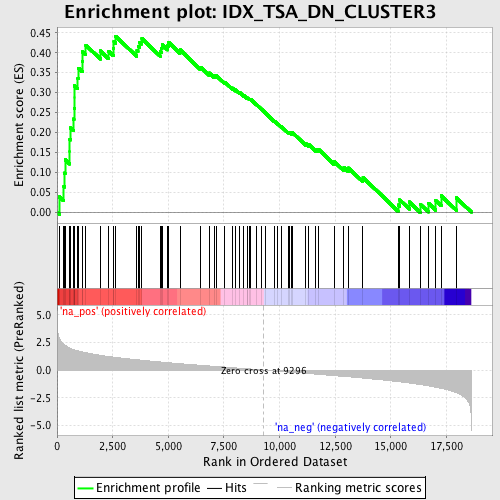
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | set03_absentNotch_versus_wtNotch |
| Phenotype | NoPhenotypeAvailable |
| Upregulated in class | na_pos |
| GeneSet | IDX_TSA_DN_CLUSTER3 |
| Enrichment Score (ES) | 0.4418739 |
| Normalized Enrichment Score (NES) | 1.8944687 |
| Nominal p-value | 0.0 |
| FDR q-value | 0.09701317 |
| FWER p-Value | 0.596 |

| PROBE | GENE SYMBOL | GENE_TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|---|
| 1 | SLC8A1 | 110 | 2.843 | 0.0384 | Yes | ||
| 2 | VCAM1 | 302 | 2.331 | 0.0645 | Yes | ||
| 3 | KLF4 | 335 | 2.277 | 0.0983 | Yes | ||
| 4 | DDEF1 | 362 | 2.232 | 0.1317 | Yes | ||
| 5 | RASA4 | 569 | 2.001 | 0.1518 | Yes | ||
| 6 | CDK4 | 572 | 1.997 | 0.1829 | Yes | ||
| 7 | IFI30 | 608 | 1.963 | 0.2116 | Yes | ||
| 8 | EXOC4 | 733 | 1.858 | 0.2339 | Yes | ||
| 9 | ARMCX2 | 762 | 1.841 | 0.2611 | Yes | ||
| 10 | EIF4B | 780 | 1.829 | 0.2887 | Yes | ||
| 11 | SPP1 | 781 | 1.828 | 0.3173 | Yes | ||
| 12 | CSF1 | 932 | 1.733 | 0.3362 | Yes | ||
| 13 | STXBP3 | 960 | 1.718 | 0.3615 | Yes | ||
| 14 | KIFAP3 | 1134 | 1.634 | 0.3777 | Yes | ||
| 15 | THBS2 | 1147 | 1.630 | 0.4025 | Yes | ||
| 16 | HEBP1 | 1291 | 1.573 | 0.4193 | Yes | ||
| 17 | NEDD9 | 1945 | 1.334 | 0.4049 | Yes | ||
| 18 | PLAT | 2323 | 1.231 | 0.4038 | Yes | ||
| 19 | EPHX1 | 2528 | 1.174 | 0.4111 | Yes | ||
| 20 | CITED2 | 2554 | 1.168 | 0.4280 | Yes | ||
| 21 | CDKN2B | 2631 | 1.152 | 0.4419 | Yes | ||
| 22 | CYR61 | 3587 | 0.939 | 0.4050 | No | ||
| 23 | TPD52L1 | 3637 | 0.930 | 0.4169 | No | ||
| 24 | CEBPD | 3715 | 0.912 | 0.4270 | No | ||
| 25 | RNF13 | 3811 | 0.892 | 0.4358 | No | ||
| 26 | CXCL12 | 4635 | 0.731 | 0.4028 | No | ||
| 27 | JUP | 4687 | 0.724 | 0.4113 | No | ||
| 28 | CNN3 | 4717 | 0.717 | 0.4210 | No | ||
| 29 | RASSF5 | 4977 | 0.672 | 0.4175 | No | ||
| 30 | AKAP12 | 5011 | 0.666 | 0.4261 | No | ||
| 31 | SPG20 | 5524 | 0.576 | 0.4075 | No | ||
| 32 | NME4 | 6463 | 0.434 | 0.3637 | No | ||
| 33 | CDKN2A | 6837 | 0.377 | 0.3494 | No | ||
| 34 | DPP7 | 7057 | 0.345 | 0.3430 | No | ||
| 35 | PLEKHC1 | 7161 | 0.330 | 0.3426 | No | ||
| 36 | BCAR1 | 7545 | 0.278 | 0.3263 | No | ||
| 37 | ANXA11 | 7882 | 0.222 | 0.3116 | No | ||
| 38 | PTGS2 | 8026 | 0.202 | 0.3071 | No | ||
| 39 | PLK2 | 8208 | 0.174 | 0.3000 | No | ||
| 40 | SNX6 | 8384 | 0.147 | 0.2929 | No | ||
| 41 | F3 | 8535 | 0.126 | 0.2868 | No | ||
| 42 | PHLDA1 | 8576 | 0.121 | 0.2865 | No | ||
| 43 | PRKCD | 8655 | 0.109 | 0.2840 | No | ||
| 44 | RECK | 8700 | 0.101 | 0.2832 | No | ||
| 45 | CSRP1 | 8944 | 0.055 | 0.2710 | No | ||
| 46 | DDAH2 | 9195 | 0.016 | 0.2577 | No | ||
| 47 | NDRG4 | 9360 | -0.011 | 0.2491 | No | ||
| 48 | FADS1 | 9758 | -0.068 | 0.2287 | No | ||
| 49 | SKAP2 | 9903 | -0.090 | 0.2223 | No | ||
| 50 | FZD2 | 10087 | -0.122 | 0.2144 | No | ||
| 51 | GSTT1 | 10385 | -0.171 | 0.2010 | No | ||
| 52 | GALK2 | 10462 | -0.182 | 0.1998 | No | ||
| 53 | ACVR2A | 10541 | -0.193 | 0.1986 | No | ||
| 54 | SIRT2 | 10562 | -0.195 | 0.2005 | No | ||
| 55 | TPM1 | 11180 | -0.286 | 0.1717 | No | ||
| 56 | GEM | 11303 | -0.306 | 0.1699 | No | ||
| 57 | CCT3 | 11631 | -0.360 | 0.1579 | No | ||
| 58 | MAGED2 | 11733 | -0.376 | 0.1583 | No | ||
| 59 | PSCD3 | 12452 | -0.494 | 0.1273 | No | ||
| 60 | PLA2G4A | 12892 | -0.569 | 0.1125 | No | ||
| 61 | SFRP2 | 13097 | -0.601 | 0.1109 | No | ||
| 62 | DNM1 | 13748 | -0.720 | 0.0871 | No | ||
| 63 | GPC4 | 15330 | -1.040 | 0.0181 | No | ||
| 64 | DOK1 | 15402 | -1.057 | 0.0307 | No | ||
| 65 | GAS2 | 15834 | -1.167 | 0.0257 | No | ||
| 66 | ROCK2 | 16323 | -1.303 | 0.0197 | No | ||
| 67 | CNN2 | 16685 | -1.423 | 0.0224 | No | ||
| 68 | MAOA | 17017 | -1.553 | 0.0288 | No | ||
| 69 | GGTA1 | 17267 | -1.649 | 0.0411 | No | ||
| 70 | LATS2 | 17947 | -2.023 | 0.0361 | No |
