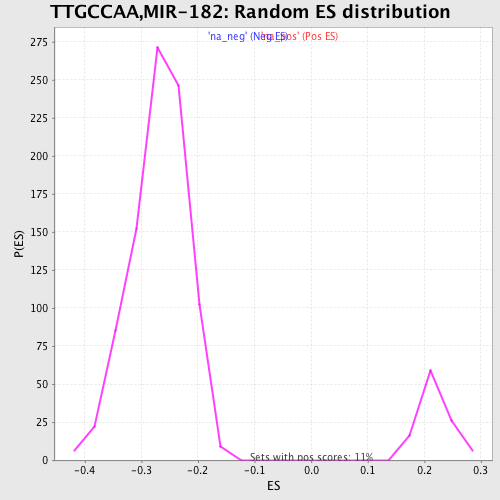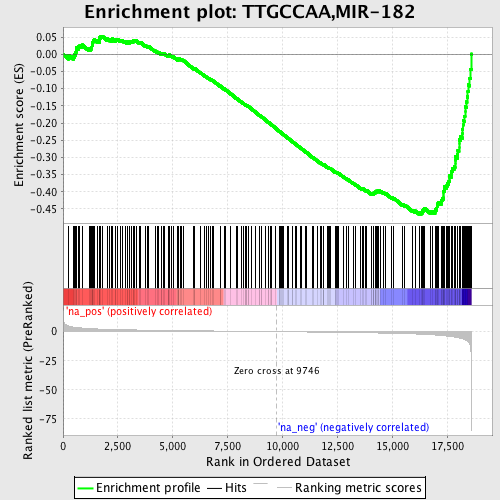
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | set03_absentNotch_versus_normalThy |
| Phenotype | NoPhenotypeAvailable |
| Upregulated in class | na_neg |
| GeneSet | TTGCCAA,MIR-182 |
| Enrichment Score (ES) | -0.46596944 |
| Normalized Enrichment Score (NES) | -1.7329378 |
| Nominal p-value | 0.0 |
| FDR q-value | 0.08710033 |
| FWER p-Value | 0.729 |

| PROBE | GENE SYMBOL | GENE_TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|---|
| 1 | PPIL1 | 259 | 4.787 | -0.0035 | No | ||
| 2 | CFL1 | 458 | 3.955 | -0.0055 | No | ||
| 3 | CEBPA | 514 | 3.808 | -0.0000 | No | ||
| 4 | MYOHD1 | 577 | 3.646 | 0.0047 | No | ||
| 5 | P2RX4 | 596 | 3.606 | 0.0117 | No | ||
| 6 | ACACA | 606 | 3.579 | 0.0191 | No | ||
| 7 | MTCH2 | 692 | 3.421 | 0.0221 | No | ||
| 8 | YAF2 | 764 | 3.302 | 0.0256 | No | ||
| 9 | NCALD | 860 | 3.139 | 0.0274 | No | ||
| 10 | RHOBTB1 | 1181 | 2.691 | 0.0159 | No | ||
| 11 | KPNA3 | 1257 | 2.592 | 0.0176 | No | ||
| 12 | DSCAM | 1305 | 2.544 | 0.0207 | No | ||
| 13 | ARHGEF3 | 1316 | 2.527 | 0.0257 | No | ||
| 14 | ZMPSTE24 | 1337 | 2.507 | 0.0302 | No | ||
| 15 | EVI5 | 1344 | 2.503 | 0.0354 | No | ||
| 16 | EIF3S10 | 1365 | 2.478 | 0.0398 | No | ||
| 17 | DENR | 1414 | 2.435 | 0.0426 | No | ||
| 18 | KTN1 | 1553 | 2.299 | 0.0402 | No | ||
| 19 | IBRDC2 | 1660 | 2.224 | 0.0394 | No | ||
| 20 | PPP3R1 | 1664 | 2.223 | 0.0441 | No | ||
| 21 | VLDLR | 1667 | 2.221 | 0.0490 | No | ||
| 22 | PYGO2 | 1718 | 2.184 | 0.0511 | No | ||
| 23 | ANK3 | 1775 | 2.140 | 0.0528 | No | ||
| 24 | BRE | 2000 | 2.009 | 0.0450 | No | ||
| 25 | AUP1 | 2122 | 1.940 | 0.0428 | No | ||
| 26 | MTSS1 | 2224 | 1.888 | 0.0415 | No | ||
| 27 | ATP2A2 | 2231 | 1.884 | 0.0453 | No | ||
| 28 | RDX | 2392 | 1.804 | 0.0406 | No | ||
| 29 | THBS2 | 2404 | 1.796 | 0.0440 | No | ||
| 30 | CLOCK | 2497 | 1.756 | 0.0429 | No | ||
| 31 | UBXD3 | 2601 | 1.714 | 0.0411 | No | ||
| 32 | FZD3 | 2685 | 1.674 | 0.0403 | No | ||
| 33 | STARD13 | 2822 | 1.617 | 0.0365 | No | ||
