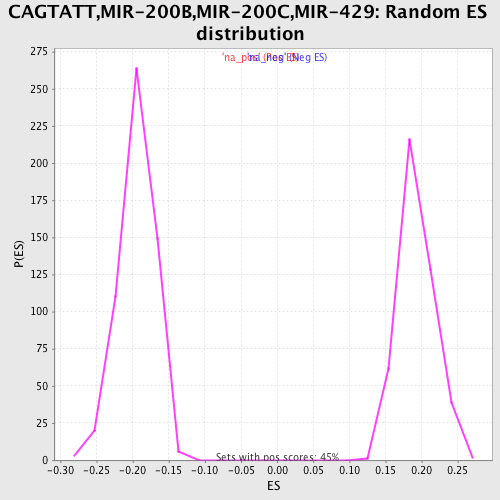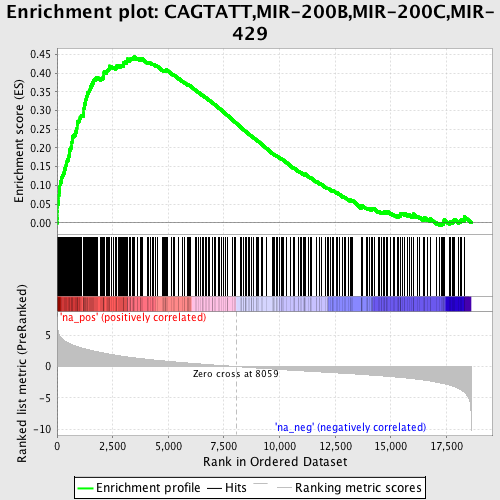
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | set03_absentNotch_versus_truncNotch |
| Phenotype | NoPhenotypeAvailable |
| Upregulated in class | na_pos |
| GeneSet | CAGTATT,MIR-200B,MIR-200C,MIR-429 |
| Enrichment Score (ES) | 0.44336918 |
| Normalized Enrichment Score (NES) | 2.299069 |
| Nominal p-value | 0.0 |
| FDR q-value | 0.0 |
| FWER p-Value | 0.0 |

| PROBE | GENE SYMBOL | GENE_TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|---|
| 1 | FREQ | 8 | 6.962 | 0.0111 | Yes | ||
| 2 | UBQLN1 | 19 | 6.067 | 0.0207 | Yes | ||
| 3 | CDYL | 20 | 6.035 | 0.0307 | Yes | ||
| 4 | SLC6A6 | 30 | 5.883 | 0.0400 | Yes | ||
| 5 | CALU | 34 | 5.802 | 0.0495 | Yes | ||
| 6 | ZFYVE20 | 51 | 5.476 | 0.0577 | Yes | ||
| 7 | ICK | 66 | 5.291 | 0.0657 | Yes | ||
| 8 | ITPR1 | 69 | 5.257 | 0.0744 | Yes | ||
| 9 | HS2ST1 | 87 | 5.048 | 0.0818 | Yes | ||
| 10 | DDEF1 | 106 | 4.920 | 0.0890 | Yes | ||
| 11 | CFL2 | 122 | 4.838 | 0.0963 | Yes | ||
| 12 | BRMS1L | 129 | 4.772 | 0.1039 | Yes | ||
| 13 | ARID2 | 159 | 4.666 | 0.1100 | Yes | ||
| 14 | RHOT1 | 201 | 4.531 | 0.1153 | Yes | ||
| 15 | SFRS1 | 209 | 4.486 | 0.1224 | Yes | ||
| 16 | PRKACB | 246 | 4.348 | 0.1277 | Yes | ||
| 17 | RAB6IP1 | 273 | 4.274 | 0.1334 | Yes | ||
| 18 | ATP2A2 | 317 | 4.171 | 0.1379 | Yes | ||
| 19 | BCL11B | 324 | 4.156 | 0.1445 | Yes | ||
| 20 | ACACA | 383 | 4.013 | 0.1480 | Yes | ||
| 21 | HNRPK | 396 | 3.977 | 0.1540 | Yes | ||
| 22 | NUDT4 | 426 | 3.911 | 0.1589 | Yes | ||
| 23 | ARL8B | 440 | 3.891 | 0.1646 | Yes | ||
| 24 | PPP2CA | 467 | 3.839 | 0.1696 | Yes | ||
| 25 | YWHAB | 527 | 3.736 | 0.1726 | Yes | ||
| 26 | TBK1 | 530 | 3.732 | 0.1787 | Yes | ||
| 27 | RAP1B | 546 | 3.696 | 0.1840 | Yes | ||
| 28 | CNOT7 | 557 | 3.681 | 0.1896 | Yes | ||
| 29 | RANBP9 | 570 | 3.662 | 0.1950 | Yes | ||
| 30 | KRAS | 590 | 3.637 | 0.2000 | Yes | ||
| 31 | CNN3 | 630 | 3.581 | 0.2038 | Yes | ||
| 32 | SLC38A2 | 645 | 3.552 | 0.2090 | Yes | ||
| 33 | KHDRBS1 | 646 | 3.552 | 0.2149 | Yes | ||
| 34 | OSBPL11 | 673 | 3.502 | 0.2193 | Yes | ||
