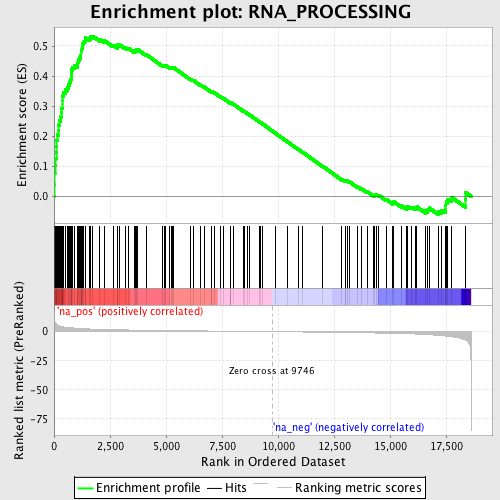
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | set03_absentNotch_versus_normalThy |
| Phenotype | NoPhenotypeAvailable |
| Upregulated in class | na_pos |
| GeneSet | RNA_PROCESSING |
| Enrichment Score (ES) | 0.53285265 |
| Normalized Enrichment Score (NES) | 2.2259927 |
| Nominal p-value | 0.0 |
| FDR q-value | 1.34901E-4 |
| FWER p-Value | 0.0010 |

| PROBE | GENE SYMBOL | GENE_TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|---|
| 1 | NOLA2 | 4 | 11.884 | 0.0404 | Yes | ||
| 2 | FBL | 8 | 10.854 | 0.0773 | Yes | ||
| 3 | SNRPD3 | 41 | 7.780 | 0.1021 | Yes | ||
| 4 | NOLC1 | 54 | 7.270 | 0.1263 | Yes | ||
| 5 | METTL1 | 92 | 6.421 | 0.1462 | Yes | ||
| 6 | U2AF1 | 107 | 6.165 | 0.1665 | Yes | ||
| 7 | PPAN | 114 | 6.041 | 0.1868 | Yes | ||
| 8 | RRP9 | 142 | 5.644 | 0.2047 | Yes | ||
| 9 | SFRS5 | 190 | 5.273 | 0.2201 | Yes | ||
| 10 | NSUN2 | 199 | 5.191 | 0.2374 | Yes | ||
| 11 | SNRP70 | 237 | 4.903 | 0.2522 | Yes | ||
| 12 | PHF5A | 303 | 4.556 | 0.2642 | Yes | ||
| 13 | SNRPA1 | 319 | 4.448 | 0.2786 | Yes | ||
| 14 | NOL5A | 328 | 4.416 | 0.2932 | Yes | ||
| 15 | EXOSC2 | 377 | 4.233 | 0.3051 | Yes | ||
| 16 | PABPC4 | 378 | 4.228 | 0.3195 | Yes | ||
| 17 | POP4 | 381 | 4.221 | 0.3339 | Yes | ||
| 18 | PABPC1 | 419 | 4.086 | 0.3458 | Yes | ||
| 19 | CPSF1 | 494 | 3.855 | 0.3550 | Yes | ||
| 20 | CSTF3 | 580 | 3.645 | 0.3628 | Yes | ||
| 21 | PRPF31 | 644 | 3.505 | 0.3714 | Yes | ||
| 22 | DDX20 | 695 | 3.419 | 0.3804 | Yes | ||
| 23 | DDX39 | 746 | 3.331 | 0.3890 | Yes | ||
| 24 | EFTUD2 | 768 | 3.290 | 0.3991 | Yes | ||
| 25 | TRNT1 | 775 | 3.280 | 0.4100 | Yes | ||
| 26 | AARS | 786 | 3.263 | 0.4206 | Yes | ||
| 27 | SNRPF | 831 | 3.189 | 0.4291 | Yes | ||
| 28 | TXNL4A | 908 | 3.057 | 0.4355 | Yes | ||
| 29 | SFRS2IP | 1051 | 2.871 | 0.4376 | Yes | ||
| 30 | SFRS6 | 1062 | 2.851 | 0.4468 | Yes | ||
| 31 | SYNCRIP | 1090 | 2.817 | 0.4550 | Yes | ||
| 32 | CDC2L2 | 1150 | 2.730 | 0.4611 | Yes | ||
| 33 | DGCR8 | 1188 | 2.679 | 0.4682 | Yes | ||
| 34 | DBR1 | 1200 | 2.664 | 0.4767 | Yes | ||
| 35 | EXOSC7 | 1201 | 2.663 | 0.4858 | Yes | ||
