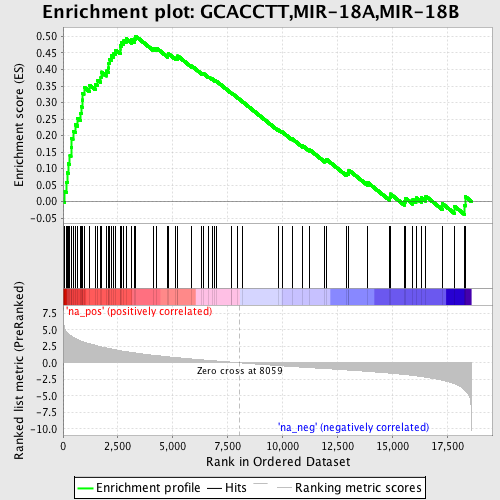
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | set03_absentNotch_versus_truncNotch |
| Phenotype | NoPhenotypeAvailable |
| Upregulated in class | na_pos |
| GeneSet | GCACCTT,MIR-18A,MIR-18B |
| Enrichment Score (ES) | 0.5010114 |
| Normalized Enrichment Score (NES) | 2.1418548 |
| Nominal p-value | 0.0 |
| FDR q-value | 2.3267716E-4 |
| FWER p-Value | 0.0030 |

| PROBE | GENE SYMBOL | GENE_TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|---|
| 1 | WSB1 | 75 | 5.191 | 0.0314 | Yes | ||
| 2 | JARID1B | 164 | 4.650 | 0.0584 | Yes | ||
| 3 | RHOT1 | 201 | 4.531 | 0.0874 | Yes | ||
| 4 | PRKACB | 246 | 4.348 | 0.1147 | Yes | ||
| 5 | ZFP36L1 | 311 | 4.187 | 0.1398 | Yes | ||
| 6 | MAN1A2 | 361 | 4.064 | 0.1649 | Yes | ||
| 7 | STK4 | 384 | 4.010 | 0.1911 | Yes | ||
| 8 | EHMT1 | 452 | 3.857 | 0.2138 | Yes | ||
| 9 | NEDD9 | 559 | 3.680 | 0.2332 | Yes | ||
| 10 | FCHSD2 | 651 | 3.547 | 0.2525 | Yes | ||
| 11 | RABGAP1 | 799 | 3.317 | 0.2672 | Yes | ||
| 12 | SMAD2 | 827 | 3.278 | 0.2881 | Yes | ||
| 13 | CTDSPL | 871 | 3.220 | 0.3078 | Yes | ||
| 14 | SAR1A | 888 | 3.198 | 0.3287 | Yes | ||
| 15 | HSF2 | 953 | 3.125 | 0.3466 | Yes | ||
| 16 | PARP6 | 1200 | 2.878 | 0.3530 | Yes | ||
| 17 | ASXL2 | 1479 | 2.617 | 0.3559 | Yes | ||
| 18 | SIM2 | 1575 | 2.532 | 0.3680 | Yes | ||
| 19 | IGF1 | 1722 | 2.417 | 0.3766 | Yes | ||
| 20 | NCOA1 | 1735 | 2.409 | 0.3924 | Yes | ||
| 21 | ALCAM | 1963 | 2.249 | 0.3955 | Yes | ||
| 22 | RUNX1 | 2071 | 2.175 | 0.4046 | Yes | ||
| 23 | NR3C1 | 2083 | 2.168 | 0.4188 | Yes | ||
| 24 | CDC42 | 2118 | 2.139 | 0.4316 | Yes | ||
| 25 | PURB | 2200 | 2.082 | 0.4414 | Yes | ||
| 26 | TEX2 | 2311 | 2.005 | 0.4492 | Yes | ||
| 27 | ANKRD50 | 2388 | 1.957 | 0.4584 | Yes | ||
| 28 | RAB5C | 2603 | 1.843 | 0.4595 | Yes | ||
| 29 | GAB1 | 2606 | 1.840 | 0.4719 | Yes | ||
| 30 | NAV1 | 2670 | 1.795 | 0.4808 | Yes | ||
| 31 | PHF2 | 2771 | 1.731 | 0.4872 | Yes | ||
| 32 | IRF2 | 2872 | 1.678 | 0.4933 | Yes | ||
| 33 | SON | 3127 | 1.555 | 0.4902 | Yes | ||