| 35 | CRKL | 677 | 3.501 | 0.2250 | Yes | ||
| 36 | HIPK1 | 686 | 3.486 | 0.2303 | Yes | ||
| 37 | LFNG | 720 | 3.426 | 0.2342 | Yes | ||
| 38 | KLF9 | 775 | 3.344 | 0.2368 | Yes | ||
| 39 | SFRS2 | 836 | 3.269 | 0.2390 | Yes | ||
| 40 | DDX3X | 840 | 3.265 | 0.2442 | Yes | ||
| 41 | TCEB1 | 886 | 3.203 | 0.2471 | Yes | ||
| 42 | ANLN | 887 | 3.202 | 0.2524 | Yes | ||
| 43 | TIAL1 | 894 | 3.194 | 0.2574 | Yes | ||
| 44 | SLC23A2 | 895 | 3.193 | 0.2627 | Yes | ||
| 45 | NFYA | 906 | 3.180 | 0.2674 | Yes | ||
| 46 | SERINC1 | 920 | 3.165 | 0.2720 | Yes | ||
| 47 | TOB1 | 982 | 3.095 | 0.2738 | Yes | ||
| 48 | RAP2C | 1021 | 3.061 | 0.2768 | Yes | ||
| 49 | APRIN | 1027 | 3.058 | 0.2816 | Yes | ||
| 50 | RTF1 | 1070 | 3.013 | 0.2843 | Yes | ||
| 51 | SESN1 | 1099 | 2.980 | 0.2878 | Yes | ||
| 52 | NBR1 | 1173 | 2.899 | 0.2886 | Yes | ||
| 53 | HNRPU | 1176 | 2.897 | 0.2933 | Yes | ||
| 54 | PARP6 | 1200 | 2.878 | 0.2968 | Yes | ||
| 55 | NEDD1 | 1202 | 2.878 | 0.3016 | Yes | ||
| 56 | RECK | 1203 | 2.876 | 0.3063 | Yes | ||
| 57 | MYB | 1222 | 2.858 | 0.3101 | Yes | ||
| 58 | RANBP10 | 1230 | 2.847 | 0.3145 | Yes | ||
| 59 | ADIPOR2 | 1236 | 2.838 | 0.3189 | Yes | ||
| 60 | PUM2 | 1273 | 2.810 | 0.3216 | Yes | ||
| 61 | FBXW7 | 1279 | 2.806 | 0.3260 | Yes | ||
| 62 | TRIM33 | 1293 | 2.795 | 0.3299 | Yes | ||
| 63 | AKAP7 | 1326 | 2.767 | 0.3328 | Yes | ||
| 64 | ANK3 | 1329 | 2.765 | 0.3373 | Yes | ||
| 65 | CLASP1 | 1347 | 2.744 | 0.3409 | Yes | ||
| 66 | TSGA14 | 1369 | 2.724 | 0.3443 | Yes | ||
| 67 | SEMA3F | 1371 | 2.723 | 0.3488 | Yes | ||
| 68 | LRP4 | 1432 | 2.667 | 0.3499 | Yes | ||
| 69 | MMD | 1437 | 2.659 | 0.3541 | Yes | ||
| 70 | LASS6 | 1476 | 2.625 | 0.3564 | Yes | ||
| 71 | MARCKS | 1488 | 2.603 | 0.3601 | Yes | ||
| 72 | MSN | 1501 | 2.593 | 0.3638 | Yes | ||
| 73 | PHTF2 | 1535 | 2.567 | 0.3662 | Yes | ||
| 74 | BIRC4 | 1553 | 2.550 | 0.3695 | Yes | ||
| 75 | CNOT4 | 1591 | 2.520 | 0.3717 | Yes | ||
| 76 | CDR2 | 1593 | 2.517 | 0.3758 | Yes | ||
| 77 | TBP | 1642 | 2.479 | 0.3773 | Yes | ||
| 78 | PDPK1 | 1643 | 2.477 | 0.3815 | Yes | ||
| 79 | GABPA | 1689 | 2.441 | 0.3831 | Yes | ||
| 80 | NIN | 1734 | 2.410 | 0.3847 | Yes | ||
