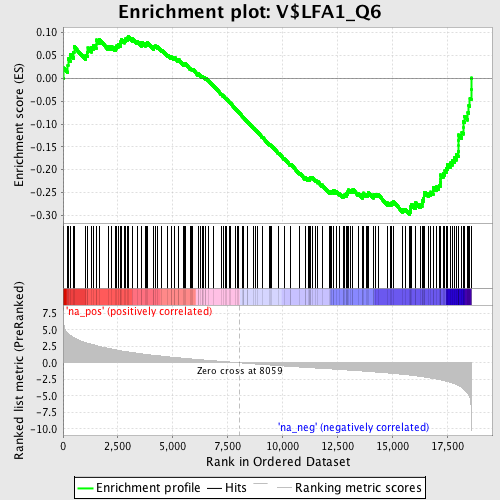
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | set03_absentNotch_versus_truncNotch |
| Phenotype | NoPhenotypeAvailable |
| Upregulated in class | na_neg |
| GeneSet | V$LFA1_Q6 |
| Enrichment Score (ES) | -0.29798457 |
| Normalized Enrichment Score (NES) | -1.4178932 |
| Nominal p-value | 0.0074074073 |
| FDR q-value | 0.5214564 |
| FWER p-Value | 1.0 |

| PROBE | GENE SYMBOL | GENE_TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|---|
| 1 | M6PR | 12 | 6.683 | 0.0228 | No | ||
| 2 | RBM12 | 219 | 4.466 | 0.0274 | No | ||
| 3 | BCL6 | 225 | 4.442 | 0.0427 | No | ||
| 4 | BCL11B | 324 | 4.156 | 0.0520 | No | ||
| 5 | CXCR4 | 479 | 3.818 | 0.0570 | No | ||
| 6 | UBL3 | 499 | 3.780 | 0.0693 | No | ||
| 7 | GSK3B | 1038 | 3.046 | 0.0508 | No | ||
| 8 | ARID1A | 1116 | 2.949 | 0.0570 | No | ||
| 9 | IMMT | 1133 | 2.934 | 0.0664 | No | ||
| 10 | TCF7 | 1301 | 2.788 | 0.0672 | No | ||
| 11 | CCNL1 | 1384 | 2.713 | 0.0723 | No | ||
| 12 | NXF1 | 1505 | 2.591 | 0.0749 | No | ||
| 13 | GNAO1 | 1507 | 2.590 | 0.0839 | No | ||
| 14 | EMILIN1 | 1654 | 2.468 | 0.0847 | No | ||
| 15 | CKB | 2061 | 2.181 | 0.0703 | No | ||
| 16 | RBBP6 | 2208 | 2.077 | 0.0697 | No | ||
| 17 | BRPF1 | 2378 | 1.962 | 0.0674 | No | ||
| 18 | NYX | 2441 | 1.928 | 0.0708 | No | ||
| 19 | NACA | 2505 | 1.889 | 0.0741 | No | ||
| 20 | RAB5C | 2603 | 1.843 | 0.0753 | No | ||
| 21 | FLI1 | 2632 | 1.817 | 0.0801 | No | ||
| 22 | ANXA4 | 2649 | 1.807 | 0.0856 | No | ||
| 23 | CLTC | 2805 | 1.715 | 0.0832 | No | ||
| 24 | LRP8 | 2860 | 1.683 | 0.0862 | No | ||
| 25 | UPF2 | 2929 | 1.648 | 0.0883 | No | ||
| 26 | FES | 2962 | 1.634 | 0.0923 | No | ||
| 27 | GAD1 | 3169 | 1.537 | 0.0866 | No | ||
| 28 | ETV1 | 3371 | 1.448 | 0.0808 | No | ||
| 29 | PCBP2 | 3566 | 1.361 | 0.0750 | No | ||
| 30 | DHH | 3580 | 1.355 | 0.0791 | No | ||
| 31 | THRAP1 | 3741 | 1.280 | 0.0749 | No | ||
| 32 | EDG1 | 3797 | 1.256 | 0.0763 | No | ||
| 33 | AKT3 | 3842 | 1.238 | 0.0783 | No | ||
| 34 | CPNE1 | 4128 | 1.129 | 0.0668 | No | ||
| 35 | CRMP1 | 4134 | 1.128 | 0.0705 | No | ||
| 36 | PYGO2 | 4197 | 1.107 | 0.0710 | No | ||
| 37 | NPC2 | 4298 | 1.070 | 0.0694 | No | ||
| 38 | PRRX1 | 4501 | 0.999 | 0.0619 | No | ||
| 39 | NFAT5 | 4779 | 0.904 | 0.0501 | No | ||
