCAP | 14671 | -1.086 | -0.3861 | No | ||
| 134 | NCK2 | 14803 | -1.136 | -0.3895 | No | ||
| 135 | TIMP3 | 14815 | -1.139 | -0.3864 | No | ||
| 136 | HMGN1 | 14895 | -1.165 | -0.3869 | No | ||
| 137 | PFTK1 | 14897 | -1.166 | -0.3831 | No | ||
| 138 | HIAT1 | 14987 | -1.203 | -0.3840 | No | ||
| 139 | LIN7C | 15135 | -1.262 | -0.3878 | No | ||
| 140 | GPD2 | 15288 | -1.322 | -0.3917 | No | ||
| 141 | UBE2H | 15315 | -1.330 | -0.3888 | No | ||
| 142 | BPNT1 | 15611 | -1.449 | -0.4000 | Yes | ||
| 143 | SULF1 | 15616 | -1.452 | -0.3955 | Yes | ||
| 144 | UST | 15697 | -1.489 | -0.3949 | Yes | ||
| 145 | RRBP1 | 15707 | -1.492 | -0.3905 | Yes | ||
| 146 | PICALM | 15769 | -1.519 | -0.3889 | Yes | ||
| 147 | QKI | 15817 | -1.540 | -0.3864 | Yes | ||
| 148 | AP3D1 | 15892 | -1.576 | -0.3852 | Yes | ||
| 149 | EIF2C1 | 15942 | -1.597 | -0.3826 | Yes | ||
| 150 | DDX5 | 16021 | -1.638 | -0.3815 | Yes | ||
| 151 | BCL7A | 16072 | -1.659 | -0.3787 | Yes | ||
| 152 | SLC7A6 | 16111 | -1.678 | -0.3753 | Yes | ||
| 153 | ARRDC3 | 16120 | -1.680 | -0.3702 | Yes | ||
| 154 | MMD | 16151 | -1.693 | -0.3663 | Yes | ||
| 155 | ARCN1 | 16230 | -1.727 | -0.3649 | Yes | ||
| 156 | SMAP1 | 16231 | -1.727 | -0.3592 | Yes | ||
| 157 | ARF3 | 16234 | -1.728 | -0.3536 | Yes | ||
| 158 | FBXW7 | 16248 | -1.732 | -0.3486 | Yes | ||
| 159 | SEC63 | 16256 | -1.735 | -0.3433 | Yes | ||
| 160 | FNDC3A | 16264 | -1.741 | -0.3380 | Yes | ||
| 161 | POLDIP3 | 16295 | -1.758 | -0.3338 | Yes | ||
| 162 | AZIN1 | 16311 | -1.768 | -0.3288 | Yes | ||
| 163 | ZIC4 | 16338 | -1.782 | -0.3244 | Yes | ||
| 164 | USP33 | 16351 | -1.789 | -0.3192 | Yes | ||
| 165 | RSBN1 | 16360 | -1.791 | -0.3137 | Yes | ||
| 166 | GNPDA2 | 16380 | -1.801 | -0.3088 | Yes | ||
| 167 | YWHAZ | 16431 | -1.824 | -0.3056 | Yes | ||
| 168 | OTX2 | 16479 | -1.843 | -0.3021 | Yes | ||
| 169 | TBP | 16552 | -1.880 | -0.2998 | Yes | ||
| 170 | CDC42 | 16568 | -1.892 | -0.2944 | Yes | ||
| 171 | AKAP12 | 16571 | -1.893 | -0.2883 | Yes | ||
| 172 | SMARCA4 | 16653 | -1.934 | -0.2864 | Yes | ||
| 173 | HNRPK | 16662 | -1.937 | -0.2804 | Yes | ||
| 174 | MATR3 | 16687 | -1.952 | -0.2753 | Yes | ||
| 175 | DNAJC13 | 16694 | -1.955 | -0.2692 | Yes | ||