| 81 | SMARCAD1 | 1756 | 2.394 | 0.3875 | Yes | ||
| 82 | BCAP29 | 1818 | 2.351 | 0.3881 | Yes | ||
| 83 | VLDLR | 1959 | 2.251 | 0.3841 | Yes | ||
| 84 | CYP1B1 | 1986 | 2.232 | 0.3864 | Yes | ||
| 85 | E2F3 | 2042 | 2.193 | 0.3871 | Yes | ||
| 86 | NR3C1 | 2083 | 2.168 | 0.3885 | Yes | ||
| 87 | PHF6 | 2088 | 2.166 | 0.3919 | Yes | ||
| 88 | ZIC3 | 2100 | 2.160 | 0.3949 | Yes | ||
| 89 | PPM1B | 2103 | 2.157 | 0.3983 | Yes | ||
| 90 | YTHDF3 | 2104 | 2.156 | 0.4019 | Yes | ||
| 91 | WDFY3 | 2108 | 2.150 | 0.4053 | Yes | ||
| 92 | MAFG | 2230 | 2.065 | 0.4021 | Yes | ||
| 93 | OGT | 2242 | 2.058 | 0.4050 | Yes | ||
| 94 | FBXO33 | 2248 | 2.054 | 0.4081 | Yes | ||
| 95 | KLF4 | 2293 | 2.019 | 0.4091 | Yes | ||
| 96 | TSC22D1 | 2331 | 1.992 | 0.4103 | Yes | ||
| 97 | PCDH7 | 2338 | 1.986 | 0.4133 | Yes | ||
| 98 | NCOA2 | 2348 | 1.980 | 0.4161 | Yes | ||
| 99 | ELF2 | 2357 | 1.974 | 0.4190 | Yes | ||
| 100 | RAC1 | 2457 | 1.919 | 0.4167 | Yes | ||
| 101 | PPAP2B | 2543 | 1.868 | 0.4152 | Yes | ||
| 102 | FLI1 | 2632 | 1.817 | 0.4134 | Yes | ||
| 103 | SLC6A1 | 2654 | 1.805 | 0.4152 | Yes | ||
| 104 | FN1 | 2668 | 1.796 | 0.4175 | Yes | ||
| 105 | PSIP1 | 2689 | 1.781 | 0.4194 | Yes | ||
| 106 | NTF3 | 2741 | 1.751 | 0.4195 | Yes | ||
| 107 | DMRT2 | 2810 | 1.712 | 0.4186 | Yes | ||
| 108 | SMURF2 | 2861 | 1.683 | 0.4187 | Yes | ||
| 109 | RABIF | 2878 | 1.676 | 0.4206 | Yes | ||
| 110 | CSNK1G3 | 2955 | 1.636 | 0.4192 | Yes | ||
| 111 | CLIC4 | 2982 | 1.624 | 0.4204 | Yes | ||
| 112 | DNAJB6 | 2988 | 1.620 | 0.4229 | Yes | ||
| 113 | GOLGA7 | 2996 | 1.616 | 0.4252 | Yes | ||
| 114 | MATR3 | 2999 | 1.615 | 0.4277 | Yes | ||
| 115 | OLIG3 | 3037 | 1.599 | 0.4284 | Yes | ||
| 116 | CHD9 | 3054 | 1.591 | 0.4302 | Yes | ||
| 117 | HNRPH2 | 3134 | 1.551 | 0.4284 | Yes | ||
| 118 | NRIP1 | 3141 | 1.549 | 0.4307 | Yes | ||
| 119 | PIP5K3 | 3142 | 1.549 | 0.4332 | Yes | ||
| 120 | ULK2 | 3146 | 1.548 | 0.4356 | Yes | ||
| 121 | EVI5 | 3154 | 1.543 | 0.4378 | Yes | ||
| 122 | AEBP2 | 3273 | 1.494 | 0.4339 | Yes | ||
| 123 | LAMC1 | 3280 | 1.489 | 0.4360 | Yes | ||
| 124 | DACT1 | 3297 | 1.480 | 0.4376 | Yes | ||