| 40 | PAX7 | 4920 | 0.860 | 0.0455 | No | ||
| 41 | TSP50 | 4950 | 0.852 | 0.0469 | No | ||
| 42 | NPAS3 | 5063 | 0.815 | 0.0437 | No | ||
| 43 | IL1F10 | 5098 | 0.803 | 0.0447 | No | ||
| 44 | SLC2A4 | 5248 | 0.761 | 0.0393 | No | ||
| 45 | NFKBIA | 5250 | 0.761 | 0.0419 | No | ||
| 46 | NKX2-3 | 5504 | 0.679 | 0.0306 | No | ||
| 47 | MGEA5 | 5536 | 0.669 | 0.0312 | No | ||
| 48 | CACNB1 | 5567 | 0.657 | 0.0319 | No | ||
| 49 | DLX1 | 5811 | 0.592 | 0.0208 | No | ||
| 50 | MYOG | 5862 | 0.578 | 0.0201 | No | ||
| 51 | BNC2 | 5900 | 0.569 | 0.0201 | No | ||
| 52 | SMARCD2 | 6149 | 0.494 | 0.0084 | No | ||
| 53 | PHF12 | 6151 | 0.493 | 0.0101 | No | ||
| 54 | LZTS2 | 6254 | 0.466 | 0.0062 | No | ||
| 55 | ADM | 6332 | 0.444 | 0.0036 | No | ||
| 56 | NPTXR | 6373 | 0.431 | 0.0029 | No | ||
| 57 | LIN28 | 6405 | 0.424 | 0.0027 | No | ||
| 58 | CHX10 | 6502 | 0.400 | -0.0011 | No | ||
| 59 | ACVR1C | 6508 | 0.398 | 0.0001 | No | ||
| 60 | NGFB | 6606 | 0.373 | -0.0039 | No | ||
| 61 | PPY | 6833 | 0.311 | -0.0150 | No | ||
| 62 | CASQ2 | 7223 | 0.209 | -0.0354 | No | ||
| 63 | NEURL | 7291 | 0.194 | -0.0384 | No | ||
| 64 | PLA2G10 | 7413 | 0.163 | -0.0443 | No | ||
| 65 | DUOX2 | 7465 | 0.149 | -0.0466 | No | ||
| 66 | FXYD3 | 7584 | 0.119 | -0.0526 | No | ||
| 67 | JPH4 | 7647 | 0.104 | -0.0556 | No | ||
| 68 | WNT4 | 7865 | 0.050 | -0.0671 | No | ||
| 69 | KLK9 | 7966 | 0.022 | -0.0725 | No | ||
| 70 | TAL1 | 7998 | 0.014 | -0.0741 | No | ||
| 71 | CTGF | 8162 | -0.024 | -0.0829 | No | ||
| 72 | HEBP1 | 8198 | -0.033 | -0.0847 | No | ||
| 73 | SP7 | 8240 | -0.044 | -0.0867 | No | ||
| 74 | OLFML2A | 8385 | -0.083 | -0.0942 | No | ||
| 75 | CAV1 | 8390 | -0.085 | -0.0942 | No | ||
| 76 | XYLT1 | 8683 | -0.156 | -0.1094 | No | ||
| 77 | PAK1IP1 | 8773 | -0.180 | -0.1136 | No | ||
| 78 | KCNQ4 | 8837 | -0.196 | -0.1164 | No | ||
| 79 | MYBPH | 9097 | -0.258 | -0.1295 | No | ||
| 80 | S100A5 | 9419 | -0.322 | -0.1458 | No | ||
| 81 | GABARAP | 9440 | -0.328 | -0.1457 | No | ||
| 82 | GPR4 | 9514 | -0.345 | -0.1485 | No | ||
| 83 | GPR3 | 9826 | -0.422 | -0.1638 | No | ||
| 84 | TBR1 | 10086 | -0.483 | -0.1762 | No | ||
| 85 | PNOC | 10361 | -0.544 | -0.1891 | No | ||
| 86 | NDUFS2 | 10380 | -0.549 | -0.1882 | No | ||
