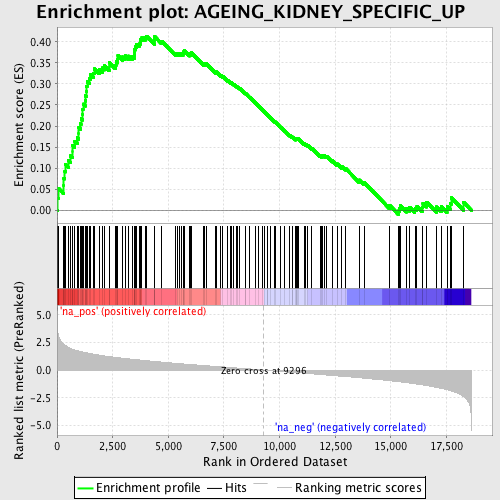
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | set03_absentNotch_versus_wtNotch |
| Phenotype | NoPhenotypeAvailable |
| Upregulated in class | na_pos |
| GeneSet | AGEING_KIDNEY_SPECIFIC_UP |
| Enrichment Score (ES) | 0.4135263 |
| Normalized Enrichment Score (NES) | 1.988917 |
| Nominal p-value | 0.0 |
| FDR q-value | 0.07352822 |
| FWER p-Value | 0.255 |

| PROBE | GENE SYMBOL | GENE_TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|---|
| 1 | SLC40A1 | 16 | 3.686 | 0.0289 | Yes | ||
| 2 | CYFIP1 | 49 | 3.147 | 0.0526 | Yes | ||
| 3 | AXL | 296 | 2.336 | 0.0581 | Yes | ||
| 4 | VCAM1 | 302 | 2.331 | 0.0766 | Yes | ||
| 5 | GGA2 | 348 | 2.259 | 0.0924 | Yes | ||
| 6 | CLDN1 | 378 | 2.213 | 0.1087 | Yes | ||
| 7 | ANTXR1 | 514 | 2.053 | 0.1180 | Yes | ||
| 8 | PRICKLE1 | 600 | 1.970 | 0.1293 | Yes | ||
| 9 | C1QC | 694 | 1.889 | 0.1395 | Yes | ||
| 10 | MGLL | 696 | 1.887 | 0.1547 | Yes | ||
| 11 | P4HB | 796 | 1.820 | 0.1640 | Yes | ||
| 12 | C1QB | 903 | 1.746 | 0.1724 | Yes | ||
| 13 | RPL35 | 971 | 1.714 | 0.1826 | Yes | ||
| 14 | TRIM47 | 975 | 1.713 | 0.1963 | Yes | ||
| 15 | NFIX | 1050 | 1.676 | 0.2058 | Yes | ||
| 16 | TPBG | 1078 | 1.665 | 0.2178 | Yes | ||
| 17 | TNC | 1146 | 1.631 | 0.2273 | Yes | ||
| 18 | SSR4 | 1154 | 1.627 | 0.2400 | Yes | ||
| 19 | GPNMB | 1169 | 1.622 | 0.2524 | Yes | ||
| 20 | CXCL14 | 1262 | 1.585 | 0.2602 | Yes | ||
| 21 | RPS25 | 1280 | 1.577 | 0.2720 | Yes | ||
| 22 | RBMS3 | 1315 | 1.559 | 0.2827 | Yes | ||
| 23 | FN1 | 1326 | 1.556 | 0.2948 | Yes | ||
| 24 | CALU | 1350 | 1.546 | 0.3060 | Yes | ||
| 25 | BICC1 | 1460 | 1.497 | 0.3122 | Yes | ||
| 26 | SCN5A | 1494 | 1.483 | 0.3224 | Yes | ||
| 27 | COL3A1 | 1642 | 1.426 | 0.3259 | Yes | ||
| 28 | GGTLA1 | 1670 | 1.417 | 0.3359 | Yes | ||
| 29 | LIX1 | 1895 | 1.349 | 0.3347 | Yes | ||
| 30 | SPON1 | 2027 | 1.306 | 0.3381 | Yes | ||
| 31 | BNC2 | 2122 | 1.280 | 0.3434 | Yes | ||
| 32 | SETD7 | 2343 | 1.225 | 0.3413 | Yes | ||
| 33 | RBP1 | 2346 | 1.224 | 0.3511 | Yes | ||
| 34 | PLOD3 | 2634 | 1.151 | 0.3449 | Yes | ||
| 35 | ABCC3 | 2650 | 1.147 | 0.3533 | Yes | ||
| 36 | OCLN | 2719 | 1.127 | 0.3587 | Yes | ||
| 37 | COL1A2 | 2733 | 1.124 | 0.3671 | Yes | ||
| 38 | PHCA | 2947 | 1.068 | 0.3642 | Yes | ||