| 125 | MAP4K4 | 3316 | 1.473 | 0.4390 | Yes | ||
| 126 | ZC3H6 | 3376 | 1.447 | 0.4382 | Yes | ||
| 127 | HNRPD | 3413 | 1.430 | 0.4386 | Yes | ||
| 128 | RPS6KA3 | 3437 | 1.415 | 0.4397 | Yes | ||
| 129 | CITED2 | 3446 | 1.410 | 0.4416 | Yes | ||
| 130 | EGLN1 | 3458 | 1.405 | 0.4434 | Yes | ||
| 131 | MAP4K3 | 3592 | 1.348 | 0.4383 | No | ||
| 132 | ASF1A | 3617 | 1.333 | 0.4392 | No | ||
| 133 | THRAP1 | 3741 | 1.280 | 0.4346 | No | ||
| 134 | OXR1 | 3755 | 1.275 | 0.4360 | No | ||
| 135 | YWHAG | 3760 | 1.274 | 0.4379 | No | ||
| 136 | ARID4B | 3811 | 1.250 | 0.4373 | No | ||
| 137 | CNTN4 | 3824 | 1.245 | 0.4387 | No | ||
| 138 | PCNP | 4062 | 1.160 | 0.4277 | No | ||
| 139 | GOSR2 | 4068 | 1.157 | 0.4293 | No | ||
| 140 | NOG | 4113 | 1.136 | 0.4288 | No | ||
| 141 | GDI2 | 4179 | 1.115 | 0.4271 | No | ||
| 142 | QKI | 4270 | 1.078 | 0.4240 | No | ||
| 143 | KCNJ2 | 4302 | 1.069 | 0.4240 | No | ||
| 144 | PDCD10 | 4342 | 1.054 | 0.4237 | No | ||
| 145 | ORMDL3 | 4439 | 1.021 | 0.4201 | No | ||
| 146 | DCUN1D1 | 4504 | 0.998 | 0.4183 | No | ||
| 147 | UBE2I | 4730 | 0.920 | 0.4075 | No | ||
| 148 | AMFR | 4785 | 0.901 | 0.4060 | No | ||
| 149 | PALM2 | 4793 | 0.897 | 0.4071 | No | ||
| 150 | UBE2D1 | 4847 | 0.885 | 0.4057 | No | ||
| 151 | SULF1 | 4849 | 0.885 | 0.4071 | No | ||
| 152 | TUBB | 4892 | 0.869 | 0.4063 | No | ||
| 153 | NGEF | 4893 | 0.869 | 0.4077 | No | ||
| 154 | DDX3Y | 4895 | 0.868 | 0.4091 | No | ||
| 155 | NPM1 | 4948 | 0.854 | 0.4077 | No | ||
| 156 | OCLN | 5142 | 0.790 | 0.3984 | No | ||
| 157 | TARDBP | 5221 | 0.768 | 0.3954 | No | ||
| 158 | TMEFF2 | 5280 | 0.751 | 0.3935 | No | ||
| 159 | THAP1 | 5456 | 0.691 | 0.3851 | No | ||
| 160 | ESRRG | 5457 | 0.691 | 0.3862 | No | ||
| 161 | GEM | 5613 | 0.642 | 0.3788 | No | ||
| 162 | PAPOLA | 5716 | 0.615 | 0.3743 | No | ||
| 163 | NIPA1 | 5723 | 0.615 | 0.3750 | No | ||
| 164 | DAG1 | 5858 | 0.579 | 0.3686 | No | ||
| 165 | SYVN1 | 5882 | 0.573 | 0.3683 | No | ||
| 166 | SSH2 | 5898 | 0.569 | 0.3684 | No | ||
| 167 | KCTD15 | 5910 | 0.566 | 0.3687 | No | ||
| 168 | DGKA | 5967 | 0.546 | 0.3666 | No | ||
