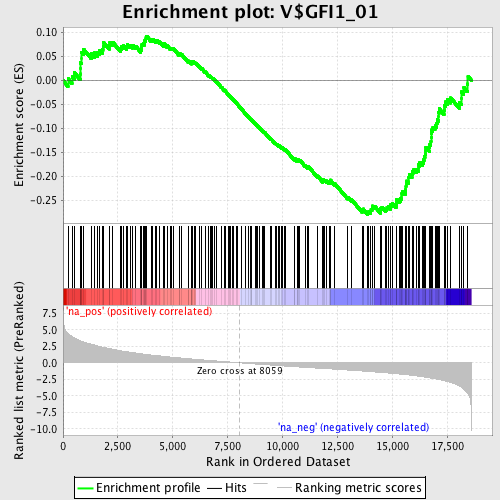
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | set03_absentNotch_versus_truncNotch |
| Phenotype | NoPhenotypeAvailable |
| Upregulated in class | na_neg |
| GeneSet | V$GFI1_01 |
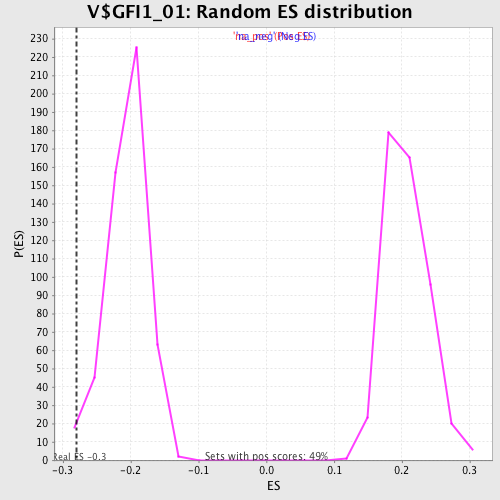
| Enrichment Score (ES) | -0.27932712 |
| Normalized Enrichment Score (NES) | -1.3596617 |
| Nominal p-value | 0.01764706 |
| FDR q-value | 0.55171466 |
| FWER p-Value | 1.0 |

| PROBE | GENE SYMBOL | GENE_TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|---|
| 1 | BCL6 | 225 | 4.442 | 0.0039 | No | ||
| 2 | SLC4A7 | 408 | 3.959 | 0.0084 | No | ||
| 3 | TLK1 | 510 | 3.763 | 0.0165 | No | ||
| 4 | BRD2 | 782 | 3.336 | 0.0139 | No | ||
| 5 | ITGA10 | 789 | 3.326 | 0.0256 | No | ||
| 6 | MBNL1 | 795 | 3.320 | 0.0374 | No | ||
| 7 | SFRS2 | 836 | 3.269 | 0.0471 | No | ||
| 8 | POLH | 839 | 3.265 | 0.0588 | No | ||
| 9 | HBP1 | 926 | 3.155 | 0.0656 | No | ||
| 10 | NCAM1 | 1286 | 2.800 | 0.0562 | No | ||
| 11 | HSD11B1 | 1431 | 2.668 | 0.0581 | No | ||
| 12 | YWHAE | 1571 | 2.534 | 0.0597 | No | ||
| 13 | TOP1 | 1664 | 2.462 | 0.0636 | No | ||
| 14 | TCF12 | 1786 | 2.371 | 0.0657 | No | ||
| 15 | STMN1 | 1820 | 2.349 | 0.0724 | No | ||
| 16 | PDC | 1839 | 2.328 | 0.0799 | No | ||
| 17 | ZIC3 | 2100 | 2.160 | 0.0736 | No | ||
| 18 | SOX4 | 2131 | 2.128 | 0.0797 | No | ||
| 19 | DNAJB4 | 2257 | 2.049 | 0.0803 | No | ||
| 20 | MYH10 | 2624 | 1.822 | 0.0670 | No | ||
| 21 | MCTS1 | 2673 | 1.792 | 0.0709 | No | ||
| 22 | GPR20 | 2754 | 1.740 | 0.0729 | No | ||
| 23 | CNKSR1 | 2904 | 1.664 | 0.0708 | No | ||
| 24 | TMPRSS2 | 2936 | 1.646 | 0.0751 | No | ||
| 25 | ATF2 | 3062 | 1.588 | 0.0741 | No | ||
| 26 | CDH13 | 3180 | 1.530 | 0.0733 | No | ||
| 27 | PRIMA1 | 3310 | 1.475 | 0.0716 | No | ||
| 28 | TRIM23 | 3542 | 1.369 | 0.0641 | No | ||
| 29 | PDE4D | 3575 | 1.357 | 0.0672 | No | ||
| 30 | ELMO3 | 3585 | 1.352 | 0.0717 | No | ||
| 31 | MAP4K3 | 3592 | 1.348 | 0.0762 | No | ||
| 32 | TCF4 | 3652 | 1.316 | 0.0778 | No | ||
| 33 | DSCR1L1 | 3722 | 1.287 | 0.0787 | No | ||
| 34 | ICAM5 | 3724 | 1.286 | 0.0833 | No | ||
| 35 | SLC25A18 | 3770 | 1.269 | 0.0855 | No | ||
| 36 | LUZP1 | 3776 | 1.268 | 0.0898 | No | ||
| 37 | TPBG | 3804 | 1.254 | 0.0929 | No | ||
| 38 | PPARGC1A | 4010 | 1.174 | 0.0860 | No | ||
| 39 | MYO1C | 4084 | 1.147 | 0.0862 | No | ||
| 40 | TFB1M | 4215 | 1.102 | 0.0831 | No | ||
| 41 | PPARGC1B | 4266 | 1.079 | 0.0843 | No | ||
| 42 | BAI3 | 4396 | 1.034 | 0.0811 | No | ||
