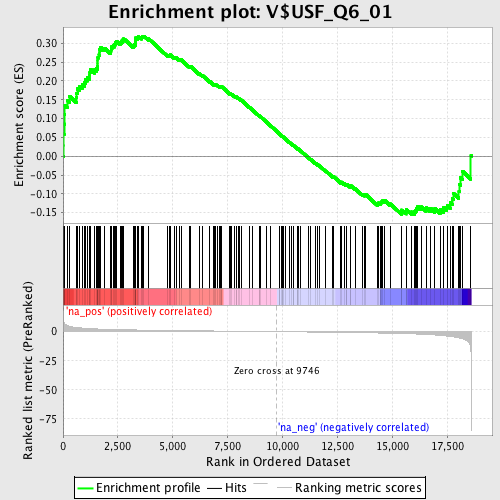
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | set03_absentNotch_versus_normalThy |
| Phenotype | NoPhenotypeAvailable |
| Upregulated in class | na_pos |
| GeneSet | V$USF_Q6_01 |
| Enrichment Score (ES) | 0.319071 |
| Normalized Enrichment Score (NES) | 1.3884776 |
| Nominal p-value | 0.011494253 |
| FDR q-value | 0.18701306 |
| FWER p-Value | 0.994 |

| PROBE | GENE SYMBOL | GENE_TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|---|
| 1 | EIF5A | 26 | 8.461 | 0.0294 | Yes | ||
| 2 | BAX | 28 | 8.257 | 0.0594 | Yes | ||
| 3 | HMOX1 | 47 | 7.454 | 0.0856 | Yes | ||
| 4 | APEX1 | 57 | 7.195 | 0.1113 | Yes | ||
| 5 | PFDN2 | 74 | 6.806 | 0.1352 | Yes | ||
| 6 | EIF3S9 | 187 | 5.303 | 0.1485 | Yes | ||
| 7 | AATF | 299 | 4.568 | 0.1591 | Yes | ||
| 8 | ACACA | 606 | 3.579 | 0.1556 | Yes | ||
| 9 | GTF2A1 | 621 | 3.547 | 0.1677 | Yes | ||
| 10 | AKAP12 | 650 | 3.499 | 0.1789 | Yes | ||
| 11 | MRPL27 | 751 | 3.320 | 0.1856 | Yes | ||
| 12 | EIF4E | 869 | 3.123 | 0.1907 | Yes | ||
| 13 | ABR | 982 | 2.945 | 0.1953 | Yes | ||
| 14 | CLN3 | 1012 | 2.914 | 0.2044 | Yes | ||
| 15 | RANBP1 | 1098 | 2.805 | 0.2100 | Yes | ||
| 16 | HEXA | 1207 | 2.652 | 0.2138 | Yes | ||
| 17 | SYT13 | 1219 | 2.638 | 0.2228 | Yes | ||
| 18 | SSR1 | 1245 | 2.606 | 0.2309 | Yes | ||
| 19 | RPS2 | 1435 | 2.416 | 0.2295 | Yes | ||
| 20 | RAB3IL1 | 1514 | 2.334 | 0.2338 | Yes | ||
| 21 | IRS4 | 1568 | 2.290 | 0.2392 | Yes | ||
| 22 | PER1 | 1581 | 2.278 | 0.2469 | Yes | ||
| 23 | KLF4 | 1583 | 2.277 | 0.2551 | Yes | ||
| 24 | SFXN2 | 1586 | 2.276 | 0.2633 | Yes | ||
| 25 | NR3C2 | 1634 | 2.242 | 0.2689 | Yes | ||
| 26 | VLDLR | 1667 | 2.221 | 0.2753 | Yes | ||
| 27 | SNTB2 | 1670 | 2.219 | 0.2832 | Yes | ||
| 28 | KRTCAP2 | 1711 | 2.189 | 0.2890 | Yes | ||
| 29 | CCAR1 | 1883 | 2.075 | 0.2873 | Yes | ||
| 30 | GTF2H1 | 2175 | 1.914 | 0.2786 | Yes | ||
| 31 | ETV1 | 2183 | 1.911 | 0.2851 | Yes | ||
| 32 | GADD45G | 2212 | 1.894 | 0.2905 | Yes | ||
| 33 | KCTD15 | 2273 | 1.857 | 0.2940 | Yes | ||
| 34 | SYT3 | 2348 | 1.827 | 0.2967 | Yes | ||
| 35 | SC5DL | 2373 | 1.815 | 0.3020 | Yes | ||
| 36 | EEF1B2 | 2420 | 1.789 | 0.3060 | Yes | ||
| 37 | UBXD3 | 2601 | 1.714 | 0.3025 | Yes | ||
| 38 | STMN1 | 2649 | 1.690 | 0.3061 | Yes | ||
| 39 | COPS7A | 2711 | 1.662 | 0.3089 | Yes | ||
| 40 | SGK | 2769 | 1.637 | 0.3117 | Yes | ||
| 41 | LTBR | 3198 | 1.474 | 0.2939 | Yes | ||
| 42 | CHD4 | 3237 | 1.460 | 0.2972 | Yes | ||
| 43 | EIF4G1 | 3294 | 1.441 | 0.2994 | Yes | ||
| 44 | POGK | 3300 | 1.437 | 0.3044 | Yes | ||
| 45 | RAPGEF3 | 3308 | 1.434 | 0.3092 | Yes | ||
| 46 | HNRPDL | 3313 | 1.432 | 0.3142 | Yes | ||
