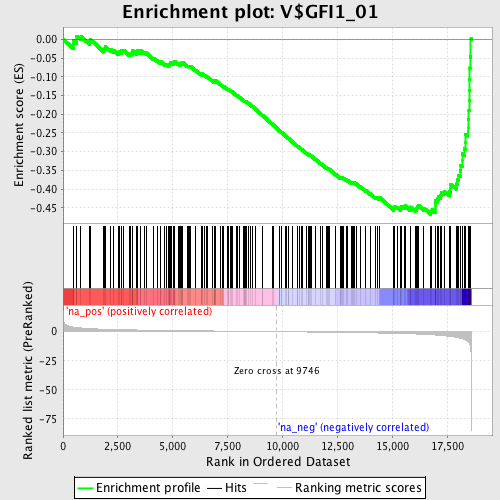
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | set03_absentNotch_versus_normalThy |
| Phenotype | NoPhenotypeAvailable |
| Upregulated in class | na_neg |
| GeneSet | V$GFI1_01 |
| Enrichment Score (ES) | -0.46755293 |
| Normalized Enrichment Score (NES) | -1.7198952 |
| Nominal p-value | 0.0011520737 |
| FDR q-value | 0.08873499 |
| FWER p-Value | 0.768 |

| PROBE | GENE SYMBOL | GENE_TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|---|
| 1 | ATP5G2 | 470 | 3.913 | -0.0143 | No | ||
| 2 | SREBF2 | 483 | 3.872 | -0.0039 | No | ||
| 3 | TMPRSS2 | 626 | 3.544 | -0.0015 | No | ||
| 4 | ABCF2 | 627 | 3.544 | 0.0086 | No | ||
| 5 | CNKSR1 | 813 | 3.209 | 0.0077 | No | ||
| 6 | CSAD | 1217 | 2.639 | -0.0067 | No | ||
| 7 | MYO1C | 1230 | 2.624 | 0.0002 | No | ||
| 8 | DCX | 1860 | 2.087 | -0.0280 | No | ||
| 9 | PDC | 1904 | 2.064 | -0.0244 | No | ||
| 10 | USP5 | 1914 | 2.058 | -0.0191 | No | ||
| 11 | HMGA1 | 2154 | 1.926 | -0.0265 | No | ||
| 12 | MYH10 | 2317 | 1.841 | -0.0301 | No | ||
| 13 | ICAM5 | 2526 | 1.743 | -0.0364 | No | ||
| 14 | CDH13 | 2547 | 1.734 | -0.0325 | No | ||
| 15 | STMN1 | 2649 | 1.690 | -0.0332 | No | ||
| 16 | SLC25A18 | 2675 | 1.680 | -0.0297 | No | ||
| 17 | TOP1 | 2747 | 1.646 | -0.0289 | No | ||
| 18 | BAI3 | 3027 | 1.533 | -0.0397 | No | ||
| 19 | YWHAE | 3091 | 1.513 | -0.0388 | No | ||
| 20 | TPBG | 3149 | 1.492 | -0.0376 | No | ||
| 21 | SFRS2 | 3178 | 1.480 | -0.0349 | No | ||
| 22 | CNTN6 | 3180 | 1.480 | -0.0307 | No | ||
| 23 | TCF12 | 3355 | 1.417 | -0.0361 | No | ||
| 24 | HOXA3 | 3392 | 1.402 | -0.0341 | No | ||
| 25 | EXOC7 | 3393 | 1.402 | -0.0301 | No | ||
| 26 | TRIM23 | 3526 | 1.357 | -0.0334 | No | ||
| 27 | MCTS1 | 3536 | 1.355 | -0.0300 | No | ||
| 28 | GAS2 | 3689 | 1.302 | -0.0345 | No | ||
| 29 | AVP | 3789 | 1.270 | -0.0363 | No | ||
| 30 | BRUNOL4 | 4114 | 1.177 | -0.0505 | No | ||
| 31 | RPS6KB1 | 4288 | 1.133 | -0.0567 | No | ||
| 32 | FOXP2 | 4445 | 1.089 | -0.0620 | No | ||
| 33 | ZIC3 | 4454 | 1.087 | -0.0593 | No | ||
| 34 | MAP2K7 | 4635 | 1.048 | -0.0661 | No | ||
| 35 | NFIA | 4728 | 1.021 | -0.0682 | No | ||
| 36 | JPH4 | 4808 | 1.003 | -0.0696 | No | ||
| 37 | DNAJB4 | 4835 | 0.997 | -0.0682 | No | ||
| 38 | PBX1 | 4846 | 0.994 | -0.0659 | No | ||
| 39 | XPO5 | 4870 | 0.989 | -0.0643 | No | ||
| 40 | NRXN3 | 4871 | 0.989 | -0.0615 | No | ||
| 41 | PPARGC1A | 4930 | 0.974 | -0.0619 | No | ||
| 42 | OTP | 5019 | 0.954 | -0.0639 | No | ||