| 43 | DEPDC2 | 4577 | 0.971 | 0.0748 | No | ||
| 44 | PANK1 | 4602 | 0.962 | 0.0770 | No | ||
| 45 | LAMA3 | 4741 | 0.917 | 0.0728 | No | ||
| 46 | NGEF | 4893 | 0.869 | 0.0678 | No | ||
| 47 | TFEC | 4951 | 0.852 | 0.0678 | No | ||
| 48 | PTEN | 5008 | 0.829 | 0.0678 | No | ||
| 49 | ABCF2 | 5283 | 0.749 | 0.0556 | No | ||
| 50 | XPO5 | 5323 | 0.737 | 0.0562 | No | ||
| 51 | SPAG7 | 5384 | 0.716 | 0.0555 | No | ||
| 52 | CSAD | 5726 | 0.614 | 0.0392 | No | ||
| 53 | CGN | 5733 | 0.611 | 0.0411 | No | ||
| 54 | PREP | 5855 | 0.581 | 0.0366 | No | ||
| 55 | CRYZL1 | 5861 | 0.578 | 0.0385 | No | ||
| 56 | SH3BGRL2 | 5863 | 0.577 | 0.0405 | No | ||
| 57 | BNC2 | 5900 | 0.569 | 0.0406 | No | ||
| 58 | HOXA7 | 5993 | 0.539 | 0.0376 | No | ||
| 59 | AVP | 6042 | 0.525 | 0.0369 | No | ||
| 60 | KLF15 | 6201 | 0.480 | 0.0300 | No | ||
| 61 | IRX3 | 6321 | 0.445 | 0.0252 | No | ||
| 62 | MEF2C | 6474 | 0.406 | 0.0184 | No | ||
| 63 | PEA15 | 6641 | 0.360 | 0.0107 | No | ||
| 64 | OTP | 6703 | 0.345 | 0.0086 | No | ||
| 65 | PBX1 | 6757 | 0.333 | 0.0070 | No | ||
| 66 | RARB | 6819 | 0.316 | 0.0048 | No | ||
| 67 | TRDN | 6878 | 0.296 | 0.0027 | No | ||
| 68 | GAS2 | 6969 | 0.273 | -0.0012 | No | ||
| 69 | MEIS1 | 7214 | 0.211 | -0.0136 | No | ||
| 70 | SGK2 | 7334 | 0.184 | -0.0194 | No | ||
| 71 | HOXD11 | 7379 | 0.172 | -0.0212 | No | ||
| 72 | TLR1 | 7547 | 0.129 | -0.0298 | No | ||
| 73 | CNTN6 | 7576 | 0.123 | -0.0309 | No | ||
| 74 | JPH4 | 7647 | 0.104 | -0.0343 | No | ||
| 75 | ZIC4 | 7739 | 0.081 | -0.0389 | No | ||
| 76 | SERTAD4 | 7784 | 0.073 | -0.0410 | No | ||
| 77 | EVA1 | 7882 | 0.046 | -0.0461 | No | ||
| 78 | PROCR | 7962 | 0.023 | -0.0503 | No | ||
| 79 | PHF8 | 8119 | -0.012 | -0.0588 | No | ||
| 80 | ZFPM2 | 8298 | -0.059 | -0.0682 | No | ||
| 81 | GRPR | 8438 | -0.097 | -0.0754 | No | ||
| 82 | CACNG3 | 8555 | -0.128 | -0.0812 | No | ||
| 83 | MSR1 | 8572 | -0.132 | -0.0816 | No | ||
| 84 | POU3F1 | 8587 | -0.135 | -0.0819 | No | ||
| 85 | BPGM | 8748 | -0.173 | -0.0900 | No | ||
| 86 | CDH6 | 8806 | -0.188 | -0.0924 | No | ||
| 87 | RKHD1 | 8869 | -0.203 | -0.0950 | No | ||
| 88 | NKX2-8 | 8945 | -0.219 | -0.0983 | No | ||
| 89 | EDN1 | 9106 | -0.260 | -0.1060 | No | ||
| 90 | HOXD10 | 9116 | -0.262 | -0.1056 | No | ||
| 91 | ITSN1 | 9186 | -0.277 | -0.1083 | No | ||
| 92 | GABARAP | 9440 | -0.328 | -0.1208 | No | ||
| 93 | MAP2K7 | 9488 | -0.337 | -0.1222 | No | ||
| 94 | CTLA4 | 9669 | -0.384 | -0.1306 | No | ||
| 95 | DSPP | 9732 | -0.398 | -0.1325 | No | ||