| 47 | HOXA3 | 3392 | 1.402 | 0.3151 | Yes | ||
| 48 | COMMD3 | 3444 | 1.382 | 0.3174 | Yes | ||
| 49 | SOCS5 | 3558 | 1.346 | 0.3161 | Yes | ||
| 50 | ARTN | 3595 | 1.329 | 0.3190 | Yes | ||
| 51 | HTF9C | 3683 | 1.304 | 0.3191 | Yes | ||
| 52 | HMGN2 | 3884 | 1.241 | 0.3128 | No | ||
| 53 | NRAS | 4773 | 1.011 | 0.2683 | No | ||
| 54 | ADAM10 | 4832 | 0.997 | 0.2688 | No | ||
| 55 | SCHIP1 | 4887 | 0.985 | 0.2695 | No | ||
| 56 | SLC26A2 | 5068 | 0.943 | 0.2632 | No | ||
| 57 | TAF6L | 5153 | 0.923 | 0.2620 | No | ||
| 58 | FGF5 | 5293 | 0.886 | 0.2577 | No | ||
| 59 | HOXA9 | 5376 | 0.866 | 0.2564 | No | ||
| 60 | ANKRD17 | 5746 | 0.790 | 0.2393 | No | ||
| 61 | NEUROD2 | 5814 | 0.776 | 0.2385 | No | ||
| 62 | DNMT3A | 6216 | 0.696 | 0.2193 | No | ||
| 63 | MYST3 | 6354 | 0.664 | 0.2143 | No | ||
| 64 | PICALM | 6681 | 0.598 | 0.1988 | No | ||
| 65 | STC2 | 6866 | 0.559 | 0.1909 | No | ||
| 66 | HOXD10 | 6913 | 0.548 | 0.1904 | No | ||
| 67 | CGN | 6929 | 0.544 | 0.1916 | No | ||
| 68 | SLC16A1 | 7026 | 0.525 | 0.1883 | No | ||
| 69 | PCDHA10 | 7146 | 0.502 | 0.1837 | No | ||
| 70 | HOXC5 | 7170 | 0.497 | 0.1842 | No | ||
| 71 | HNRPH2 | 7224 | 0.487 | 0.1831 | No | ||
| 72 | PTMA | 7233 | 0.486 | 0.1845 | No | ||
| 73 | HOXC12 | 7600 | 0.412 | 0.1661 | No | ||
| 74 | GLA | 7614 | 0.411 | 0.1669 | No | ||
| 75 | NR5A1 | 7693 | 0.397 | 0.1642 | No | ||
| 76 | RNF12 | 7821 | 0.371 | 0.1586 | No | ||
| 77 | HNRPD | 7882 | 0.358 | 0.1567 | No | ||
| 78 | APLN | 7885 | 0.357 | 0.1579 | No | ||
| 79 | EN1 | 7971 | 0.341 | 0.1545 | No | ||
| 80 | LIN28 | 8035 | 0.329 | 0.1523 | No | ||
| 81 | SIRT1 | 8112 | 0.315 | 0.1493 | No | ||
| 82 | WEE1 | 8488 | 0.246 | 0.1299 | No | ||
| 83 | PANK3 | 8627 | 0.216 | 0.1232 | No | ||
| 84 | NPM1 | 8937 | 0.166 | 0.1071 | No | ||
| 85 | NPTX1 | 8966 | 0.160 | 0.1062 | No | ||
| 86 | SLC1A7 | 8982 | 0.157 | 0.1059 | No | ||
| 87 | ZFX | 9266 | 0.104 | 0.0910 | No | ||
| 88 | CDH1 | 9473 | 0.061 | 0.0800 | No | ||
| 89 | TRIB1 | 9842 | -0.020 | 0.0602 | No | ||
| 90 | HOOK2 | 9850 | -0.022 | 0.0599 | No | ||
| 91 | UTP14C | 9956 | -0.048 | 0.0544 | No | ||
| 92 | BHLHB3 | 9995 | -0.058 | 0.0525 | No | ||
| 93 | GNA13 | 10003 | -0.059 | 0.0524 | No | ||
| 94 | AGRP | 10035 | -0.066 | 0.0509 | No | ||
| 95 | HOXB6 | 10122 | -0.084 | 0.0466 | No | ||
| 96 | NEUROD1 | 10311 | -0.126 | 0.0369 | No | ||
| 97 | ELOVL3 | 10424 | -0.153 | 0.0313 | No | ||
| 98 | MAX | 10487 | -0.167 | 0.0286 | No | ||
| 99 | ATP6V1H | 10511 | -0.170 | 0.0280 | No | ||
| 100 | LAMP1 | 10676 | -0.201 | 0.0198 | No | ||
| 101 | RHOQ | 10696 | -0.205 | 0.0195 | No | ||
| 102 | KCNE4 | 10712 | -0.207 | 0.0195 | No | ||