| 39 | POLE4 | 3056 | 1.048 | 0.3668 | Yes | ||
| 40 | FCAMR | 3217 | 1.013 | 0.3663 | Yes | ||
| 41 | RGS1 | 3375 | 0.977 | 0.3657 | Yes | ||
| 42 | AEBP1 | 3466 | 0.961 | 0.3686 | Yes | ||
| 43 | NFKBIZ | 3481 | 0.959 | 0.3756 | Yes | ||
| 44 | FHL1 | 3490 | 0.957 | 0.3829 | Yes | ||
| 45 | TNFRSF11B | 3535 | 0.950 | 0.3882 | Yes | ||
| 46 | TFPI | 3558 | 0.945 | 0.3946 | Yes | ||
| 47 | RAD54L | 3687 | 0.918 | 0.3951 | Yes | ||
| 48 | LHFP | 3733 | 0.909 | 0.4000 | Yes | ||
| 49 | PIGR | 3747 | 0.906 | 0.4066 | Yes | ||
| 50 | SERPINE2 | 3812 | 0.892 | 0.4104 | Yes | ||
| 51 | CCL2 | 3954 | 0.862 | 0.4097 | Yes | ||
| 52 | CKAP4 | 4011 | 0.851 | 0.4135 | Yes | ||
| 53 | STAB1 | 4371 | 0.782 | 0.4004 | No | ||
| 54 | SLC1A3 | 4374 | 0.782 | 0.4066 | No | ||
| 55 | CHRNA3 | 4388 | 0.778 | 0.4122 | No | ||
| 56 | PGM2L1 | 4703 | 0.720 | 0.4010 | No | ||
| 57 | NNMT | 5316 | 0.613 | 0.3728 | No | ||
| 58 | DCDC2 | 5422 | 0.593 | 0.3719 | No | ||
| 59 | CFB | 5516 | 0.577 | 0.3716 | No | ||
| 60 | CDO1 | 5603 | 0.567 | 0.3715 | No | ||
| 61 | NR2F1 | 5661 | 0.557 | 0.3729 | No | ||
| 62 | COL6A3 | 5685 | 0.553 | 0.3761 | No | ||
| 63 | ANXA2 | 5713 | 0.549 | 0.3791 | No | ||
| 64 | LTF | 5952 | 0.508 | 0.3703 | No | ||
| 65 | PTRF | 5999 | 0.501 | 0.3719 | No | ||
| 66 | ITGB6 | 6031 | 0.496 | 0.3742 | No | ||
| 67 | COL1A1 | 6590 | 0.413 | 0.3474 | No | ||
| 68 | C3 | 6644 | 0.406 | 0.3478 | No | ||
| 69 | GPR61 | 6698 | 0.398 | 0.3481 | No | ||
| 70 | PDGFC | 7118 | 0.337 | 0.3281 | No | ||
| 71 | TIMP1 | 7157 | 0.331 | 0.3288 | No | ||
| 72 | RPS27L | 7356 | 0.306 | 0.3205 | No | ||
| 73 | AATF | 7454 | 0.293 | 0.3176 | No | ||
| 74 | SERPING1 | 7678 | 0.258 | 0.3077 | No | ||
| 75 | SYTL2 | 7798 | 0.237 | 0.3031 | No | ||
| 76 | HEXB | 7825 | 0.232 | 0.3036 | No | ||
| 77 | LOXL1 | 7941 | 0.213 | 0.2991 | No | ||
| 78 | PROM1 | 8063 | 0.195 | 0.2941 | No | ||
| 79 | EBF1 | 8126 | 0.185 | 0.2923 | No | ||
| 80 | PLK2 | 8208 | 0.174 | 0.2893 | No | ||
| 81 | COL5A1 | 8454 | 0.137 | 0.2771 | No | ||
| 82 | CXCL2 | 8471 | 0.134 | 0.2773 | No | ||
| 83 | CTHRC1 | 8657 | 0.109 | 0.2682 | No | ||
| 84 | RPL35A | 8921 | 0.062 | 0.2545 | No | ||
| 85 | VEGFC | 9066 | 0.035 | 0.2470 | No | ||
| 86 | ITM2C | 9214 | 0.013 | 0.2391 | No | ||
| 87 | PAM | 9316 | -0.003 | 0.2337 | No | ||