| 43 | DPF3 | 5037 | 0.950 | -0.0621 | No | ||
| 44 | NCAM1 | 5087 | 0.939 | -0.0621 | No | ||
| 45 | ZFPM2 | 5090 | 0.938 | -0.0596 | No | ||
| 46 | ZIC1 | 5244 | 0.898 | -0.0653 | No | ||
| 47 | CACNB2 | 5311 | 0.882 | -0.0664 | No | ||
| 48 | GPR20 | 5364 | 0.869 | -0.0667 | No | ||
| 49 | HOXA9 | 5376 | 0.866 | -0.0648 | No | ||
| 50 | ICAM4 | 5391 | 0.864 | -0.0631 | No | ||
| 51 | DDR2 | 5444 | 0.852 | -0.0635 | No | ||
| 52 | PRIMA1 | 5461 | 0.849 | -0.0620 | No | ||
| 53 | ANGPT1 | 5675 | 0.805 | -0.0712 | No | ||
| 54 | MBNL1 | 5726 | 0.794 | -0.0717 | No | ||
| 55 | PI15 | 5763 | 0.786 | -0.0714 | No | ||
| 56 | TLK1 | 5802 | 0.779 | -0.0712 | No | ||
| 57 | SOX9 | 6052 | 0.727 | -0.0827 | No | ||
| 58 | CLMN | 6290 | 0.679 | -0.0936 | No | ||
| 59 | PPARGC1B | 6312 | 0.674 | -0.0928 | No | ||
| 60 | CDH6 | 6337 | 0.668 | -0.0922 | No | ||
| 61 | ASB4 | 6442 | 0.644 | -0.0960 | No | ||
| 62 | SLC4A7 | 6545 | 0.625 | -0.0998 | No | ||
| 63 | NGEF | 6596 | 0.616 | -0.1007 | No | ||
| 64 | PRKAG1 | 6793 | 0.571 | -0.1097 | No | ||
| 65 | HOXB8 | 6884 | 0.554 | -0.1130 | No | ||
| 66 | ITGA10 | 6911 | 0.548 | -0.1129 | No | ||
| 67 | HOXD10 | 6913 | 0.548 | -0.1114 | No | ||
| 68 | CAMK2D | 6927 | 0.544 | -0.1105 | No | ||
| 69 | CGN | 6929 | 0.544 | -0.1090 | No | ||
| 70 | BNC2 | 7154 | 0.501 | -0.1197 | No | ||
| 71 | HOXC8 | 7277 | 0.478 | -0.1250 | No | ||
| 72 | C1QTNF7 | 7330 | 0.465 | -0.1265 | No | ||
| 73 | TCF4 | 7482 | 0.437 | -0.1334 | No | ||
| 74 | PROCR | 7544 | 0.426 | -0.1355 | No | ||
| 75 | SERTAD4 | 7554 | 0.424 | -0.1348 | No | ||
| 76 | PCDH8 | 7642 | 0.406 | -0.1384 | No | ||
| 77 | CRYZL1 | 7683 | 0.400 | -0.1394 | No | ||
| 78 | CACNG3 | 7710 | 0.392 | -0.1397 | No | ||
| 79 | HOXD11 | 7903 | 0.353 | -0.1491 | No | ||
| 80 | KLF15 | 7969 | 0.341 | -0.1517 | No | ||
| 81 | MTMR4 | 8046 | 0.328 | -0.1549 | No | ||
| 82 | EBF2 | 8223 | 0.296 | -0.1636 | No | ||
| 83 | DEPDC2 | 8259 | 0.290 | -0.1646 | No | ||
| 84 | DSCR1L1 | 8283 | 0.286 | -0.1651 | No | ||
| 85 | HIST1H1E | 8305 | 0.282 | -0.1654 | No | ||
| 86 | TBR1 | 8315 | 0.281 | -0.1651 | No | ||
| 87 | MEF2C | 8378 | 0.266 | -0.1677 | No | ||
| 88 | ITSN1 | 8453 | 0.255 | -0.1710 | No | ||
| 89 | SH3BGRL2 | 8525 | 0.238 | -0.1742 | No | ||
| 90 | SP8 | 8618 | 0.217 | -0.1785 | No | ||
| 91 | GPR17 | 8780 | 0.191 | -0.1867 | No | ||
| 92 | HOXC4 | 9075 | 0.139 | -0.2023 | No | ||
| 93 | TFEC | 9104 | 0.133 | -0.2034 | No | ||
| 94 | PLCD4 | 9521 | 0.052 | -0.2259 | No | ||