| 103 | TLL1 | 10831 | -0.234 | 0.0139 | No | ||
| 104 | FGF14 | 11201 | -0.310 | -0.0049 | No | ||
| 105 | SLC31A2 | 11293 | -0.330 | -0.0087 | No | ||
| 106 | JUP | 11496 | -0.372 | -0.0182 | No | ||
| 107 | EGLN2 | 11601 | -0.395 | -0.0224 | No | ||
| 108 | IPO7 | 11662 | -0.410 | -0.0242 | No | ||
| 109 | NFX1 | 11979 | -0.480 | -0.0396 | No | ||
| 110 | TGIF2 | 12299 | -0.556 | -0.0548 | No | ||
| 111 | STX6 | 12333 | -0.565 | -0.0545 | No | ||
| 112 | WDR20 | 12664 | -0.643 | -0.0701 | No | ||
| 113 | SEPT3 | 12680 | -0.647 | -0.0685 | No | ||
| 114 | GNB2 | 12806 | -0.679 | -0.0728 | No | ||
| 115 | BDNF | 12902 | -0.703 | -0.0754 | No | ||
| 116 | PPM1A | 12935 | -0.712 | -0.0746 | No | ||
| 117 | RAMP2 | 13084 | -0.756 | -0.0798 | No | ||
| 118 | DCTN4 | 13091 | -0.757 | -0.0774 | No | ||
| 119 | MCOLN1 | 13318 | -0.815 | -0.0867 | No | ||
| 120 | NCKIPSD | 13659 | -0.919 | -0.1017 | No | ||
| 121 | ATP7B | 13738 | -0.943 | -0.1025 | No | ||
| 122 | HOXA1 | 13771 | -0.955 | -0.1008 | No | ||
| 123 | ILF3 | 14335 | -1.146 | -0.1271 | No | ||
| 124 | MPV17 | 14359 | -1.153 | -0.1241 | No | ||
| 125 | TAC1 | 14458 | -1.189 | -0.1251 | No | ||
| 126 | NDUFS1 | 14487 | -1.202 | -0.1223 | No | ||
| 127 | HIF1A | 14529 | -1.213 | -0.1201 | No | ||
| 128 | DUSP7 | 14571 | -1.226 | -0.1178 | No | ||
| 129 | TRIM46 | 14653 | -1.256 | -0.1176 | No | ||
| 130 | ZNF318 | 14915 | -1.364 | -0.1268 | No | ||
| 131 | WDR23 | 15429 | -1.609 | -0.1487 | No | ||
| 132 | HPGD | 15431 | -1.610 | -0.1429 | No | ||
| 133 | CENTB5 | 15630 | -1.712 | -0.1474 | No | ||
| 134 | UBE2B | 15649 | -1.725 | -0.1421 | No | ||
| 135 | HOXB7 | 15886 | -1.873 | -0.1480 | No | ||
| 136 | CSRP2BP | 16008 | -1.942 | -0.1475 | No | ||
| 137 | SMYD4 | 16061 | -1.973 | -0.1432 | No | ||
| 138 | RPL19 | 16118 | -2.024 | -0.1388 | No | ||
| 139 | ARL3 | 16165 | -2.072 | -0.1338 | No | ||
| 140 | HRH3 | 16331 | -2.214 | -0.1346 | No | ||
| 141 | ATF7IP | 16541 | -2.408 | -0.1372 | No | ||
| 142 | TCF7 | 16737 | -2.603 | -0.1383 | No | ||
| 143 | IGF2R | 16943 | -2.855 | -0.1390 | No | ||
| 144 | LZTS2 | 17184 | -3.207 | -0.1403 | No | ||
| 145 | VPS16 | 17345 | -3.485 | -0.1363 | No | ||
| 146 | GNAS | 17496 | -3.817 | -0.1305 | No | ||
| 147 | AXUD1 | 17645 | -4.123 | -0.1235 | No | ||
| 148 | CNOT4 | 17745 | -4.368 | -0.1130 | No | ||
| 149 | WBP2 | 17796 | -4.515 | -0.0992 | No | ||
| 150 | KBTBD2 | 18041 | -5.284 | -0.0932 | No | ||
| 151 | WDR1 | 18069 | -5.373 | -0.0751 | No | ||
| 152 | RGL2 | 18106 | -5.554 | -0.0568 | No | ||
| 153 | RNF44 | 18193 | -5.987 | -0.0397 | No | ||
| 154 | HOXA7 | 18589 | -17.167 | 0.0015 | No |
