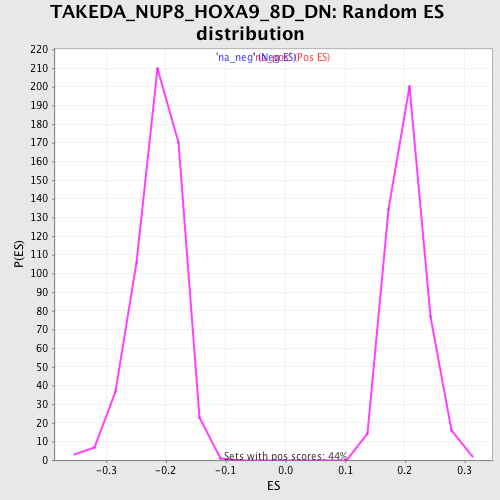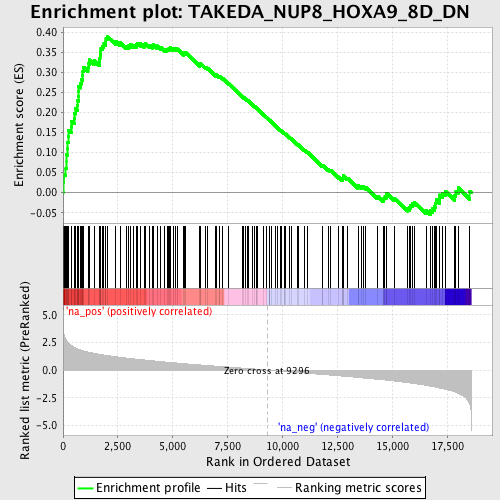
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | set03_absentNotch_versus_wtNotch |
| Phenotype | NoPhenotypeAvailable |
| Upregulated in class | na_pos |
| GeneSet | TAKEDA_NUP8_HOXA9_8D_DN |
| Enrichment Score (ES) | 0.3900322 |
| Normalized Enrichment Score (NES) | 1.9177793 |
| Nominal p-value | 0.0 |
| FDR q-value | 0.108842686 |
| FWER p-Value | 0.504 |

| PROBE | GENE SYMBOL | GENE_TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|---|
| 1 | CLEC7A | 4 | 4.013 | 0.0266 | Yes | ||
| 2 | SGK | 40 | 3.258 | 0.0465 | Yes | ||
| 3 | CUTL1 | 127 | 2.767 | 0.0603 | Yes | ||
| 4 | IL1R2 | 144 | 2.711 | 0.0776 | Yes | ||
| 5 | FCGR2B | 146 | 2.702 | 0.0956 | Yes | ||
| 6 | MMP14 | 209 | 2.497 | 0.1089 | Yes | ||
| 7 | NCF2 | 213 | 2.488 | 0.1254 | Yes | ||
| 8 | HIPK1 | 230 | 2.457 | 0.1409 | Yes | ||
| 9 | EMP1 | 262 | 2.386 | 0.1552 | Yes | ||
| 10 | UHMK1 | 369 | 2.221 | 0.1643 | Yes | ||
| 11 | ELOVL7 | 386 | 2.203 | 0.1782 | Yes | ||
| 12 | ANTXR1 | 514 | 2.053 | 0.1850 | Yes | ||
| 13 | DAB2 | 529 | 2.043 | 0.1979 | Yes | ||
| 14 | SLC7A7 | 567 | 2.003 | 0.2093 | Yes | ||
| 15 | PPP1R3B | 662 | 1.909 | 0.2170 | Yes | ||
| 16 | CYBB | 672 | 1.905 | 0.2292 | Yes | ||
| 17 | C1QC | 694 | 1.889 | 0.2407 | Yes | ||
| 18 | LGMN | 702 | 1.884 | 0.2529 | Yes | ||
| 19 | TRPA1 | 708 | 1.881 | 0.2652 | Yes | ||
| 20 | SLC11A1 | 792 | 1.822 | 0.2729 | Yes | ||
| 21 | LFNG | 834 | 1.795 | 0.2827 | Yes | ||
| 22 | CD9 | 880 | 1.766 | 0.2921 | Yes | ||
| 23 | C1QB | 903 | 1.746 | 0.3025 | Yes | ||
| 24 | C1QA | 928 | 1.734 | 0.3128 | Yes | ||
| 25 | ARPC5 | 1135 | 1.634 | 0.3126 | Yes | ||
| 26 | GPNMB | 1169 | 1.622 | 0.3217 | Yes | ||
| 27 | CD24 | 1201 | 1.611 | 0.3307 | Yes | ||
| 28 | CXCL5 | 1413 | 1.519 | 0.3295 | Yes | ||
| 29 | SLCO2B1 | 1657 | 1.421 | 0.3258 | Yes | ||
| 30 | CDH8 | 1665 | 1.419 | 0.3349 | Yes | ||
| 31 | MMP12 | 1682 | 1.414 | 0.3435 | Yes | ||
| 32 | CLU | 1718 | 1.402 | 0.3510 | Yes | ||
| 33 | PRNP | 1724 | 1.401 | 0.3601 | Yes | ||
| 34 | HTR2C | 1805 | 1.375 | 0.3649 | Yes | ||
| 35 | TGFBI | 1861 | 1.359 | 0.3710 | Yes | ||
| 36 | EXT1 | 1946 | 1.333 | 0.3754 | Yes | ||
| 37 | ENC1 | 1953 | 1.330 | 0.3840 | Yes | ||
| 38 | HK3 | 2004 | 1.313 | 0.3900 | Yes | ||
| 39 | PRDM2 | 2391 | 1.210 | 0.3772 | No | ||
| 40 | SLC25A27 | 2605 | 1.157 | 0.3734 | No | ||
| 41 | RAP1A | 2888 | 1.081 | 0.3654 | No | ||
| 42 | SLAMF8 | 3001 | 1.059 | 0.3664 | No | ||
| 43 | FYB | 3078 | 1.043 | 0.3693 | No | ||
