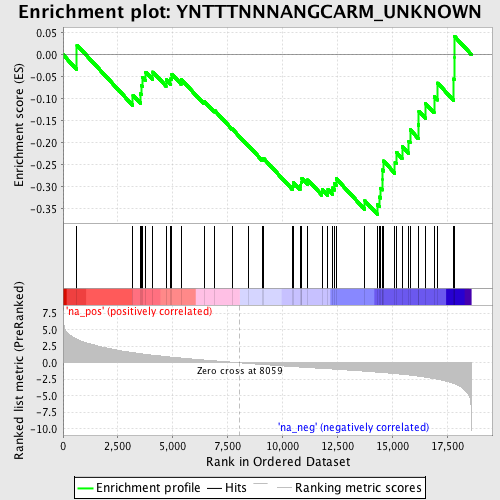
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | set03_absentNotch_versus_truncNotch |
| Phenotype | NoPhenotypeAvailable |
| Upregulated in class | na_neg |
| GeneSet | YNTTTNNNANGCARM_UNKNOWN |
| Enrichment Score (ES) | -0.36286724 |
| Normalized Enrichment Score (NES) | -1.3922567 |
| Nominal p-value | 0.052529182 |
| FDR q-value | 0.58415407 |
| FWER p-Value | 1.0 |

| PROBE | GENE SYMBOL | GENE_TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|---|
| 1 | CNN3 | 630 | 3.581 | 0.0220 | No | ||
| 2 | NRF1 | 3183 | 1.530 | -0.0916 | No | ||
| 3 | KCTD12 | 3528 | 1.374 | -0.0887 | No | ||
| 4 | PDE4D | 3575 | 1.357 | -0.0700 | No | ||
| 5 | KCNA2 | 3613 | 1.336 | -0.0511 | No | ||
| 6 | FGD4 | 3768 | 1.270 | -0.0396 | No | ||
| 7 | GNAI1 | 4092 | 1.143 | -0.0391 | No | ||
| 8 | TLL2 | 4693 | 0.935 | -0.0568 | No | ||
| 9 | DMD | 4909 | 0.863 | -0.0549 | No | ||
| 10 | GABRG1 | 4953 | 0.850 | -0.0440 | No | ||
| 11 | EVX1 | 5377 | 0.718 | -0.0555 | No | ||
| 12 | SFRP2 | 6422 | 0.419 | -0.1052 | No | ||
| 13 | OMG | 6886 | 0.294 | -0.1256 | No | ||
| 14 | B3GNT7 | 7705 | 0.089 | -0.1682 | No | ||
| 15 | ADAMTS5 | 8444 | -0.100 | -0.2064 | No | ||
| 16 | SMPX | 9076 | -0.253 | -0.2364 | No | ||
| 17 | NRK | 9137 | -0.266 | -0.2355 | No | ||
| 18 | KIF13A | 10467 | -0.569 | -0.2982 | No | ||
| 19 | DNAH5 | 10486 | -0.572 | -0.2903 | No | ||
| 20 | MAFB | 10801 | -0.643 | -0.2971 | No | ||
| 21 | GPM6A | 10842 | -0.651 | -0.2891 | No | ||
| 22 | C1S | 10879 | -0.659 | -0.2808 | No | ||
| 23 | HOXC4 | 11153 | -0.714 | -0.2843 | No | ||
| 24 | HCRTR2 | 11799 | -0.847 | -0.3058 | No | ||
| 25 | SYNPR | 12069 | -0.900 | -0.3063 | No | ||
| 26 | GABRA1 | 12273 | -0.943 | -0.3025 | No | ||
| 27 | FGF14 | 12367 | -0.964 | -0.2924 | No | ||
| 28 | LRRTM1 | 12448 | -0.979 | -0.2815 | No | ||
| 29 | HOXC5 | 13754 | -1.268 | -0.3320 | No | ||
| 30 | KLHL5 | 14329 | -1.412 | -0.3408 | Yes | ||
| 31 | EYA1 | 14416 | -1.435 | -0.3231 | Yes | ||
| 32 | DCX | 14465 | -1.447 | -0.3031 | Yes | ||
| 33 | GRK5 | 14544 | -1.469 | -0.2843 | Yes | ||
| 34 | GCDH | 14548 | -1.471 | -0.2615 | Yes | ||
| 35 | ELAVL2 | 14602 | -1.483 | -0.2412 | Yes | ||
| 36 | PVRL1 | 15125 | -1.630 | -0.2439 | Yes | ||
| 37 | HOXB6 | 15192 | -1.651 | -0.2217 | Yes | ||
| 38 | MSX2 | 15451 | -1.731 | -0.2085 | Yes | ||
| 39 | CDX2 | 15756 | -1.837 | -0.1962 | Yes | ||
| 40 | VDR | 15817 | -1.865 | -0.1704 | Yes | ||
| 41 | LEMD2 | 16188 | -2.022 | -0.1587 | Yes | ||
| 42 | FGF12 | 16216 | -2.030 | -0.1285 | Yes | ||
| 43 | GPR158 | 16526 | -2.172 | -0.1112 | Yes | ||
| 44 | VAX1 | 16909 | -2.385 | -0.0946 | Yes | ||
| 45 | LDB2 | 17065 | -2.486 | -0.0641 | Yes | ||
| 46 | IL10 | 17806 | -3.137 | -0.0550 | Yes | ||
| 47 | ETV5 | 17825 | -3.153 | -0.0067 | Yes | ||
| 48 | ETV6 | 17829 | -3.156 | 0.0424 | Yes |