| 95 | CALN1 | 9580 | 0.039 | -0.2289 | No | ||
| 96 | HOXB3 | 9844 | -0.021 | -0.2431 | No | ||
| 97 | HIST1H1T | 9877 | -0.028 | -0.2448 | No | ||
| 98 | PURA | 9884 | -0.030 | -0.2450 | No | ||
| 99 | PEA15 | 9951 | -0.047 | -0.2485 | No | ||
| 100 | GPRC5D | 9974 | -0.053 | -0.2495 | No | ||
| 101 | HOXB6 | 10122 | -0.084 | -0.2572 | No | ||
| 102 | DSPP | 10184 | -0.098 | -0.2603 | No | ||
| 103 | PTHLH | 10250 | -0.112 | -0.2635 | No | ||
| 104 | SOX15 | 10266 | -0.117 | -0.2640 | No | ||
| 105 | PHF8 | 10437 | -0.155 | -0.2727 | No | ||
| 106 | GRK6 | 10681 | -0.202 | -0.2854 | No | ||
| 107 | SPAG7 | 10703 | -0.206 | -0.2859 | No | ||
| 108 | GRPR | 10767 | -0.220 | -0.2887 | No | ||
| 109 | TFB1M | 10881 | -0.245 | -0.2941 | No | ||
| 110 | GFRA3 | 10892 | -0.247 | -0.2940 | No | ||
| 111 | BMP4 | 11108 | -0.292 | -0.3048 | No | ||
| 112 | RKHD3 | 11185 | -0.308 | -0.3080 | No | ||
| 113 | SGK2 | 11190 | -0.309 | -0.3074 | No | ||
| 114 | SLC26A3 | 11217 | -0.314 | -0.3079 | No | ||
| 115 | BCL9 | 11279 | -0.327 | -0.3103 | No | ||
| 116 | CDON | 11321 | -0.335 | -0.3115 | No | ||
| 117 | HOXA6 | 11494 | -0.371 | -0.3198 | No | ||
| 118 | POU3F1 | 11725 | -0.424 | -0.3311 | No | ||
| 119 | SOSTDC1 | 11837 | -0.448 | -0.3358 | No | ||
| 120 | ELAVL4 | 11988 | -0.482 | -0.3426 | No | ||
| 121 | CDX2 | 12059 | -0.496 | -0.3450 | No | ||
| 122 | MRGPRF | 12093 | -0.504 | -0.3453 | No | ||
| 123 | CHRM1 | 12158 | -0.520 | -0.3473 | No | ||
| 124 | PHYHIP | 12436 | -0.589 | -0.3607 | No | ||
| 125 | PHACTR3 | 12640 | -0.637 | -0.3699 | No | ||
| 126 | MSR1 | 12644 | -0.637 | -0.3682 | No | ||
| 127 | RARB | 12667 | -0.644 | -0.3676 | No | ||
| 128 | EDN1 | 12729 | -0.661 | -0.3690 | No | ||
| 129 | CREB5 | 12801 | -0.678 | -0.3709 | No | ||
| 130 | BDNF | 12902 | -0.703 | -0.3744 | No | ||
| 131 | NEXN | 12976 | -0.723 | -0.3763 | No | ||
| 132 | GREM1 | 13152 | -0.775 | -0.3835 | No | ||
| 133 | NKX2-8 | 13175 | -0.780 | -0.3825 | No | ||
| 134 | HOXA10 | 13212 | -0.788 | -0.3822 | No | ||
| 135 | PDLIM1 | 13266 | -0.801 | -0.3828 | No | ||
| 136 | DSG3 | 13361 | -0.829 | -0.3855 | No | ||
| 137 | TLR1 | 13532 | -0.880 | -0.3923 | No | ||
| 138 | SOX5 | 13777 | -0.957 | -0.4028 | No | ||
| 139 | NR2F2 | 14000 | -1.030 | -0.4119 | No | ||
| 140 | RKHD1 | 14237 | -1.113 | -0.4215 | No | ||
| 141 | TCF1 | 14307 | -1.137 | -0.4220 | No | ||
| 142 | LUZP1 | 14405 | -1.170 | -0.4239 | No | ||
| 143 | MEIS1 | 14412 | -1.172 | -0.4209 | No | ||
| 144 | SOX4 | 15077 | -1.432 | -0.4529 | No | ||