| 169 | TEAD1 | 5982 | 0.542 | 0.3667 | No | ||
| 170 | STX1A | 6213 | 0.476 | 0.3549 | No | ||
| 171 | SCHIP1 | 6279 | 0.458 | 0.3521 | No | ||
| 172 | RND3 | 6352 | 0.439 | 0.3489 | No | ||
| 173 | EIF2S1 | 6466 | 0.408 | 0.3434 | No | ||
| 174 | SNAP25 | 6512 | 0.397 | 0.3416 | No | ||
| 175 | GATA2 | 6542 | 0.390 | 0.3407 | No | ||
| 176 | IER5 | 6588 | 0.377 | 0.3388 | No | ||
| 177 | PAM | 6677 | 0.351 | 0.3346 | No | ||
| 178 | CRH | 6727 | 0.340 | 0.3325 | No | ||
| 179 | FHL1 | 6822 | 0.315 | 0.3279 | No | ||
| 180 | NPTX1 | 6864 | 0.300 | 0.3261 | No | ||
| 181 | CLASP2 | 6981 | 0.268 | 0.3202 | No | ||
| 182 | UBE2B | 7000 | 0.263 | 0.3197 | No | ||
| 183 | YPEL2 | 7003 | 0.263 | 0.3200 | No | ||
| 184 | FHOD1 | 7060 | 0.248 | 0.3174 | No | ||
| 185 | LRP1 | 7097 | 0.238 | 0.3158 | No | ||
| 186 | RAB21 | 7273 | 0.198 | 0.3065 | No | ||
| 187 | EDG2 | 7313 | 0.188 | 0.3047 | No | ||
| 188 | BAP1 | 7367 | 0.176 | 0.3021 | No | ||
| 189 | EFNB2 | 7464 | 0.149 | 0.2971 | No | ||
| 190 | PKD1 | 7580 | 0.122 | 0.2910 | No | ||
| 191 | PLEKHC1 | 7659 | 0.100 | 0.2869 | No | ||
| 192 | DNAJB5 | 7676 | 0.096 | 0.2862 | No | ||
| 193 | EFNA1 | 7677 | 0.096 | 0.2863 | No | ||
| 194 | SEC23A | 7894 | 0.042 | 0.2746 | No | ||
| 195 | PCTK2 | 7984 | 0.019 | 0.2698 | No | ||
| 196 | KLF10 | 8004 | 0.013 | 0.2687 | No | ||
| 197 | ST6GALNAC5 | 8222 | -0.038 | 0.2569 | No | ||
| 198 | KCTD8 | 8283 | -0.055 | 0.2537 | No | ||
| 199 | ZFPM2 | 8298 | -0.059 | 0.2531 | No | ||
| 200 | PLS3 | 8367 | -0.078 | 0.2495 | No | ||
| 201 | CCNJ | 8391 | -0.085 | 0.2484 | No | ||
| 202 | GLRA2 | 8450 | -0.101 | 0.2454 | No | ||
| 203 | DUSP1 | 8464 | -0.107 | 0.2448 | No | ||
| 204 | NUP153 | 8465 | -0.107 | 0.2450 | No | ||
| 205 | PARD6B | 8504 | -0.116 | 0.2431 | No | ||
| 206 | RGL1 | 8623 | -0.144 | 0.2369 | No | ||
| 207 | CAB39 | 8650 | -0.150 | 0.2357 | No | ||
| 208 | FBXW11 | 8723 | -0.167 | 0.2321 | No | ||
| 209 | HIC2 | 8744 | -0.172 | 0.2313 | No | ||
| 210 | B3GNT1 | 8758 | -0.178 | 0.2308 | No | ||
| 211 | KCNQ4 | 8837 | -0.196 | 0.2269 | No | ||
| 212 | AFF3 | 8843 | -0.198 | 0.2270 | No | ||
| 213 | ACY1L2 | 8952 | -0.219 | 0.2214 | No | ||