| 34 | AEBP2 | 3273 | 1.494 | 0.4926 | Yes | ||
| 35 | GLRB | 3304 | 1.476 | 0.5010 | Yes | ||
| 36 | AKR1D1 | 4133 | 1.128 | 0.4641 | No | ||
| 37 | MESP1 | 4278 | 1.076 | 0.4636 | No | ||
| 38 | NFAT5 | 4779 | 0.904 | 0.4428 | No | ||
| 39 | FNBP1 | 4789 | 0.899 | 0.4485 | No | ||
| 40 | BHLHB5 | 5144 | 0.789 | 0.4348 | No | ||
| 41 | TARDBP | 5221 | 0.768 | 0.4359 | No | ||
| 42 | DPP10 | 5226 | 0.767 | 0.4409 | No | ||
| 43 | ESR1 | 5859 | 0.578 | 0.4108 | No | ||
| 44 | GCLC | 6325 | 0.445 | 0.3888 | No | ||
| 45 | LIN28 | 6405 | 0.424 | 0.3874 | No | ||
| 46 | ARL15 | 6644 | 0.360 | 0.3770 | No | ||
| 47 | KIT | 6786 | 0.325 | 0.3716 | No | ||
| 48 | BTG3 | 6918 | 0.286 | 0.3665 | No | ||
| 49 | CLASP2 | 6981 | 0.268 | 0.3650 | No | ||
| 50 | PRICKLE2 | 7682 | 0.095 | 0.3279 | No | ||
| 51 | SNURF | 7685 | 0.095 | 0.3284 | No | ||
| 52 | FRMD4A | 7959 | 0.024 | 0.3139 | No | ||
| 53 | CTGF | 8162 | -0.024 | 0.3031 | No | ||
| 54 | MEF2D | 9812 | -0.420 | 0.2171 | No | ||
| 55 | ATXN1 | 9983 | -0.460 | 0.2110 | No | ||
| 56 | SH3BP4 | 10439 | -0.562 | 0.1903 | No | ||
| 57 | TRIB2 | 10916 | -0.667 | 0.1692 | No | ||
| 58 | ARHGEF5 | 11241 | -0.735 | 0.1568 | No | ||
| 59 | PHF19 | 11930 | -0.874 | 0.1256 | No | ||
| 60 | PERQ1 | 11994 | -0.885 | 0.1283 | No | ||
| 61 | MDGA1 | 12925 | -1.079 | 0.0855 | No | ||
| 62 | RAB11FIP2 | 13008 | -1.094 | 0.0885 | No | ||
| 63 | ZBTB4 | 13024 | -1.096 | 0.0952 | No | ||
| 64 | XYLT2 | 13867 | -1.298 | 0.0586 | No | ||
| 65 | CLK2 | 14896 | -1.561 | 0.0138 | No | ||
| 66 | PTGFRN | 14922 | -1.571 | 0.0232 | No | ||
| 67 | TRIOBP | 15551 | -1.758 | 0.0013 | No | ||
| 68 | PACSIN1 | 15603 | -1.776 | 0.0107 | No | ||
| 69 | NR1H2 | 15943 | -1.914 | 0.0055 | No | ||
| 70 | SOCS5 | 16084 | -1.974 | 0.0114 | No | ||
| 71 | HIF1A | 16331 | -2.081 | 0.0123 | No | ||
| 72 | ADD3 | 16532 | -2.177 | 0.0164 | No | ||
| 73 | PIAS3 | 17287 | -2.648 | -0.0062 | No | ||
| 74 | ETV6 | 17829 | -3.156 | -0.0138 | No | ||
| 75 | TRIM2 | 18296 | -4.052 | -0.0113 | No | ||
| 76 | CRIM1 | 18334 | -4.175 | 0.0152 | No |