| 87 | CNKSR2 | 10778 | -0.636 | -0.2075 | No | ||
| 88 | CREB3L2 | 11031 | -0.687 | -0.2187 | No | ||
| 89 | LCAT | 11049 | -0.692 | -0.2172 | No | ||
| 90 | DPF2 | 11174 | -0.719 | -0.2214 | No | ||
| 91 | PTRF | 11224 | -0.730 | -0.2215 | No | ||
| 92 | GRIN2A | 11231 | -0.732 | -0.2193 | No | ||
| 93 | ACCN1 | 11255 | -0.737 | -0.2179 | No | ||
| 94 | GBA2 | 11286 | -0.742 | -0.2169 | No | ||
| 95 | CEBPB | 11345 | -0.756 | -0.2174 | No | ||
| 96 | ZFPM1 | 11499 | -0.786 | -0.2230 | No | ||
| 97 | HNRPR | 11614 | -0.811 | -0.2263 | No | ||
| 98 | ZNF385 | 11801 | -0.847 | -0.2334 | No | ||
| 99 | HOXC10 | 12138 | -0.914 | -0.2484 | No | ||
| 100 | ZFP36 | 12200 | -0.925 | -0.2485 | No | ||
| 101 | OTOP2 | 12246 | -0.935 | -0.2476 | No | ||
| 102 | IFRD2 | 12316 | -0.951 | -0.2480 | No | ||
| 103 | RFX4 | 12334 | -0.956 | -0.2456 | No | ||
| 104 | BOC | 12445 | -0.978 | -0.2481 | No | ||
| 105 | MRPL40 | 12586 | -1.010 | -0.2521 | No | ||
| 106 | FLT1 | 12764 | -1.044 | -0.2581 | No | ||
| 107 | SIX4 | 12786 | -1.048 | -0.2555 | No | ||
| 108 | NR4A1 | 12844 | -1.060 | -0.2549 | No | ||
| 109 | NR5A2 | 12904 | -1.072 | -0.2543 | No | ||
| 110 | PPP1R16A | 12915 | -1.074 | -0.2511 | No | ||
| 111 | KCNAB1 | 12968 | -1.086 | -0.2501 | No | ||
| 112 | POU4F1 | 12980 | -1.089 | -0.2469 | No | ||
| 113 | LAG3 | 13003 | -1.093 | -0.2442 | No | ||
| 114 | SLC7A10 | 13111 | -1.114 | -0.2461 | No | ||
| 115 | MSX1 | 13198 | -1.135 | -0.2468 | No | ||
| 116 | ARID4A | 13208 | -1.137 | -0.2433 | No | ||
| 117 | LMOD3 | 13448 | -1.195 | -0.2520 | No | ||
| 118 | USH1G | 13638 | -1.240 | -0.2579 | No | ||
| 119 | ADAMTS4 | 13657 | -1.245 | -0.2545 | No | ||
| 120 | TNRC4 | 13673 | -1.250 | -0.2510 | No | ||
| 121 | STOML2 | 13809 | -1.284 | -0.2538 | No | ||
| 122 | EPO | 13893 | -1.304 | -0.2537 | No | ||
| 123 | NRGN | 13912 | -1.308 | -0.2501 | No | ||
| 124 | RHO | 14135 | -1.360 | -0.2573 | No | ||
| 125 | SMAD3 | 14165 | -1.369 | -0.2541 | No | ||
| 126 | SLC26A9 | 14250 | -1.390 | -0.2538 | No | ||
| 127 | KIF1C | 14359 | -1.421 | -0.2546 | No | ||
| 128 | SLC13A5 | 14790 | -1.530 | -0.2726 | No | ||
| 129 | SOX5 | 14917 | -1.569 | -0.2739 | No | ||
| 130 | FKBP2 | 14987 | -1.588 | -0.2720 | No | ||
| 131 | SLC25A29 | 15053 | -1.608 | -0.2699 | No | ||