| 145 | IRX3 | 15078 | -1.433 | -0.4488 | No | ||
| 146 | TRDN | 15123 | -1.452 | -0.4470 | No | ||
| 147 | CPEB3 | 15222 | -1.495 | -0.4481 | No | ||
| 148 | ARHGEF2 | 15378 | -1.586 | -0.4520 | No | ||
| 149 | CTLA4 | 15398 | -1.593 | -0.4484 | No | ||
| 150 | WDR23 | 15429 | -1.609 | -0.4455 | No | ||
| 151 | MAP2K5 | 15558 | -1.674 | -0.4476 | No | ||
| 152 | TSGA10 | 15584 | -1.690 | -0.4442 | No | ||
| 153 | RORA | 15836 | -1.839 | -0.4526 | No | ||
| 154 | PTEN | 15842 | -1.843 | -0.4476 | No | ||
| 155 | LAMA3 | 16048 | -1.969 | -0.4531 | No | ||
| 156 | BCL9L | 16103 | -2.004 | -0.4503 | No | ||
| 157 | HNRPA0 | 16132 | -2.037 | -0.4460 | No | ||
| 158 | PREP | 16203 | -2.098 | -0.4438 | No | ||
| 159 | ETS1 | 16445 | -2.310 | -0.4503 | No | ||
| 160 | KLF12 | 16764 | -2.638 | -0.4600 | Yes | ||
| 161 | HIVEP3 | 16812 | -2.695 | -0.4549 | Yes | ||
| 162 | TCTA | 16967 | -2.886 | -0.4550 | Yes | ||
| 163 | JMJD1C | 16969 | -2.887 | -0.4468 | Yes | ||
| 164 | MAP4K3 | 16972 | -2.892 | -0.4387 | Yes | ||
| 165 | ATF2 | 16984 | -2.909 | -0.4310 | Yes | ||
| 166 | ELMO3 | 17053 | -2.996 | -0.4261 | Yes | ||
| 167 | ADD3 | 17108 | -3.081 | -0.4203 | Yes | ||
| 168 | IRF2BP1 | 17203 | -3.241 | -0.4161 | Yes | ||
| 169 | RAB6B | 17230 | -3.281 | -0.4082 | Yes | ||
| 170 | PPP2R2A | 17365 | -3.532 | -0.4053 | Yes | ||
| 171 | GABARAP | 17603 | -4.008 | -0.4068 | Yes | ||
| 172 | POLH | 17664 | -4.165 | -0.3981 | Yes | ||
| 173 | AGPAT4 | 17669 | -4.180 | -0.3864 | Yes | ||
| 174 | CREM | 17917 | -4.880 | -0.3859 | Yes | ||
| 175 | BPGM | 17961 | -4.996 | -0.3740 | Yes | ||
| 176 | MAP3K3 | 18023 | -5.201 | -0.3625 | Yes | ||
| 177 | BRD2 | 18095 | -5.480 | -0.3507 | Yes | ||
| 178 | ZIC4 | 18117 | -5.585 | -0.3359 | Yes | ||
| 179 | PANK1 | 18186 | -5.948 | -0.3226 | Yes | ||
| 180 | RHOA | 18189 | -5.967 | -0.3057 | Yes | ||
| 181 | HSD11B1 | 18294 | -6.651 | -0.2923 | Yes | ||
| 182 | HBP1 | 18344 | -7.095 | -0.2747 | Yes | ||
| 183 | PDE4D | 18349 | -7.162 | -0.2545 | Yes | ||
| 184 | TRAF4 | 18453 | -8.305 | -0.2364 | Yes | ||
| 185 | MAP4K2 | 18455 | -8.325 | -0.2127 | Yes | ||
| 186 | GADD45A | 18496 | -9.058 | -0.1890 | Yes | ||
| 187 | BCL6 | 18511 | -9.435 | -0.1629 | Yes | ||
| 188 | CCR7 | 18521 | -9.794 | -0.1354 | Yes | ||
| 189 | BTG1 | 18531 | -10.306 | -0.1065 | Yes | ||
| 190 | EVA1 | 18540 | -10.654 | -0.0765 | Yes | ||
| 191 | IL16 | 18549 | -11.052 | -0.0454 | Yes | ||
| 192 | HOXA7 | 18589 | -17.167 | 0.0015 | Yes |