| 214 | EPS8 | 8977 | -0.225 | 0.2205 | No | ||
| 215 | MYT1 | 8989 | -0.228 | 0.2202 | No | ||
| 216 | PHF21A | 9066 | -0.250 | 0.2165 | No | ||
| 217 | PRDM1 | 9208 | -0.281 | 0.2093 | No | ||
| 218 | CBX4 | 9252 | -0.289 | 0.2074 | No | ||
| 219 | GNAI3 | 9433 | -0.326 | 0.1981 | No | ||
| 220 | ARHGEF17 | 9665 | -0.383 | 0.1861 | No | ||
| 221 | SYT1 | 9720 | -0.395 | 0.1838 | No | ||
| 222 | CORO1C | 9738 | -0.399 | 0.1835 | No | ||
| 223 | AKAP2 | 9791 | -0.412 | 0.1813 | No | ||
| 224 | LEPR | 9850 | -0.427 | 0.1789 | No | ||
| 225 | TRAM1L1 | 9864 | -0.431 | 0.1789 | No | ||
| 226 | MFAP5 | 9906 | -0.443 | 0.1774 | No | ||
| 227 | ATXN1 | 9983 | -0.460 | 0.1740 | No | ||
| 228 | PTBP1 | 10011 | -0.467 | 0.1733 | No | ||
| 229 | USP25 | 10063 | -0.477 | 0.1713 | No | ||
| 230 | TAF4 | 10103 | -0.489 | 0.1700 | No | ||
| 231 | DACH1 | 10125 | -0.494 | 0.1696 | No | ||
| 232 | SEMA6D | 10187 | -0.506 | 0.1671 | No | ||
| 233 | SLC38A4 | 10322 | -0.537 | 0.1607 | No | ||
| 234 | TNFRSF11B | 10500 | -0.574 | 0.1520 | No | ||
| 235 | NEGR1 | 10603 | -0.598 | 0.1474 | No | ||
| 236 | PKIA | 10656 | -0.611 | 0.1456 | No | ||
| 237 | TCF2 | 10687 | -0.616 | 0.1449 | No | ||
| 238 | GPM6A | 10842 | -0.651 | 0.1376 | No | ||
| 239 | HDHD2 | 10857 | -0.655 | 0.1379 | No | ||
| 240 | LHFP | 10930 | -0.669 | 0.1351 | No | ||
| 241 | KCNK2 | 10978 | -0.677 | 0.1336 | No | ||
| 242 | GIT2 | 11083 | -0.699 | 0.1291 | No | ||
| 243 | HLF | 11123 | -0.709 | 0.1282 | No | ||
| 244 | SDC2 | 11125 | -0.710 | 0.1293 | No | ||
| 245 | PLCL1 | 11139 | -0.712 | 0.1298 | No | ||
| 246 | HMGB3 | 11143 | -0.713 | 0.1308 | No | ||
| 247 | BHLHB3 | 11277 | -0.741 | 0.1247 | No | ||
| 248 | NAP1L5 | 11386 | -0.766 | 0.1201 | No | ||
| 249 | MPP5 | 11407 | -0.770 | 0.1203 | No | ||
| 250 | NXPH1 | 11442 | -0.777 | 0.1197 | No | ||
| 251 | GATA4 | 11636 | -0.815 | 0.1105 | No | ||
| 252 | NAB1 | 11669 | -0.821 | 0.1101 | No | ||
| 253 | TUBB2A | 11798 | -0.847 | 0.1045 | No | ||
| 254 | ACVR2A | 11808 | -0.850 | 0.1054 | No | ||
| 255 | AMOTL2 | 11865 | -0.862 | 0.1038 | No | ||
| 256 | EIF5B | 12050 | -0.897 | 0.0952 | No | ||
