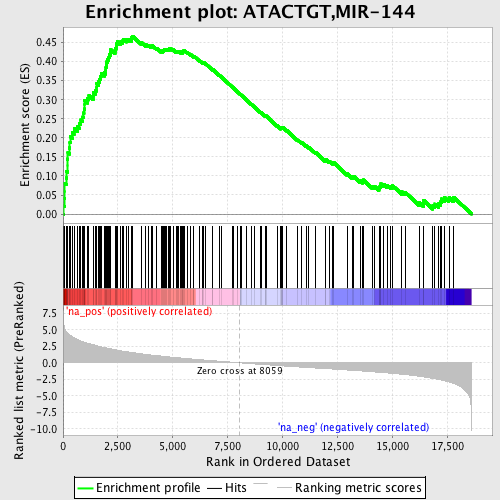
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | set03_absentNotch_versus_truncNotch |
| Phenotype | NoPhenotypeAvailable |
| Upregulated in class | na_pos |
| GeneSet | ATACTGT,MIR-144 |
| Enrichment Score (ES) | 0.4669647 |
| Normalized Enrichment Score (NES) | 2.1995726 |
| Nominal p-value | 0.0 |
| FDR q-value | 2.9084642E-4 |
| FWER p-Value | 0.0030 |

| PROBE | GENE SYMBOL | GENE_TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|---|
| 1 | APPBP2 | 21 | 5.998 | 0.0220 | Yes | ||
| 2 | SS18 | 56 | 5.427 | 0.0411 | Yes | ||
| 3 | GLCCI1 | 74 | 5.228 | 0.0603 | Yes | ||
| 4 | PPP3R1 | 77 | 5.168 | 0.0802 | Yes | ||
| 5 | SIN3A | 142 | 4.711 | 0.0949 | Yes | ||
| 6 | ARID2 | 159 | 4.666 | 0.1120 | Yes | ||
| 7 | SLC12A2 | 203 | 4.524 | 0.1272 | Yes | ||
| 8 | ACBD3 | 204 | 4.515 | 0.1446 | Yes | ||
| 9 | RBM12 | 219 | 4.466 | 0.1610 | Yes | ||
| 10 | MAPK6 | 289 | 4.244 | 0.1737 | Yes | ||
| 11 | PAFAH1B1 | 309 | 4.189 | 0.1888 | Yes | ||
| 12 | PPP1R16B | 338 | 4.131 | 0.2032 | Yes | ||
| 13 | GSPT1 | 424 | 3.914 | 0.2137 | Yes | ||
| 14 | HAT1 | 501 | 3.775 | 0.2242 | Yes | ||
| 15 | EIF4G2 | 660 | 3.525 | 0.2292 | Yes | ||
| 16 | PKNOX1 | 756 | 3.371 | 0.2371 | Yes | ||
| 17 | CCNE2 | 810 | 3.300 | 0.2469 | Yes | ||
| 18 | SLC23A2 | 895 | 3.193 | 0.2547 | Yes | ||
| 19 | PFKFB2 | 924 | 3.157 | 0.2654 | Yes | ||
| 20 | HSF2 | 953 | 3.125 | 0.2759 | Yes | ||
| 21 | CUGBP2 | 980 | 3.096 | 0.2864 | Yes | ||
| 22 | FBXO32 | 984 | 3.093 | 0.2982 | Yes | ||
| 23 | ARID1A | 1116 | 2.949 | 0.3025 | Yes | ||
| 24 | HNRPU | 1176 | 2.897 | 0.3105 | Yes | ||
| 25 | PUM1 | 1370 | 2.724 | 0.3105 | Yes | ||
| 26 | MTPN | 1392 | 2.703 | 0.3198 | Yes | ||
| 27 | MARCKS | 1488 | 2.603 | 0.3247 | Yes | ||
| 28 | PABPN1 | 1528 | 2.574 | 0.3325 | Yes | ||
| 29 | PHTF2 | 1535 | 2.567 | 0.3421 | Yes | ||
| 30 | PTP4A1 | 1618 | 2.497 | 0.3473 | Yes | ||
| 31 | SLC7A11 | 1671 | 2.458 | 0.3540 | Yes | ||
| 32 | RNF111 | 1694 | 2.437 | 0.3622 | Yes | ||
| 33 | NIN | 1734 | 2.410 | 0.3694 | Yes | ||
| 34 | CLK1 | 1871 | 2.306 | 0.3709 | Yes | ||
| 35 | EIF5 | 1942 | 2.258 | 0.3758 | Yes | ||
| 36 | CBX1 | 1950 | 2.254 | 0.3841 | Yes | ||
| 37 | KPNA1 | 1965 | 2.248 | 0.3921 | Yes | ||
| 38 | IDH2 | 1993 | 2.227 | 0.3992 | Yes | ||
| 39 | ZFX | 2014 | 2.215 | 0.4067 | Yes | ||
| 40 | NR3C1 | 2083 | 2.168 | 0.4113 | Yes | ||
| 41 | WDFY3 | 2108 | 2.150 | 0.4183 | Yes | ||
| 42 | STAG1 | 2159 | 2.109 | 0.4238 | Yes | ||
| 43 | UBE2D2 | 2174 | 2.100 | 0.4311 | Yes | ||
| 44 | ITSN2 | 2367 | 1.969 | 0.4283 | Yes | ||
| 45 | BRPF1 | 2378 | 1.962 | 0.4353 | Yes | ||