| 44 | IFNGR1 | 3229 | 1.012 | 0.3679 | No | ||
| 45 | JUN | 3331 | 0.989 | 0.3690 | No | ||
| 46 | PYGL | 3384 | 0.976 | 0.3727 | No | ||
| 47 | IGF2R | 3525 | 0.952 | 0.3715 | No | ||
| 48 | S100A8 | 3690 | 0.918 | 0.3688 | No | ||
| 49 | F5 | 3736 | 0.909 | 0.3724 | No | ||
| 50 | GPR34 | 3944 | 0.863 | 0.3670 | No | ||
| 51 | SLC30A7 | 4070 | 0.839 | 0.3658 | No | ||
| 52 | ATP6V0D2 | 4112 | 0.831 | 0.3692 | No | ||
| 53 | CTSB | 4279 | 0.800 | 0.3655 | No | ||
| 54 | ABP1 | 4434 | 0.770 | 0.3623 | No | ||
| 55 | IL10 | 4621 | 0.733 | 0.3572 | No | ||
| 56 | BCL6 | 4734 | 0.715 | 0.3559 | No | ||
| 57 | KCNMA1 | 4783 | 0.708 | 0.3580 | No | ||
| 58 | S100A9 | 4836 | 0.698 | 0.3599 | No | ||
| 59 | CD14 | 4878 | 0.691 | 0.3623 | No | ||
| 60 | TCERG1 | 5035 | 0.663 | 0.3582 | No | ||
| 61 | RHOH | 5105 | 0.650 | 0.3589 | No | ||
| 62 | MARCO | 5199 | 0.634 | 0.3581 | No | ||
| 63 | ETS1 | 5492 | 0.581 | 0.3461 | No | ||
| 64 | IL5RA | 5519 | 0.577 | 0.3486 | No | ||
| 65 | GPR137B | 5561 | 0.572 | 0.3502 | No | ||
| 66 | SLC16A6 | 6222 | 0.467 | 0.3176 | No | ||
| 67 | LCP2 | 6243 | 0.463 | 0.3196 | No | ||
| 68 | UTS2 | 6264 | 0.460 | 0.3216 | No | ||
| 69 | MS4A2 | 6476 | 0.432 | 0.3130 | No | ||
| 70 | RAB3C | 6588 | 0.413 | 0.3098 | No | ||
| 71 | C5AR1 | 6959 | 0.358 | 0.2922 | No | ||
| 72 | TEX15 | 6982 | 0.355 | 0.2933 | No | ||
| 73 | SLC26A11 | 7108 | 0.338 | 0.2888 | No | ||
| 74 | SLC16A7 | 7133 | 0.335 | 0.2898 | No | ||
| 75 | SFRS6 | 7260 | 0.321 | 0.2851 | No | ||
| 76 | GJA4 | 7558 | 0.277 | 0.2709 | No | ||
| 77 | MMP9 | 8168 | 0.179 | 0.2391 | No | ||
| 78 | ADORA3 | 8237 | 0.169 | 0.2365 | No | ||
| 79 | PPBP | 8334 | 0.155 | 0.2324 | No | ||
| 80 | PNPLA1 | 8398 | 0.145 | 0.2299 | No | ||
| 81 | KLF2 | 8432 | 0.141 | 0.2291 | No | ||
| 82 | ANKRD29 | 8614 | 0.115 | 0.2201 | No | ||
| 83 | CST7 | 8740 | 0.093 | 0.2139 | No | ||
| 84 | PLA2G7 | 8805 | 0.083 | 0.2110 | No | ||
| 85 | IL3RA | 8831 | 0.079 | 0.2102 | No | ||
| 86 | ABI1 | 8862 | 0.072 | 0.2090 | No | ||
| 87 | CD200R1 | 9152 | 0.021 | 0.1935 | No | ||
| 88 | STAC | 9247 | 0.008 | 0.1885 | No | ||
| 89 | DOCK4 | 9394 | -0.016 | 0.1807 | No | ||
| 90 | HTRA1 | 9410 | -0.018 | 0.1800 | No | ||
| 91 | HRH4 | 9413 | -0.019 | 0.1800 | No | ||
| 92 | NDRG2 | 9493 | -0.030 | 0.1760 | No | ||
| 93 | EPAS1 | 9684 | -0.058 | 0.1661 | No | ||
| 94 | ADAM9 | 9751 | -0.066 | 0.1629 | No | ||
| 95 | ST6GALNAC5 | 9913 | -0.092 | 0.1548 | No | ||
| 96 | ENTPD1 | 9958 | -0.099 | 0.1531 | No | ||
