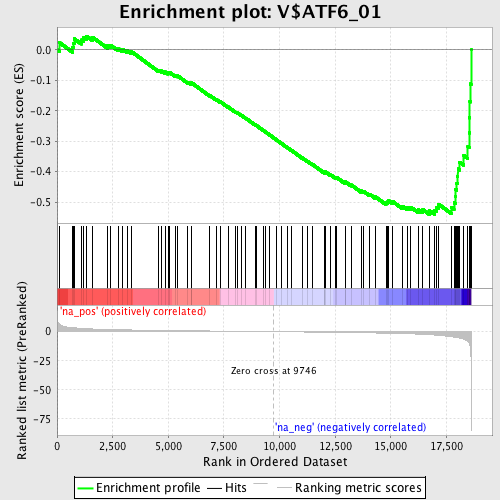
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | set03_absentNotch_versus_normalThy |
| Phenotype | NoPhenotypeAvailable |
| Upregulated in class | na_neg |
| GeneSet | V$ATF6_01 |
| Enrichment Score (ES) | -0.5406688 |
| Normalized Enrichment Score (NES) | -1.7928684 |
| Nominal p-value | 0.0025740026 |
| FDR q-value | 0.11024182 |
| FWER p-Value | 0.54 |

| PROBE | GENE SYMBOL | GENE_TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|---|
| 1 | MRPS18B | 106 | 6.167 | 0.0238 | No | ||
| 2 | KDELR2 | 698 | 3.414 | 0.0082 | No | ||
| 3 | GRB2 | 736 | 3.346 | 0.0223 | No | ||
| 4 | EIF4A1 | 769 | 3.290 | 0.0363 | No | ||
| 5 | SYNCRIP | 1090 | 2.817 | 0.0325 | No | ||
| 6 | TPM4 | 1183 | 2.688 | 0.0404 | No | ||
| 7 | KDELR3 | 1321 | 2.523 | 0.0451 | No | ||
| 8 | PER1 | 1581 | 2.278 | 0.0420 | No | ||
| 9 | STAG1 | 2259 | 1.867 | 0.0144 | No | ||
| 10 | GOLGB1 | 2413 | 1.793 | 0.0147 | No | ||
| 11 | TOP1 | 2747 | 1.646 | 0.0047 | No | ||
| 12 | KDELR1 | 2929 | 1.572 | 0.0024 | No | ||
| 13 | TYRO3 | 3165 | 1.485 | -0.0032 | No | ||
| 14 | EGR3 | 3322 | 1.429 | -0.0047 | No | ||
| 15 | DLST | 4566 | 1.062 | -0.0667 | No | ||
| 16 | HCRTR2 | 4702 | 1.028 | -0.0691 | No | ||
| 17 | FLNC | 4854 | 0.991 | -0.0725 | No | ||
| 18 | TWIST1 | 4985 | 0.959 | -0.0749 | No | ||
| 19 | ATBF1 | 5036 | 0.950 | -0.0731 | No | ||
| 20 | NANS | 5322 | 0.880 | -0.0842 | No | ||
| 21 | ICA1 | 5415 | 0.859 | -0.0851 | No | ||
| 22 | TLE3 | 5880 | 0.763 | -0.1065 | No | ||
| 23 | USP2 | 6024 | 0.731 | -0.1107 | No | ||
| 24 | XPNPEP1 | 6047 | 0.728 | -0.1084 | No | ||
| 25 | NKX2-2 | 6855 | 0.561 | -0.1493 | No | ||
| 26 | LMX1B | 7167 | 0.498 | -0.1637 | No | ||
| 27 | C1QTNF7 | 7330 | 0.465 | -0.1702 | No | ||
| 28 | SRP19 | 7687 | 0.399 | -0.1875 | No | ||
| 29 | SFRS1 | 8028 | 0.331 | -0.2042 | No | ||
| 30 | LMO4 | 8110 | 0.315 | -0.2071 | No | ||
| 31 | DUSP4 | 8297 | 0.283 | -0.2158 | No | ||
| 32 | TLOC1 | 8475 | 0.249 | -0.2241 | No | ||
| 33 | GRIN2B | 8897 | 0.172 | -0.2460 | No | ||
| 34 | NPTX1 | 8966 | 0.160 | -0.2489 | No | ||
| 35 | GCC1 | 9262 | 0.105 | -0.2643 | No | ||
| 36 | NTRK2 | 9358 | 0.084 | -0.2691 | No | ||
| 37 | VCP | 9566 | 0.041 | -0.2800 | No | ||
| 38 | ARF4 | 9848 | -0.021 | -0.2951 | No | ||
| 39 | GSC | 10070 | -0.073 | -0.3067 | No | ||