| 34 | KDELR1 | 2929 | 1.572 | 0.0342 | No | ||
| 35 | EXOC4 | 2950 | 1.563 | 0.0366 | No | ||
| 36 | EDNRB | 3029 | 1.533 | 0.0357 | No | ||
| 37 | TEX2 | 3040 | 1.529 | 0.0386 | No | ||
| 38 | PPP1R2 | 3103 | 1.510 | 0.0385 | No | ||
| 39 | ARMC1 | 3188 | 1.477 | 0.0372 | No | ||
| 40 | NTNG1 | 3189 | 1.477 | 0.0405 | No | ||
| 41 | JUB | 3233 | 1.464 | 0.0414 | No | ||
| 42 | EGR3 | 3322 | 1.429 | 0.0398 | No | ||
| 43 | MYADM | 3488 | 1.369 | 0.0339 | No | ||
| 44 | INSIG1 | 3515 | 1.361 | 0.0355 | No | ||
| 45 | ZCCHC14 | 3761 | 1.282 | 0.0250 | No | ||
| 46 | DAZAP2 | 3867 | 1.248 | 0.0221 | No | ||
| 47 | CORO1C | 3886 | 1.241 | 0.0238 | No | ||
| 48 | HMGCLL1 | 4194 | 1.154 | 0.0097 | No | ||
| 49 | AKAP7 | 4297 | 1.130 | 0.0066 | No | ||
| 50 | BHLHB5 | 4364 | 1.111 | 0.0055 | No | ||
| 51 | TXNL1 | 4478 | 1.082 | 0.0018 | No | ||
| 52 | BCL2L13 | 4493 | 1.079 | 0.0034 | No | ||
| 53 | GMFB | 4570 | 1.061 | 0.0016 | No | ||
| 54 | PCDH18 | 4599 | 1.056 | 0.0024 | No | ||
| 55 | FGF9 | 4787 | 1.008 | -0.0055 | No | ||
| 56 | PLD1 | 4809 | 1.003 | -0.0044 | No | ||
| 57 | CDV3 | 4810 | 1.003 | -0.0022 | No | ||
| 58 | ADAM10 | 4832 | 0.997 | -0.0011 | No | ||
| 59 | EBF3 | 4957 | 0.967 | -0.0057 | No | ||
| 60 | GPR85 | 5026 | 0.953 | -0.0073 | No | ||
| 61 | ACVR1 | 5221 | 0.905 | -0.0158 | No | ||
| 62 | CTTN | 5240 | 0.900 | -0.0148 | No | ||
| 63 | EIF5 | 5246 | 0.898 | -0.0131 | No | ||
| 64 | DCUN1D1 | 5259 | 0.894 | -0.0118 | No | ||
| 65 | ARRDC3 | 5353 | 0.872 | -0.0149 | No | ||
| 66 | HOXA9 | 5376 | 0.866 | -0.0142 | No | ||
| 67 | PRDM1 | 5478 | 0.845 | -0.0178 | No | ||
| 68 | RAC1 | 5946 | 0.749 | -0.0415 | No | ||
| 69 | SEPT7 | 5966 | 0.745 | -0.0409 | No | ||
| 70 | PDZD4 | 5996 | 0.738 | -0.0408 | No | ||
| 71 | RGS17 | 6241 | 0.690 | -0.0526 | No | ||
| 72 | NFASC | 6447 | 0.644 | -0.0623 | No | ||
| 73 | SLC4A7 | 6545 | 0.625 | -0.0662 | No | ||
| 74 | SUHW2 | 6633 | 0.608 | -0.0696 | No | ||
| 75 | UBE2R2 | 6733 | 0.586 | -0.0737 | No | ||
| 76 | SP3 | 6806 | 0.568 | -0.0763 | No | ||