| 88 | MFGE8 | 9441 | -0.022 | 0.2272 | No | ||
| 89 | ADRA2A | 9459 | -0.025 | 0.2264 | No | ||
| 90 | CLDN3 | 9610 | -0.048 | 0.2187 | No | ||
| 91 | COL14A1 | 9783 | -0.073 | 0.2100 | No | ||
| 92 | MOXD1 | 9799 | -0.075 | 0.2098 | No | ||
| 93 | HOMER3 | 10045 | -0.114 | 0.1975 | No | ||
| 94 | SPARCL1 | 10239 | -0.147 | 0.1882 | No | ||
| 95 | COL27A1 | 10461 | -0.182 | 0.1777 | No | ||
| 96 | RAB34 | 10558 | -0.195 | 0.1741 | No | ||
| 97 | PHLDA3 | 10601 | -0.202 | 0.1734 | No | ||
| 98 | STS | 10726 | -0.222 | 0.1685 | No | ||
| 99 | TSPAN1 | 10741 | -0.225 | 0.1696 | No | ||
| 100 | NOV | 10775 | -0.230 | 0.1697 | No | ||
| 101 | RPN1 | 10823 | -0.237 | 0.1690 | No | ||
| 102 | PKIB | 10831 | -0.238 | 0.1706 | No | ||
| 103 | DPP3 | 11105 | -0.276 | 0.1580 | No | ||
| 104 | TPM1 | 11180 | -0.286 | 0.1563 | No | ||
| 105 | LUM | 11257 | -0.299 | 0.1546 | No | ||
| 106 | GPR56 | 11455 | -0.332 | 0.1467 | No | ||
| 107 | BHLHB3 | 11818 | -0.390 | 0.1302 | No | ||
| 108 | MMP11 | 11905 | -0.406 | 0.1288 | No | ||
| 109 | UBD | 11928 | -0.409 | 0.1309 | No | ||
| 110 | FBN1 | 12016 | -0.422 | 0.1296 | No | ||
| 111 | GABRP | 12102 | -0.438 | 0.1286 | No | ||
| 112 | B3GAT3 | 12363 | -0.481 | 0.1184 | No | ||
| 113 | C1S | 12595 | -0.520 | 0.1101 | No | ||
| 114 | CDC42EP1 | 12792 | -0.553 | 0.1040 | No | ||
| 115 | VSNL1 | 12951 | -0.577 | 0.1001 | No | ||
| 116 | SMPD2 | 13574 | -0.688 | 0.0720 | No | ||
| 117 | ARHGAP1 | 13795 | -0.728 | 0.0659 | No | ||
| 118 | RARRES1 | 14939 | -0.951 | 0.0118 | No | ||
| 119 | MMP7 | 15342 | -1.043 | -0.0016 | No | ||
| 120 | C1R | 15379 | -1.051 | 0.0050 | No | ||
| 121 | FEZ2 | 15428 | -1.064 | 0.0110 | No | ||
| 122 | ANXA1 | 15710 | -1.132 | 0.0049 | No | ||
| 123 | MRAS | 15841 | -1.169 | 0.0073 | No | ||
| 124 | ASNS | 16086 | -1.234 | 0.0040 | No | ||
| 125 | DEGS1 | 16172 | -1.257 | 0.0096 | No | ||
| 126 | COL15A1 | 16410 | -1.327 | 0.0075 | No | ||
| 127 | ITPR3 | 16442 | -1.342 | 0.0166 | No | ||
| 128 | MSR1 | 16609 | -1.394 | 0.0189 | No | ||
| 129 | HGF | 17043 | -1.561 | 0.0081 | No | ||
| 130 | GGTA1 | 17267 | -1.649 | 0.0093 | No | ||
| 131 | SOCS3 | 17528 | -1.766 | 0.0095 | No | ||
| 132 | FGG | 17687 | -1.859 | 0.0160 | No | ||
| 133 | ISLR | 17713 | -1.873 | 0.0297 | No | ||
| 134 | CDH11 | 18262 | -2.363 | 0.0192 | No |