| 46 | UBE2D3 | 2421 | 1.941 | 0.4405 | Yes | ||
| 47 | GCNT2 | 2444 | 1.925 | 0.4468 | Yes | ||
| 48 | VKORC1L1 | 2473 | 1.911 | 0.4526 | Yes | ||
| 49 | NARG1 | 2618 | 1.829 | 0.4519 | Yes | ||
| 50 | RIN2 | 2700 | 1.775 | 0.4544 | Yes | ||
| 51 | NFE2L2 | 2747 | 1.748 | 0.4586 | Yes | ||
| 52 | ACSL4 | 2880 | 1.676 | 0.4579 | Yes | ||
| 53 | CD28 | 2991 | 1.619 | 0.4582 | Yes | ||
| 54 | TFRC | 3112 | 1.563 | 0.4577 | Yes | ||
| 55 | SON | 3127 | 1.555 | 0.4630 | Yes | ||
| 56 | MYO1E | 3164 | 1.538 | 0.4670 | Yes | ||
| 57 | PDE4D | 3575 | 1.357 | 0.4500 | No | ||
| 58 | NEDD4 | 3754 | 1.275 | 0.4453 | No | ||
| 59 | GATA3 | 3881 | 1.221 | 0.4432 | No | ||
| 60 | TTN | 4018 | 1.171 | 0.4403 | No | ||
| 61 | ARFGEF1 | 4077 | 1.152 | 0.4416 | No | ||
| 62 | QKI | 4270 | 1.078 | 0.4354 | No | ||
| 63 | STARD8 | 4500 | 1.000 | 0.4268 | No | ||
| 64 | SNAP91 | 4542 | 0.984 | 0.4284 | No | ||
| 65 | CCNK | 4595 | 0.965 | 0.4293 | No | ||
| 66 | PANK1 | 4602 | 0.962 | 0.4327 | No | ||
| 67 | CAV2 | 4672 | 0.941 | 0.4326 | No | ||
| 68 | NGFRAP1 | 4729 | 0.920 | 0.4331 | No | ||
| 69 | PALM2 | 4793 | 0.897 | 0.4332 | No | ||
| 70 | PCDH18 | 4826 | 0.889 | 0.4349 | No | ||
| 71 | DMD | 4909 | 0.863 | 0.4337 | No | ||
| 72 | FBXL3 | 5013 | 0.828 | 0.4314 | No | ||
| 73 | FNDC3A | 5146 | 0.789 | 0.4273 | No | ||
| 74 | SMARCA1 | 5211 | 0.772 | 0.4268 | No | ||
| 75 | AHCY | 5255 | 0.760 | 0.4274 | No | ||
| 76 | ZCCHC2 | 5348 | 0.728 | 0.4252 | No | ||
| 77 | ALS2 | 5396 | 0.713 | 0.4254 | No | ||
| 78 | ESRRG | 5457 | 0.691 | 0.4248 | No | ||
| 79 | LGR4 | 5460 | 0.691 | 0.4274 | No | ||
| 80 | HNRPF | 5494 | 0.681 | 0.4282 | No | ||
| 81 | SUCLA2 | 5534 | 0.669 | 0.4287 | No | ||
| 82 | RHOBTB1 | 5654 | 0.631 | 0.4247 | No | ||
| 83 | LHX2 | 5797 | 0.596 | 0.4193 | No | ||
| 84 | ARRDC3 | 5940 | 0.555 | 0.4137 | No | ||
| 85 | FMN2 | 5963 | 0.548 | 0.4147 | No | ||
| 86 | WTAP | 6211 | 0.476 | 0.4031 | No | ||
| 87 | WIF1 | 6343 | 0.441 | 0.3977 | No | ||
| 88 | CAPZA1 | 6371 | 0.433 | 0.3979 | No | ||
| 89 | FOSB | 6417 | 0.421 | 0.3971 | No | ||
| 90 | SLITRK4 | 6511 | 0.397 | 0.3936 | No | ||
| 91 | UBE4A | 6811 | 0.319 | 0.3786 | No | ||
| 92 | RARB | 6819 | 0.316 | 0.3795 | No | ||
| 93 | JPH1 | 7106 | 0.236 | 0.3649 | No | ||
| 94 | MEIS1 | 7214 | 0.211 | 0.3599 | No | ||
| 95 | ZIC4 | 7739 | 0.081 | 0.3319 | No | ||
| 96 | RGS16 | 7769 | 0.074 | 0.3306 | No | ||
| 97 | PPARA | 7960 | 0.023 | 0.3204 | No | ||
| 98 | CALCRL | 8064 | -0.001 | 0.3148 | No | ||
| 99 | SP4 | 8066 | -0.002 | 0.3147 | No | ||
| 100 | GDF10 | 8127 | -0.014 | 0.3116 | No | ||
| 101 | SFRP1 | 8363 | -0.077 | 0.2991 | No | ||