| 77 | NKX2-2 | 6855 | 0.561 | -0.0777 | No | ||
| 78 | BNC2 | 7154 | 0.501 | -0.0928 | No | ||
| 79 | SNX4 | 7162 | 0.499 | -0.0920 | No | ||
| 80 | QKI | 7336 | 0.465 | -0.1004 | No | ||
| 81 | RIMS3 | 7420 | 0.448 | -0.1039 | No | ||
| 82 | PCDH8 | 7642 | 0.406 | -0.1151 | No | ||
| 83 | APLN | 7885 | 0.357 | -0.1274 | No | ||
| 84 | KLF15 | 7969 | 0.341 | -0.1312 | No | ||
| 85 | ANUBL1 | 8123 | 0.312 | -0.1388 | No | ||
| 86 | LPHN2 | 8231 | 0.295 | -0.1440 | No | ||
| 87 | RALGDS | 8318 | 0.280 | -0.1480 | No | ||
| 88 | ATP1B3 | 8328 | 0.277 | -0.1479 | No | ||
| 89 | MMD | 8338 | 0.275 | -0.1478 | No | ||
| 90 | MEF2C | 8378 | 0.266 | -0.1493 | No | ||
| 91 | VAMP3 | 8427 | 0.259 | -0.1513 | No | ||
| 92 | BRUNOL6 | 8429 | 0.258 | -0.1508 | No | ||
| 93 | BRMS1L | 8570 | 0.228 | -0.1579 | No | ||
| 94 | SLC1A2 | 8754 | 0.195 | -0.1674 | No | ||
| 95 | NPM1 | 8937 | 0.166 | -0.1770 | No | ||
| 96 | NPTX1 | 8966 | 0.160 | -0.1781 | No | ||
| 97 | FOXF2 | 9038 | 0.147 | -0.1817 | No | ||
| 98 | ABCC3 | 9043 | 0.146 | -0.1816 | No | ||
| 99 | ADRA2C | 9208 | 0.115 | -0.1902 | No | ||
| 100 | ANK2 | 9341 | 0.088 | -0.1972 | No | ||
| 101 | MMP16 | 9346 | 0.087 | -0.1972 | No | ||
| 102 | ADCY2 | 9451 | 0.068 | -0.2027 | No | ||
| 103 | SEPT9 | 9479 | 0.060 | -0.2041 | No | ||
| 104 | MBNL2 | 9499 | 0.056 | -0.2050 | No | ||
| 105 | EPAS1 | 9692 | 0.010 | -0.2154 | No | ||
| 106 | ARF4 | 9848 | -0.021 | -0.2238 | No | ||
| 107 | TOPBP1 | 9864 | -0.025 | -0.2245 | No | ||
| 108 | CUL5 | 9919 | -0.039 | -0.2274 | No | ||
| 109 | RARG | 9975 | -0.054 | -0.2303 | No | ||
| 110 | BHLHB3 | 9995 | -0.058 | -0.2312 | No | ||
| 111 | SLC39A9 | 10044 | -0.068 | -0.2336 | No | ||
| 112 | FMR1 | 10231 | -0.107 | -0.2435 | No | ||
| 113 | PTHLH | 10250 | -0.112 | -0.2442 | No | ||
| 114 | TMEM47 | 10262 | -0.116 | -0.2446 | No | ||
| 115 | EIF2C1 | 10443 | -0.156 | -0.2540 | No | ||
| 116 | EFNB2 | 10446 | -0.157 | -0.2538 | No | ||
| 117 | CD2AP | 10571 | -0.181 | -0.2601 | No | ||
| 118 | GRM5 | 10617 | -0.190 | -0.2621 | No | ||
| 119 | ATOH8 | 10810 | -0.229 | -0.2721 | No | ||