| 257 | RHOA | 12134 | -0.912 | 0.0922 | No | ||
| 258 | PTPN13 | 12160 | -0.918 | 0.0924 | No | ||
| 259 | PPP2R5C | 12176 | -0.921 | 0.0931 | No | ||
| 260 | CASR | 12289 | -0.945 | 0.0885 | No | ||
| 261 | NDN | 12356 | -0.961 | 0.0865 | No | ||
| 262 | LRRTM3 | 12392 | -0.968 | 0.0862 | No | ||
| 263 | PLK2 | 12442 | -0.977 | 0.0852 | No | ||
| 264 | SLC1A2 | 12568 | -1.005 | 0.0800 | No | ||
| 265 | SCD | 12580 | -1.008 | 0.0811 | No | ||
| 266 | SOX1 | 12693 | -1.032 | 0.0766 | No | ||
| 267 | CACNA1C | 12845 | -1.060 | 0.0701 | No | ||
| 268 | NR5A2 | 12904 | -1.072 | 0.0688 | No | ||
| 269 | PCTK1 | 12949 | -1.084 | 0.0681 | No | ||
| 270 | FGD1 | 13114 | -1.115 | 0.0610 | No | ||
| 271 | FBXW2 | 13169 | -1.126 | 0.0599 | No | ||
| 272 | SOX2 | 13176 | -1.128 | 0.0615 | No | ||
| 273 | FMR1 | 13235 | -1.144 | 0.0602 | No | ||
| 274 | PIN1 | 13277 | -1.154 | 0.0599 | No | ||
| 275 | ATRX | 13665 | -1.248 | 0.0408 | No | ||
| 276 | ADCY2 | 13676 | -1.251 | 0.0423 | No | ||
| 277 | KCND2 | 13678 | -1.252 | 0.0444 | No | ||
| 278 | LCP1 | 13682 | -1.252 | 0.0463 | No | ||
| 279 | HS3ST1 | 13723 | -1.261 | 0.0462 | No | ||
| 280 | PCDH8 | 13884 | -1.302 | 0.0396 | No | ||
| 281 | LIN7B | 13951 | -1.317 | 0.0382 | No | ||
| 282 | SLC14A1 | 13964 | -1.319 | 0.0397 | No | ||
| 283 | NFIA | 14031 | -1.334 | 0.0383 | No | ||
| 284 | CSMD3 | 14128 | -1.359 | 0.0353 | No | ||
| 285 | PRKAR2B | 14139 | -1.362 | 0.0370 | No | ||
| 286 | BASP1 | 14179 | -1.373 | 0.0372 | No | ||
| 287 | CDH11 | 14188 | -1.375 | 0.0390 | No | ||
| 288 | FGFR2 | 14249 | -1.390 | 0.0381 | No | ||
| 289 | NRP2 | 14423 | -1.437 | 0.0310 | No | ||
| 290 | ETS1 | 14475 | -1.449 | 0.0306 | No | ||
| 291 | GLI3 | 14588 | -1.480 | 0.0269 | No | ||
| 292 | ELAVL2 | 14602 | -1.483 | 0.0287 | No | ||
| 293 | RNF5 | 14686 | -1.505 | 0.0266 | No | ||
| 294 | NPC1 | 14722 | -1.512 | 0.0272 | No | ||
| 295 | GPR146 | 14725 | -1.513 | 0.0297 | No | ||
| 296 | LRP1B | 14806 | -1.534 | 0.0278 | No | ||
| 297 | PAK7 | 14815 | -1.536 | 0.0299 | No | ||
| 298 | TUBB3 | 14866 | -1.552 | 0.0298 | No | ||
| 299 | LYPLA2 | 14991 | -1.589 | 0.0256 | No | ||