| 102 | FOXO1A | 8601 | -0.140 | 0.2868 | No | ||
| 103 | FBXW11 | 8723 | -0.167 | 0.2809 | No | ||
| 104 | MYT1 | 8989 | -0.228 | 0.2674 | No | ||
| 105 | FAM60A | 9043 | -0.245 | 0.2655 | No | ||
| 106 | ALDH1A3 | 9201 | -0.280 | 0.2581 | No | ||
| 107 | PIM1 | 9229 | -0.285 | 0.2577 | No | ||
| 108 | CBX4 | 9252 | -0.289 | 0.2576 | No | ||
| 109 | ANK2 | 9751 | -0.402 | 0.2322 | No | ||
| 110 | ATP2B2 | 9922 | -0.445 | 0.2247 | No | ||
| 111 | PURA | 9948 | -0.451 | 0.2251 | No | ||
| 112 | PAX3 | 9951 | -0.451 | 0.2268 | No | ||
| 113 | ATXN1 | 9983 | -0.460 | 0.2269 | No | ||
| 114 | PNRC1 | 9991 | -0.462 | 0.2283 | No | ||
| 115 | SEMA6D | 10187 | -0.506 | 0.2196 | No | ||
| 116 | CPEB2 | 10697 | -0.619 | 0.1945 | No | ||
| 117 | HDHD2 | 10857 | -0.655 | 0.1884 | No | ||
| 118 | CRSP2 | 11079 | -0.698 | 0.1791 | No | ||
| 119 | CCT2 | 11192 | -0.724 | 0.1758 | No | ||
| 120 | CXCL12 | 11524 | -0.791 | 0.1609 | No | ||
| 121 | TRIB1 | 11938 | -0.876 | 0.1420 | No | ||
| 122 | ST6GALNAC3 | 11956 | -0.879 | 0.1444 | No | ||
| 123 | ARID5B | 12148 | -0.915 | 0.1376 | No | ||
| 124 | FUSIP1 | 12272 | -0.942 | 0.1346 | No | ||
| 125 | RFX4 | 12334 | -0.956 | 0.1350 | No | ||
| 126 | BACH2 | 12939 | -1.081 | 0.1064 | No | ||
| 127 | MSX1 | 13198 | -1.135 | 0.0968 | No | ||
| 128 | FMR1 | 13235 | -1.144 | 0.0993 | No | ||
| 129 | PTPN9 | 13545 | -1.217 | 0.0873 | No | ||
| 130 | ELL2 | 13658 | -1.245 | 0.0860 | No | ||
| 131 | SMPD3 | 13680 | -1.252 | 0.0897 | No | ||
| 132 | PLAG1 | 14091 | -1.351 | 0.0727 | No | ||
| 133 | CDH11 | 14188 | -1.375 | 0.0728 | No | ||
| 134 | LIFR | 14398 | -1.432 | 0.0670 | No | ||
| 135 | NRP2 | 14423 | -1.437 | 0.0712 | No | ||
| 136 | ETS1 | 14475 | -1.449 | 0.0741 | No | ||
| 137 | EDAR | 14485 | -1.451 | 0.0792 | No | ||
| 138 | TBX1 | 14610 | -1.486 | 0.0782 | No | ||
| 139 | ST18 | 14785 | -1.529 | 0.0747 | No | ||
| 140 | PTGFRN | 14922 | -1.571 | 0.0734 | No | ||
| 141 | CPEB3 | 15022 | -1.598 | 0.0742 | No | ||
| 142 | NR2F2 | 15439 | -1.727 | 0.0583 | No | ||
| 143 | SORCS3 | 15585 | -1.771 | 0.0573 | No | ||
| 144 | MYST3 | 16231 | -2.036 | 0.0302 | No | ||
| 145 | CDH2 | 16417 | -2.124 | 0.0284 | No | ||
| 146 | ATP5G2 | 16435 | -2.128 | 0.0357 | No | ||
| 147 | HOXA10 | 16854 | -2.351 | 0.0221 | No | ||
| 148 | DLG5 | 16947 | -2.413 | 0.0264 | No | ||
| 149 | PAPPA | 17121 | -2.526 | 0.0268 | No | ||
| 150 | SEMA6A | 17213 | -2.598 | 0.0319 | No | ||
| 151 | AGRN | 17257 | -2.624 | 0.0397 | No | ||
| 152 | PTHLH | 17380 | -2.723 | 0.0436 | No | ||
| 153 | PDE7B | 17587 | -2.892 | 0.0436 | No | ||
| 154 | DGCR2 | 17803 | -3.133 | 0.0440 | No |