| 36 | IVNS1ABP | 1243 | 2.607 | 0.4925 | Yes | ||
| 37 | GEMIN4 | 1280 | 2.560 | 0.4993 | Yes | ||
| 38 | CRNKL1 | 1284 | 2.557 | 0.5079 | Yes | ||
| 39 | FUSIP1 | 1332 | 2.512 | 0.5139 | Yes | ||
| 40 | SIP1 | 1379 | 2.469 | 0.5199 | Yes | ||
| 41 | LSM4 | 1383 | 2.466 | 0.5281 | Yes | ||
| 42 | SF3A3 | 1561 | 2.293 | 0.5264 | Yes | ||
| 43 | SFRS9 | 1633 | 2.242 | 0.5302 | Yes | ||
| 44 | SNRPD2 | 1723 | 2.181 | 0.5329 | Yes | ||
| 45 | PRPF3 | 2038 | 1.991 | 0.5227 | No | ||
| 46 | NCBP2 | 2232 | 1.883 | 0.5187 | No | ||
| 47 | RNMT | 2631 | 1.696 | 0.5029 | No | ||
| 48 | LSM3 | 2827 | 1.615 | 0.4979 | No | ||
| 49 | SF3A2 | 2828 | 1.615 | 0.5034 | No | ||
| 50 | CSTF2 | 2926 | 1.573 | 0.5036 | No | ||
| 51 | SFRS2 | 3178 | 1.480 | 0.4950 | No | ||
| 52 | HNRPDL | 3313 | 1.432 | 0.4927 | No | ||
| 53 | BICD1 | 3585 | 1.335 | 0.4826 | No | ||
| 54 | CSTF1 | 3635 | 1.316 | 0.4844 | No | ||
| 55 | GRSF1 | 3665 | 1.309 | 0.4873 | No | ||
| 56 | DHX15 | 3729 | 1.288 | 0.4883 | No | ||
| 57 | SFRS10 | 4111 | 1.178 | 0.4718 | No | ||
| 58 | NOL3 | 4842 | 0.995 | 0.4357 | No | ||
| 59 | PPARGC1A | 4930 | 0.974 | 0.4343 | No | ||
| 60 | SF3B4 | 4992 | 0.958 | 0.4343 | No | ||
| 61 | SNRPG | 5166 | 0.921 | 0.4281 | No | ||
| 62 | SNRPD1 | 5228 | 0.904 | 0.4279 | No | ||
| 63 | SSB | 5264 | 0.893 | 0.4290 | No | ||
| 64 | KHDRBS1 | 5317 | 0.881 | 0.4292 | No | ||
| 65 | FARS2 | 6085 | 0.722 | 0.3902 | No | ||
| 66 | HNRPF | 6202 | 0.698 | 0.3863 | No | ||
| 67 | ZNF638 | 6536 | 0.626 | 0.3705 | No | ||
| 68 | KIN | 6694 | 0.595 | 0.3640 | No | ||
| 69 | RBMS1 | 7038 | 0.523 | 0.3473 | No | ||
| 70 | SNW1 | 7044 | 0.522 | 0.3488 | No | ||
| 71 | USP39 | 7160 | 0.500 | 0.3443 | No | ||
| 72 | NUDT21 | 7436 | 0.445 | 0.3309 | No | ||
| 73 | SNRPB | 7548 | 0.425 | 0.3264 | No | ||
| 74 | GEMIN7 | 7862 | 0.362 | 0.3107 | No | ||
| 75 | NUFIP1 | 7876 | 0.359 | 0.3112 | No | ||
| 76 | HNRPD | 7882 | 0.358 | 0.3121 | No | ||
| 77 | SFRS1 | 8028 | 0.331 | 0.3054 | No | ||
| 78 | RBPMS | 8434 | 0.257 | 0.2844 | No | ||
| 79 | GEMIN5 | 8486 | 0.247 | 0.2825 | No | ||