| 132 | ELF4 | 15483 | -1.739 | -0.2871 | No | ||
| 133 | BAX | 15583 | -1.770 | -0.2862 | No | ||
| 134 | HOXC6 | 15801 | -1.856 | -0.2915 | Yes | ||
| 135 | CIC | 15841 | -1.875 | -0.2870 | Yes | ||
| 136 | SALL1 | 15849 | -1.878 | -0.2808 | Yes | ||
| 137 | DLGAP4 | 15892 | -1.892 | -0.2764 | Yes | ||
| 138 | RABGGTA | 16058 | -1.968 | -0.2784 | Yes | ||
| 139 | CACNA1G | 16075 | -1.972 | -0.2724 | Yes | ||
| 140 | OBSCN | 16282 | -2.057 | -0.2763 | Yes | ||
| 141 | PDYN | 16361 | -2.096 | -0.2732 | Yes | ||
| 142 | CLIC5 | 16389 | -2.113 | -0.2672 | Yes | ||
| 143 | BACE1 | 16431 | -2.127 | -0.2620 | Yes | ||
| 144 | SH3BP1 | 16448 | -2.133 | -0.2553 | Yes | ||
| 145 | EPHB2 | 16469 | -2.145 | -0.2489 | Yes | ||
| 146 | MCRS1 | 16656 | -2.238 | -0.2511 | Yes | ||
| 147 | MNT | 16762 | -2.299 | -0.2487 | Yes | ||
| 148 | TRIP10 | 16864 | -2.359 | -0.2459 | Yes | ||
| 149 | GABRG2 | 16877 | -2.367 | -0.2383 | Yes | ||
| 150 | UQCRC1 | 17024 | -2.462 | -0.2375 | Yes | ||
| 151 | MAZ | 17132 | -2.534 | -0.2344 | Yes | ||
| 152 | PLEC1 | 17186 | -2.570 | -0.2283 | Yes | ||
| 153 | CNN2 | 17208 | -2.594 | -0.2203 | Yes | ||
| 154 | SEMA6A | 17213 | -2.598 | -0.2114 | Yes | ||
| 155 | EFNA3 | 17319 | -2.680 | -0.2077 | Yes | ||
| 156 | DCTN1 | 17402 | -2.738 | -0.2025 | Yes | ||
| 157 | NEUROG1 | 17478 | -2.802 | -0.1967 | Yes | ||
| 158 | TITF1 | 17524 | -2.839 | -0.1892 | Yes | ||
| 159 | HIRA | 17643 | -2.964 | -0.1851 | Yes | ||
| 160 | COX7A2L | 17731 | -3.053 | -0.1791 | Yes | ||
| 161 | RAB4B | 17841 | -3.173 | -0.1739 | Yes | ||
| 162 | VASP | 17934 | -3.311 | -0.1673 | Yes | ||
| 163 | NCDN | 18013 | -3.437 | -0.1594 | Yes | ||
| 164 | SESN3 | 18025 | -3.456 | -0.1479 | Yes | ||
| 165 | PDLIM7 | 18028 | -3.461 | -0.1358 | Yes | ||
| 166 | RASGRP2 | 18034 | -3.467 | -0.1239 | Yes | ||
| 167 | GSS | 18177 | -3.726 | -0.1185 | Yes | ||
| 168 | PALM | 18242 | -3.899 | -0.1083 | Yes | ||
| 169 | TPCN1 | 18247 | -3.909 | -0.0948 | Yes | ||
| 170 | ROM1 | 18302 | -4.073 | -0.0834 | Yes | ||
| 171 | HARS2 | 18450 | -4.653 | -0.0750 | Yes | ||
| 172 | POLD4 | 18494 | -4.910 | -0.0601 | Yes | ||
| 173 | RHOG | 18544 | -5.301 | -0.0441 | Yes | ||
| 174 | HES1 | 18597 | -6.611 | -0.0237 | Yes | ||
| 175 | MOV10 | 18603 | -7.025 | 0.0007 | Yes |