| 96 | PI15 | 9822 | -0.421 | -0.1358 | No | ||
| 97 | SREBF2 | 9854 | -0.427 | -0.1359 | No | ||
| 98 | PURA | 9948 | -0.451 | -0.1393 | No | ||
| 99 | HOXA6 | 9984 | -0.460 | -0.1396 | No | ||
| 100 | TBR1 | 10086 | -0.483 | -0.1433 | No | ||
| 101 | HNRPA0 | 10123 | -0.494 | -0.1435 | No | ||
| 102 | GADD45A | 10538 | -0.584 | -0.1638 | No | ||
| 103 | C1QTNF7 | 10544 | -0.585 | -0.1620 | No | ||
| 104 | GREM1 | 10662 | -0.612 | -0.1661 | No | ||
| 105 | SLC26A3 | 10680 | -0.615 | -0.1648 | No | ||
| 106 | BMP4 | 10728 | -0.625 | -0.1651 | No | ||
| 107 | HOXA3 | 10787 | -0.638 | -0.1659 | No | ||
| 108 | CDON | 11050 | -0.692 | -0.1776 | No | ||
| 109 | HOXC4 | 11153 | -0.714 | -0.1806 | No | ||
| 110 | DDR2 | 11184 | -0.722 | -0.1796 | No | ||
| 111 | CREM | 11594 | -0.807 | -0.1988 | No | ||
| 112 | IRF2BP1 | 11837 | -0.855 | -0.2089 | No | ||
| 113 | SOX15 | 11849 | -0.857 | -0.2064 | No | ||
| 114 | ZIC1 | 11927 | -0.874 | -0.2074 | No | ||
| 115 | SOX9 | 12020 | -0.892 | -0.2091 | No | ||
| 116 | RHOA | 12134 | -0.912 | -0.2120 | No | ||
| 117 | DSG3 | 12168 | -0.920 | -0.2104 | No | ||
| 118 | CHRM1 | 12174 | -0.921 | -0.2074 | No | ||
| 119 | MTMR4 | 12380 | -0.967 | -0.2150 | No | ||
| 120 | BCL9 | 12974 | -1.087 | -0.2432 | No | ||
| 121 | JMJD1C | 13139 | -1.120 | -0.2481 | No | ||
| 122 | CCR7 | 13626 | -1.238 | -0.2700 | No | ||
| 123 | TCF1 | 13671 | -1.250 | -0.2678 | No | ||
| 124 | PCDH8 | 13884 | -1.302 | -0.2746 | Yes | ||
| 125 | BDNF | 13934 | -1.313 | -0.2725 | Yes | ||
| 126 | HOXA9 | 14019 | -1.331 | -0.2722 | Yes | ||
| 127 | NFIA | 14031 | -1.334 | -0.2680 | Yes | ||
| 128 | PHYHIP | 14081 | -1.349 | -0.2658 | Yes | ||
| 129 | IL16 | 14094 | -1.352 | -0.2615 | Yes | ||
| 130 | EXOC7 | 14199 | -1.377 | -0.2622 | Yes | ||
| 131 | DCX | 14465 | -1.447 | -0.2713 | Yes | ||
| 132 | ETS1 | 14475 | -1.449 | -0.2666 | Yes | ||
| 133 | MAP3K3 | 14527 | -1.464 | -0.2640 | Yes | ||
| 134 | FOXP2 | 14704 | -1.508 | -0.2681 | Yes | ||
| 135 | PLCD4 | 14760 | -1.523 | -0.2656 | Yes | ||
| 136 | GFRA3 | 14807 | -1.534 | -0.2625 | Yes | ||
| 137 | SOX5 | 14917 | -1.569 | -0.2627 | Yes | ||
| 138 | SOSTDC1 | 14936 | -1.575 | -0.2580 | Yes | ||
| 139 | CPEB3 | 15022 | -1.598 | -0.2568 | Yes | ||
| 140 | RKHD3 | 15178 | -1.647 | -0.2593 | Yes | ||
| 141 | HOXB6 | 15192 | -1.651 | -0.2540 | Yes | ||
| 142 | TRAF4 | 15204 | -1.659 | -0.2486 | Yes | ||
| 143 | GPRC5D | 15319 | -1.692 | -0.2486 | Yes | ||
| 144 | ARHGEF2 | 15383 | -1.710 | -0.2459 | Yes | ||