| 97 | AMFR | 10085 | -0.121 | 0.1471 | No | ||
| 98 | ALDH2 | 10095 | -0.123 | 0.1474 | No | ||
| 99 | CLEC4A | 10149 | -0.132 | 0.1455 | No | ||
| 100 | RNASE1 | 10323 | -0.160 | 0.1372 | No | ||
| 101 | CLEC5A | 10393 | -0.172 | 0.1346 | No | ||
| 102 | CAMK1D | 10688 | -0.217 | 0.1201 | No | ||
| 103 | C7ORF10 | 10707 | -0.220 | 0.1206 | No | ||
| 104 | CREB5 | 11005 | -0.260 | 0.1063 | No | ||
| 105 | CHI3L1 | 11140 | -0.281 | 0.1009 | No | ||
| 106 | SLPI | 11813 | -0.390 | 0.0671 | No | ||
| 107 | GPR97 | 11831 | -0.393 | 0.0688 | No | ||
| 108 | HSD11B1 | 12079 | -0.433 | 0.0583 | No | ||
| 109 | ADAM19 | 12177 | -0.448 | 0.0561 | No | ||
| 110 | NKTR | 12554 | -0.510 | 0.0391 | No | ||
| 111 | ST8SIA1 | 12720 | -0.541 | 0.0338 | No | ||
| 112 | PADI4 | 12738 | -0.545 | 0.0365 | No | ||
| 113 | CD163 | 12762 | -0.549 | 0.0390 | No | ||
| 114 | COLEC12 | 12768 | -0.550 | 0.0424 | No | ||
| 115 | IL7R | 12957 | -0.577 | 0.0360 | No | ||
| 116 | CXCL1 | 13440 | -0.662 | 0.0144 | No | ||
| 117 | RAPH1 | 13456 | -0.665 | 0.0180 | No | ||
| 118 | PCSK5 | 13609 | -0.694 | 0.0144 | No | ||
| 119 | ITGAM | 13674 | -0.706 | 0.0157 | No | ||
| 120 | DACH1 | 13800 | -0.729 | 0.0138 | No | ||
| 121 | TDO2 | 14323 | -0.827 | -0.0090 | No | ||
| 122 | PDK4 | 14585 | -0.875 | -0.0172 | No | ||
| 123 | LGALS12 | 14623 | -0.881 | -0.0134 | No | ||
| 124 | ATP10D | 14669 | -0.891 | -0.0098 | No | ||
| 125 | SRPX | 14739 | -0.906 | -0.0075 | No | ||
| 126 | ARHGAP18 | 14748 | -0.908 | -0.0019 | No | ||
| 127 | SLC41A2 | 15089 | -0.984 | -0.0137 | No | ||
| 128 | TSPAN15 | 15681 | -1.125 | -0.0382 | No | ||
| 129 | NEK1 | 15785 | -1.154 | -0.0361 | No | ||
| 130 | PPP1R1B | 15831 | -1.165 | -0.0307 | No | ||
| 131 | CORO2A | 15915 | -1.192 | -0.0272 | No | ||
| 132 | S100B | 16007 | -1.213 | -0.0241 | No | ||
| 133 | P2RY10 | 16540 | -1.374 | -0.0437 | No | ||
| 134 | OLIG1 | 16738 | -1.444 | -0.0447 | No | ||
| 135 | PAX2 | 16835 | -1.479 | -0.0400 | No | ||
| 136 | TSPAN2 | 16930 | -1.515 | -0.0350 | No | ||
| 137 | CAPZA2 | 16971 | -1.529 | -0.0269 | No | ||
| 138 | ITM2A | 16996 | -1.540 | -0.0179 | No | ||
| 139 | CDC42 | 17148 | -1.605 | -0.0153 | No | ||
| 140 | NPL | 17171 | -1.614 | -0.0057 | No | ||
| 141 | FPR1 | 17308 | -1.669 | -0.0020 | No | ||
| 142 | TM6SF1 | 17420 | -1.714 | 0.0035 | No | ||
| 143 | AP1S2 | 17853 | -1.959 | -0.0068 | No | ||
| 144 | RPS6KA2 | 17903 | -1.996 | 0.0039 | No | ||
| 145 | BCL2 | 18004 | -2.077 | 0.0123 | No | ||
| 146 | PADI2 | 18542 | -3.102 | 0.0040 | No |