| 300 | SCN3B | 15117 | -1.627 | 0.0215 | No | ||
| 301 | RKHD3 | 15178 | -1.647 | 0.0210 | No | ||
| 302 | BRWD1 | 15300 | -1.684 | 0.0171 | No | ||
| 303 | ARHGDIA | 15335 | -1.696 | 0.0181 | No | ||
| 304 | FOXK1 | 15346 | -1.699 | 0.0204 | No | ||
| 305 | SBF1 | 15419 | -1.723 | 0.0193 | No | ||
| 306 | PHACTR3 | 15437 | -1.727 | 0.0213 | No | ||
| 307 | HECTD2 | 15442 | -1.728 | 0.0239 | No | ||
| 308 | FEZ2 | 15454 | -1.732 | 0.0262 | No | ||
| 309 | SLC16A2 | 15506 | -1.747 | 0.0263 | No | ||
| 310 | KLF12 | 15631 | -1.785 | 0.0225 | No | ||
| 311 | AP1S2 | 15635 | -1.787 | 0.0253 | No | ||
| 312 | DNMT3A | 15769 | -1.842 | 0.0211 | No | ||
| 313 | ACE2 | 15818 | -1.866 | 0.0215 | No | ||
| 314 | PPP4R2 | 15940 | -1.914 | 0.0181 | No | ||
| 315 | MGAT2 | 16004 | -1.940 | 0.0179 | No | ||
| 316 | COPS8 | 16010 | -1.942 | 0.0208 | No | ||
| 317 | MXD3 | 16013 | -1.943 | 0.0240 | No | ||
| 318 | RPS6KA2 | 16179 | -2.017 | 0.0183 | No | ||
| 319 | RDH10 | 16291 | -2.060 | 0.0156 | No | ||
| 320 | MTUS1 | 16467 | -2.144 | 0.0096 | No | ||
| 321 | MMD2 | 16510 | -2.163 | 0.0109 | No | ||
| 322 | CNTFR | 16514 | -2.166 | 0.0144 | No | ||
| 323 | PPP1R9B | 16665 | -2.244 | 0.0099 | No | ||
| 324 | CREB5 | 16764 | -2.299 | 0.0084 | No | ||
| 325 | EGR3 | 16768 | -2.302 | 0.0120 | No | ||
| 326 | RPS6KB1 | 17050 | -2.477 | 0.0008 | No | ||
| 327 | NDST1 | 17193 | -2.579 | -0.0027 | No | ||
| 328 | SRF | 17295 | -2.658 | -0.0038 | No | ||
| 329 | RKHD2 | 17314 | -2.675 | -0.0004 | No | ||
| 330 | PPP1R10 | 17379 | -2.722 | 0.0007 | No | ||
| 331 | PTHLH | 17380 | -2.723 | 0.0052 | No | ||
| 332 | BAT3 | 17404 | -2.739 | 0.0085 | No | ||
| 333 | IHPK1 | 17623 | -2.942 | 0.0014 | No | ||
| 334 | SMARCD1 | 17673 | -2.995 | 0.0037 | No | ||
| 335 | IGSF3 | 17773 | -3.095 | 0.0035 | No | ||
| 336 | ETV5 | 17825 | -3.153 | 0.0059 | No | ||
| 337 | EIF2B5 | 17865 | -3.212 | 0.0091 | No | ||
| 338 | DLC1 | 18054 | -3.505 | 0.0047 | No | ||
| 339 | HRB | 18138 | -3.665 | 0.0062 | No | ||
| 340 | PCAF | 18187 | -3.745 | 0.0098 | No | ||
| 341 | TRIM2 | 18296 | -4.052 | 0.0107 | No | ||
| 342 | PPM1F | 18309 | -4.090 | 0.0168 | No |