| 176 | SFRS10 | 16701 | -1.959 | -0.2631 | Yes | ||
| 177 | TTC7B | 16733 | -1.980 | -0.2583 | Yes | ||
| 178 | NUP50 | 16735 | -1.980 | -0.2518 | Yes | ||
| 179 | ATP6V1A | 16736 | -1.981 | -0.2453 | Yes | ||
| 180 | STARD7 | 16966 | -2.098 | -0.2509 | Yes | ||
| 181 | PHF6 | 16970 | -2.099 | -0.2442 | Yes | ||
| 182 | CLCN3 | 16985 | -2.113 | -0.2380 | Yes | ||
| 183 | HSPD1 | 16994 | -2.116 | -0.2315 | Yes | ||
| 184 | VAMP4 | 17015 | -2.129 | -0.2256 | Yes | ||
| 185 | PAFAH1B1 | 17084 | -2.165 | -0.2221 | Yes | ||
| 186 | PUM2 | 17116 | -2.184 | -0.2167 | Yes | ||
| 187 | ANP32B | 17275 | -2.275 | -0.2178 | Yes | ||
| 188 | RNUXA | 17283 | -2.278 | -0.2107 | Yes | ||
| 189 | HMGCR | 17371 | -2.332 | -0.2077 | Yes | ||
| 190 | CNN3 | 17391 | -2.343 | -0.2011 | Yes | ||
| 191 | ABCB7 | 17430 | -2.367 | -0.1954 | Yes | ||
| 192 | KLF13 | 17576 | -2.454 | -0.1952 | Yes | ||
| 193 | NCL | 17587 | -2.462 | -0.1876 | Yes | ||
| 194 | EAF1 | 17599 | -2.476 | -0.1801 | Yes | ||
| 195 | SLC39A10 | 17667 | -2.516 | -0.1755 | Yes | ||
| 196 | HNRPU | 17736 | -2.564 | -0.1708 | Yes | ||
| 197 | RNGTT | 17757 | -2.575 | -0.1634 | Yes | ||
| 198 | SMYD4 | 17760 | -2.576 | -0.1550 | Yes | ||
| 199 | DNAJC5 | 17847 | -2.642 | -0.1510 | Yes | ||
| 200 | SFRS3 | 17871 | -2.665 | -0.1435 | Yes | ||
| 201 | SNX2 | 17908 | -2.691 | -0.1366 | Yes | ||
| 202 | OGT | 17951 | -2.718 | -0.1300 | Yes | ||
| 203 | C1GALT1 | 18044 | -2.805 | -0.1258 | Yes | ||
| 204 | PPARBP | 18053 | -2.818 | -0.1170 | Yes | ||
| 205 | DHX15 | 18058 | -2.822 | -0.1079 | Yes | ||
| 206 | FRAS1 | 18072 | -2.831 | -0.0993 | Yes | ||
| 207 | EIF4E | 18085 | -2.842 | -0.0906 | Yes | ||
| 208 | RASA1 | 18140 | -2.908 | -0.0840 | Yes | ||
| 209 | ARID2 | 18148 | -2.918 | -0.0748 | Yes | ||
| 210 | UBQLN1 | 18232 | -3.025 | -0.0694 | Yes | ||
| 211 | PDCD10 | 18239 | -3.036 | -0.0598 | Yes | ||
| 212 | TPM3 | 18287 | -3.115 | -0.0521 | Yes | ||
| 213 | SFRS1 | 18333 | -3.183 | -0.0441 | Yes | ||
| 214 | HOXB4 | 18436 | -3.412 | -0.0384 | Yes | ||
| 215 | RNF138 | 18464 | -3.488 | -0.0284 | Yes | ||
| 216 | SS18 | 18476 | -3.521 | -0.0174 | Yes | ||
| 217 | AP1G1 | 18502 | -3.623 | -0.0069 | Yes | ||
| 218 | BAG4 | 18557 | -3.972 | 0.0032 | Yes |