| 145 | PHACTR3 | 15437 | -1.727 | -0.2425 | Yes | ||
| 146 | NR2F2 | 15439 | -1.727 | -0.2363 | Yes | ||
| 147 | NEXN | 15472 | -1.736 | -0.2317 | Yes | ||
| 148 | EBF2 | 15591 | -1.772 | -0.2317 | Yes | ||
| 149 | RAB6B | 15618 | -1.780 | -0.2266 | Yes | ||
| 150 | RORA | 15626 | -1.783 | -0.2206 | Yes | ||
| 151 | KLF12 | 15631 | -1.785 | -0.2143 | Yes | ||
| 152 | GRK6 | 15662 | -1.797 | -0.2094 | Yes | ||
| 153 | DPF3 | 15754 | -1.836 | -0.2077 | Yes | ||
| 154 | CDX2 | 15756 | -1.837 | -0.2011 | Yes | ||
| 155 | HIST1H1T | 15777 | -1.845 | -0.1955 | Yes | ||
| 156 | ASB4 | 15904 | -1.899 | -0.1955 | Yes | ||
| 157 | ICAM4 | 15923 | -1.906 | -0.1895 | Yes | ||
| 158 | HIST1H1E | 15986 | -1.932 | -0.1859 | Yes | ||
| 159 | PPP2R2A | 16113 | -1.987 | -0.1855 | Yes | ||
| 160 | HOXC8 | 16196 | -2.025 | -0.1826 | Yes | ||
| 161 | CALN1 | 16202 | -2.027 | -0.1756 | Yes | ||
| 162 | MAP4K2 | 16259 | -2.046 | -0.1712 | Yes | ||
| 163 | WDR23 | 16378 | -2.106 | -0.1700 | Yes | ||
| 164 | ATP5G2 | 16435 | -2.128 | -0.1653 | Yes | ||
| 165 | BTG1 | 16454 | -2.135 | -0.1585 | Yes | ||
| 166 | ELAVL4 | 16496 | -2.156 | -0.1529 | Yes | ||
| 167 | TSGA10 | 16523 | -2.169 | -0.1465 | Yes | ||
| 168 | ADD3 | 16532 | -2.177 | -0.1390 | Yes | ||
| 169 | HOXB8 | 16694 | -2.261 | -0.1396 | Yes | ||
| 170 | ANGPT1 | 16713 | -2.268 | -0.1323 | Yes | ||
| 171 | CREB5 | 16764 | -2.299 | -0.1267 | Yes | ||
| 172 | HIVEP3 | 16770 | -2.305 | -0.1186 | Yes | ||
| 173 | HOXB3 | 16779 | -2.314 | -0.1107 | Yes | ||
| 174 | MRGPRF | 16792 | -2.321 | -0.1029 | Yes | ||
| 175 | HOXA10 | 16854 | -2.351 | -0.0977 | Yes | ||
| 176 | SP8 | 16952 | -2.417 | -0.0942 | Yes | ||
| 177 | CLMN | 17029 | -2.467 | -0.0894 | Yes | ||
| 178 | RPS6KB1 | 17050 | -2.477 | -0.0815 | Yes | ||
| 179 | GPR17 | 17111 | -2.517 | -0.0756 | Yes | ||
| 180 | CACNB2 | 17116 | -2.522 | -0.0667 | Yes | ||
| 181 | CAMK2D | 17135 | -2.535 | -0.0585 | Yes | ||
| 182 | NRXN3 | 17364 | -2.712 | -0.0610 | Yes | ||
| 183 | PTHLH | 17380 | -2.723 | -0.0520 | Yes | ||
| 184 | PRKAG1 | 17424 | -2.756 | -0.0443 | Yes | ||
| 185 | PDLIM1 | 17523 | -2.838 | -0.0394 | Yes | ||
| 186 | TCTA | 17662 | -2.985 | -0.0360 | Yes | ||
| 187 | BCL9L | 18074 | -3.539 | -0.0455 | Yes | ||
| 188 | USP5 | 18153 | -3.687 | -0.0364 | Yes | ||
| 189 | BRUNOL4 | 18160 | -3.697 | -0.0233 | Yes | ||
| 190 | MAP2K5 | 18266 | -3.966 | -0.0146 | Yes | ||
| 191 | HMGA1 | 18432 | -4.595 | -0.0069 | Yes | ||
| 192 | AGPAT4 | 18452 | -4.657 | 0.0089 | Yes |