| 40 | DHX40 | 10339 | -0.135 | -0.3205 | No | ||
| 41 | TRPC1 | 10545 | -0.177 | -0.3307 | No | ||
| 42 | OSBP | 11014 | -0.272 | -0.3547 | No | ||
| 43 | NR4A1 | 11254 | -0.322 | -0.3660 | No | ||
| 44 | LHX1 | 11481 | -0.368 | -0.3764 | No | ||
| 45 | GDNF | 12015 | -0.487 | -0.4029 | No | ||
| 46 | KCNA6 | 12039 | -0.493 | -0.4018 | No | ||
| 47 | HOXB9 | 12053 | -0.495 | -0.4001 | No | ||
| 48 | SEMA3B | 12298 | -0.556 | -0.4106 | No | ||
| 49 | PTOV1 | 12519 | -0.608 | -0.4196 | No | ||
| 50 | SP1 | 12543 | -0.615 | -0.4178 | No | ||
| 51 | NOL4 | 12946 | -0.714 | -0.4361 | No | ||
| 52 | PHC1 | 12954 | -0.717 | -0.4331 | No | ||
| 53 | ELF1 | 13220 | -0.790 | -0.4436 | No | ||
| 54 | GNG4 | 13666 | -0.920 | -0.4632 | No | ||
| 55 | SLC30A5 | 13759 | -0.951 | -0.4636 | No | ||
| 56 | GFRA1 | 14062 | -1.052 | -0.4749 | No | ||
| 57 | TPT1 | 14296 | -1.133 | -0.4820 | No | ||
| 58 | SCYL1 | 14786 | -1.309 | -0.5021 | No | ||
| 59 | PITX2 | 14855 | -1.340 | -0.4994 | No | ||
| 60 | LHX5 | 14908 | -1.362 | -0.4957 | No | ||
| 61 | IRX3 | 15078 | -1.433 | -0.4979 | No | ||
| 62 | ETV6 | 15503 | -1.645 | -0.5129 | No | ||
| 63 | SLC39A5 | 15738 | -1.777 | -0.5171 | No | ||
| 64 | GRIA3 | 15891 | -1.875 | -0.5163 | No | ||
| 65 | LENG9 | 16241 | -2.130 | -0.5249 | No | ||
| 66 | SLC22A17 | 16439 | -2.304 | -0.5245 | No | ||
| 67 | TBX20 | 16739 | -2.604 | -0.5282 | Yes | ||
| 68 | ERF | 16970 | -2.888 | -0.5268 | Yes | ||
| 69 | RORC | 17063 | -3.008 | -0.5174 | Yes | ||
| 70 | SEC24D | 17140 | -3.135 | -0.5065 | Yes | ||
| 71 | SYT11 | 17717 | -4.305 | -0.5169 | Yes | ||
| 72 | SEC23A | 17843 | -4.645 | -0.5014 | Yes | ||
| 73 | STAT3 | 17907 | -4.850 | -0.4816 | Yes | ||
| 74 | CREM | 17917 | -4.880 | -0.4588 | Yes | ||
| 75 | PPP1R10 | 17964 | -5.003 | -0.4373 | Yes | ||
| 76 | KLF13 | 17992 | -5.096 | -0.4144 | Yes | ||
| 77 | TRIM39 | 18019 | -5.193 | -0.3909 | Yes | ||
| 78 | BRD2 | 18095 | -5.480 | -0.3687 | Yes | ||
| 79 | SH2D2A | 18280 | -6.523 | -0.3474 | Yes | ||
| 80 | EGR1 | 18462 | -8.469 | -0.3167 | Yes | ||
| 81 | MADD | 18522 | -9.863 | -0.2726 | Yes | ||
| 82 | RAB6IP1 | 18535 | -10.499 | -0.2230 | Yes | ||
| 83 | PAX1 | 18552 | -11.155 | -0.1705 | Yes | ||
| 84 | ARMCX2 | 18569 | -12.449 | -0.1118 | Yes | ||
| 85 | EGR2 | 18610 | -23.873 | 0.0003 | Yes |