| 80 | RNPS1 | 8649 | 0.214 | 0.2745 | No | ||
| 81 | SRPK2 | 8729 | 0.199 | 0.2709 | No | ||
| 82 | SLBP | 9180 | 0.120 | 0.2470 | No | ||
| 83 | PTBP1 | 9194 | 0.117 | 0.2466 | No | ||
| 84 | BCAS2 | 9291 | 0.098 | 0.2418 | No | ||
| 85 | EXOSC3 | 9889 | -0.030 | 0.2096 | No | ||
| 86 | SPOP | 10432 | -0.154 | 0.1808 | No | ||
| 87 | INTS6 | 10898 | -0.248 | 0.1565 | No | ||
| 88 | RBMY1A1 | 11107 | -0.292 | 0.1463 | No | ||
| 89 | ELAVL4 | 11988 | -0.482 | 0.1004 | No | ||
| 90 | UPF1 | 12847 | -0.691 | 0.0563 | No | ||
| 91 | PRPF4B | 12998 | -0.729 | 0.0507 | No | ||
| 92 | DHX16 | 13011 | -0.731 | 0.0526 | No | ||
| 93 | LSM5 | 13080 | -0.755 | 0.0515 | No | ||
| 94 | DHX38 | 13176 | -0.780 | 0.0490 | No | ||
| 95 | RPS14 | 13536 | -0.882 | 0.0326 | No | ||
| 96 | SNRPB2 | 13743 | -0.945 | 0.0247 | No | ||
| 97 | SF3B3 | 13988 | -1.025 | 0.0150 | No | ||
| 98 | RNGTT | 14253 | -1.121 | 0.0045 | No | ||
| 99 | CPSF3 | 14323 | -1.144 | 0.0047 | No | ||
| 100 | DUSP11 | 14369 | -1.156 | 0.0062 | No | ||
| 101 | SF3B2 | 14497 | -1.205 | 0.0035 | No | ||
| 102 | PRPF8 | 14843 | -1.333 | -0.0106 | No | ||
| 103 | ASCC3L1 | 15121 | -1.452 | -0.0206 | No | ||
| 104 | SFPQ | 15167 | -1.472 | -0.0180 | No | ||
| 105 | RBM6 | 15522 | -1.652 | -0.0315 | No | ||
| 106 | SRRM1 | 15749 | -1.783 | -0.0377 | No | ||
| 107 | RBM5 | 15774 | -1.801 | -0.0328 | No | ||
| 108 | NONO | 15942 | -1.903 | -0.0354 | No | ||
| 109 | SF3A1 | 16122 | -2.027 | -0.0381 | No | ||
| 110 | SFRS7 | 16198 | -2.096 | -0.0350 | No | ||
| 111 | GEMIN6 | 16586 | -2.446 | -0.0476 | No | ||
| 112 | SRPK1 | 16686 | -2.540 | -0.0443 | No | ||
| 113 | IGF2BP3 | 16744 | -2.611 | -0.0384 | No | ||
| 114 | DHX8 | 17147 | -3.141 | -0.0494 | No | ||
| 115 | RBM3 | 17295 | -3.388 | -0.0458 | No | ||
| 116 | DDX54 | 17468 | -3.728 | -0.0424 | No | ||
| 117 | ADAT1 | 17471 | -3.735 | -0.0297 | No | ||
| 118 | HNRPUL1 | 17494 | -3.811 | -0.0179 | No | ||
| 119 | SFRS8 | 17576 | -3.965 | -0.0087 | No | ||
| 120 | CUGBP1 | 17752 | -4.386 | -0.0032 | No | ||
| 121 | SNRPN | 18357 | -7.236 | -0.0112 | No | ||
| 122 | ZNF346 | 18373 | -7.351 | 0.0131 | No |
