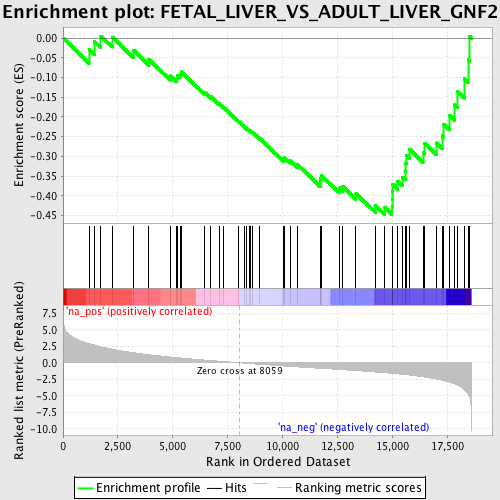
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | set03_absentNotch_versus_truncNotch |
| Phenotype | NoPhenotypeAvailable |
| Upregulated in class | na_neg |
| GeneSet | FETAL_LIVER_VS_ADULT_LIVER_GNF2 |
| Enrichment Score (ES) | -0.4473241 |
| Normalized Enrichment Score (NES) | -1.7510858 |
| Nominal p-value | 0.0056074765 |
| FDR q-value | 0.14274195 |
| FWER p-Value | 0.965 |

| PROBE | GENE SYMBOL | GENE_TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|---|
| 1 | PPP1R1A | 1181 | 2.895 | -0.0282 | No | ||
| 2 | HSD11B1 | 1431 | 2.668 | -0.0089 | No | ||
| 3 | IGF1 | 1722 | 2.417 | 0.0050 | No | ||
| 4 | CYP27A1 | 2236 | 2.062 | 0.0026 | No | ||
| 5 | PKLR | 3209 | 1.519 | -0.0312 | No | ||
| 6 | NRTN | 3906 | 1.216 | -0.0538 | No | ||
| 7 | CES1 | 4888 | 0.871 | -0.0960 | No | ||
| 8 | SLC27A5 | 5165 | 0.784 | -0.1012 | No | ||
| 9 | ITIH4 | 5207 | 0.773 | -0.0940 | No | ||
| 10 | DEXI | 5331 | 0.734 | -0.0916 | No | ||
| 11 | CD81 | 5375 | 0.719 | -0.0852 | No | ||
| 12 | RAB26 | 6440 | 0.415 | -0.1374 | No | ||
| 13 | AQP9 | 6722 | 0.341 | -0.1484 | No | ||
| 14 | LGALS1 | 7107 | 0.235 | -0.1662 | No | ||
| 15 | HGFAC | 7320 | 0.187 | -0.1753 | No | ||
| 16 | ENDOG | 8014 | 0.011 | -0.2125 | No | ||
| 17 | HPD | 8269 | -0.052 | -0.2255 | No | ||
| 18 | GPD1 | 8373 | -0.080 | -0.2301 | No | ||
| 19 | CES2 | 8473 | -0.109 | -0.2341 | No | ||
| 20 | CD14 | 8527 | -0.123 | -0.2354 | No | ||
| 21 | APOC4 | 8622 | -0.144 | -0.2388 | No | ||
| 22 | ABCC3 | 8946 | -0.219 | -0.2535 | No | ||
| 23 | MUCDHL | 10043 | -0.473 | -0.3067 | No | ||
| 24 | APCS | 10099 | -0.488 | -0.3037 | No | ||
| 25 | SDS | 10367 | -0.546 | -0.3114 | No | ||
| 26 | CLPP | 10670 | -0.613 | -0.3202 | No | ||
| 27 | NNMT | 11720 | -0.830 | -0.3665 | No | ||
| 28 | CYP1A2 | 11724 | -0.831 | -0.3565 | No | ||
| 29 | IGFALS | 11766 | -0.839 | -0.3485 | No | ||
| 30 | C1R | 12582 | -1.008 | -0.3800 | No | ||
| 31 | PTGDS | 12731 | -1.038 | -0.3753 | No | ||
| 32 | SLC22A1 | 13348 | -1.168 | -0.3942 | No | ||
| 33 | IFITM1 | 14243 | -1.389 | -0.4253 | No | ||
| 34 | CYP2E1 | 14652 | -1.495 | -0.4290 | Yes | ||
| 35 | SLC25A10 | 14989 | -1.589 | -0.4277 | Yes | ||
| 36 | PCK1 | 15000 | -1.591 | -0.4087 | Yes | ||
| 37 | NKG7 | 15005 | -1.593 | -0.3895 | Yes | ||
| 38 | UPB1 | 15024 | -1.600 | -0.3708 | Yes | ||
| 39 | GSTM2 | 15248 | -1.669 | -0.3624 | Yes | ||
| 40 | GSTM1 | 15467 | -1.735 | -0.3529 | Yes | ||
| 41 | ABHD2 | 15595 | -1.773 | -0.3381 | Yes | ||
| 42 | GSTZ1 | 15621 | -1.782 | -0.3176 | Yes | ||
| 43 | AZGP1 | 15656 | -1.795 | -0.2975 | Yes | ||
| 44 | HAMP | 15782 | -1.848 | -0.2816 | Yes | ||
| 45 | RARRES2 | 16404 | -2.119 | -0.2891 | Yes | ||
| 46 | ACY1 | 16478 | -2.148 | -0.2668 | Yes | ||
| 47 | UQCRC1 | 17024 | -2.462 | -0.2660 | Yes | ||
| 48 | GPR30 | 17273 | -2.636 | -0.2471 | Yes | ||
| 49 | NDUFA7 | 17349 | -2.703 | -0.2180 | Yes | ||
| 50 | ECHS1 | 17622 | -2.941 | -0.1967 | Yes | ||
| 51 | CD151 | 17838 | -3.169 | -0.1695 | Yes | ||
| 52 | SHMT2 | 17979 | -3.373 | -0.1357 | Yes | ||
| 53 | COMT | 18282 | -4.025 | -0.1027 | Yes | ||
| 54 | ASL | 18456 | -4.682 | -0.0547 | Yes | ||
| 55 | TXNRD2 | 18529 | -5.172 | 0.0047 | Yes |