| 120 | PRKACB | 10855 | -0.239 | -0.2739 | No | ||
| 121 | SATB2 | 11028 | -0.275 | -0.2827 | No | ||
| 122 | GRINL1A | 11090 | -0.288 | -0.2854 | No | ||
| 123 | CHES1 | 11376 | -0.345 | -0.3001 | No | ||
| 124 | BCL2L12 | 11422 | -0.356 | -0.3017 | No | ||
| 125 | KCMF1 | 11602 | -0.395 | -0.3106 | No | ||
| 126 | SCRT1 | 11710 | -0.420 | -0.3155 | No | ||
| 127 | TSNAX | 11769 | -0.431 | -0.3177 | No | ||
| 128 | WSB1 | 11858 | -0.453 | -0.3215 | No | ||
| 129 | DOK4 | 11868 | -0.455 | -0.3209 | No | ||
| 130 | EPHB1 | 11889 | -0.462 | -0.3210 | No | ||
| 131 | RET | 12052 | -0.495 | -0.3287 | No | ||
| 132 | PHF21A | 12098 | -0.505 | -0.3300 | No | ||
| 133 | SH3BP4 | 12145 | -0.516 | -0.3314 | No | ||
| 134 | TAF15 | 12178 | -0.524 | -0.3320 | No | ||
| 135 | PAIP2 | 12408 | -0.581 | -0.3431 | No | ||
| 136 | SOX2 | 12418 | -0.584 | -0.3423 | No | ||
| 137 | LIMK1 | 12451 | -0.593 | -0.3428 | No | ||
| 138 | MARK3 | 12524 | -0.609 | -0.3453 | No | ||
| 139 | FBXW11 | 12567 | -0.619 | -0.3462 | No | ||
| 140 | CCNJ | 12789 | -0.674 | -0.3568 | No | ||
| 141 | BDNF | 12902 | -0.703 | -0.3613 | No | ||
| 142 | PRPF4B | 12998 | -0.729 | -0.3648 | No | ||
| 143 | HOXA10 | 13212 | -0.788 | -0.3747 | No | ||
| 144 | NUMB | 13340 | -0.821 | -0.3798 | No | ||
| 145 | YWHAG | 13555 | -0.887 | -0.3894 | No | ||
| 146 | MITF | 13644 | -0.915 | -0.3922 | No | ||
| 147 | ARHGEF7 | 13668 | -0.920 | -0.3914 | No | ||
| 148 | GABBR1 | 13788 | -0.959 | -0.3957 | No | ||
| 149 | MFAP3 | 13827 | -0.974 | -0.3957 | No | ||
| 150 | LHX3 | 14042 | -1.045 | -0.4050 | No | ||
| 151 | EPHA7 | 14068 | -1.053 | -0.4040 | No | ||
| 152 | ISL1 | 14150 | -1.079 | -0.4060 | No | ||
| 153 | MAK | 14156 | -1.082 | -0.4039 | No | ||
| 154 | RAB10 | 14168 | -1.089 | -0.4021 | No | ||
| 155 | PRKD1 | 14218 | -1.106 | -0.4023 | No | ||
| 156 | MEF2D | 14230 | -1.112 | -0.4004 | No | ||
| 157 | PRRX1 | 14255 | -1.121 | -0.3992 | No | ||
| 158 | ADAMTS18 | 14272 | -1.126 | -0.3976 | No | ||
| 159 | ELL2 | 14333 | -1.145 | -0.3983 | No | ||
| 160 | MARCKS | 14370 | -1.156 | -0.3977 | No | ||
| 161 | BAG4 | 14377 | -1.160 | -0.3955 | No | ||
| 162 | ELL | 14486 | -1.202 | -0.3987 | No | ||
| 163 | ABLIM1 | 14606 | -1.240 | -0.4024 | No | ||
| 164 | RASA1 | 14704 | -1.279 | -0.4048 | No | ||
| 165 | SLITRK4 | 14971 | -1.387 | -0.4162 | No | ||
| 166 | SYPL1 | 15075 | -1.431 | -0.4187 | No | ||
| 167 | L1CAM | 15457 | -1.624 | -0.4358 | No | ||
| 168 | CITED2 | 15581 | -1.686 | -0.4387 | No | ||
| 169 | DR1 | 15941 | -1.903 | -0.4540 | No | ||
| 170 | TOB1 | 16052 | -1.970 | -0.4556 | No | ||
| 171 | EPM2A | 16243 | -2.131 | -0.4612 | Yes | ||
| 172 | FBXW7 | 16314 | -2.196 | -0.4602 | Yes | ||
| 173 | XPR1 | 16362 | -2.239 | -0.4578 | Yes | ||
| 174 | TRIM8 | 16391 | -2.264 | -0.4543 | Yes | ||
| 175 | ARHGDIA | 16428 | -2.296 | -0.4511 | Yes | ||
| 176 | VASP | 16467 | -2.333 | -0.4480 | Yes | ||
| 177 | BTBD11 | 16752 | -2.620 | -0.4577 | Yes | ||
| 178 | KHDRBS3 | 16830 | -2.715 | -0.4558 | Yes | ||
| 179 | RTN4 | 16951 | -2.869 | -0.4560 | Yes | ||
| 180 | PIGA | 16973 | -2.892 | -0.4507 | Yes | ||
| 181 | WASF2 | 17028 | -2.961 | -0.4471 | Yes | ||
| 182 | NAP1L1 | 17040 | -2.987 | -0.4411 | Yes | ||
| 183 | IGSF3 | 17054 | -2.996 | -0.4352 | Yes | ||
| 184 | ADD3 | 17108 | -3.081 | -0.4312 | Yes | ||
| 185 | RAB6B | 17230 | -3.281 | -0.4305 | Yes | ||
| 186 | CD47 | 17247 | -3.304 | -0.4241 | Yes | ||
| 187 | SPRY4 | 17278 | -3.353 | -0.4183 | Yes | ||
| 188 | VAV2 | 17336 | -3.460 | -0.4137 | Yes | ||
| 189 | SNAP23 | 17338 | -3.461 | -0.4061 | Yes | ||
| 190 | PHF13 | 17340 | -3.474 | -0.3984 | Yes | ||
| 191 | OGT | 17379 | -3.550 | -0.3926 | Yes | ||
| 192 | ELMO1 | 17391 | -3.567 | -0.3853 | Yes | ||
| 193 | ADCY6 | 17491 | -3.802 | -0.3823 | Yes | ||
| 194 | DYNC1LI2 | 17513 | -3.845 | -0.3749 | Yes | ||
| 195 | DOCK9 | 17579 | -3.973 | -0.3696 | Yes | ||
| 196 | MAGI1 | 17608 | -4.021 | -0.3622 | Yes | ||
| 197 | ATXN1 | 17611 | -4.029 | -0.3534 | Yes | ||
| 198 | ETS2 | 17687 | -4.238 | -0.3481 | Yes | ||
| 199 | FEM1C | 17714 | -4.300 | -0.3400 | Yes | ||
| 200 | UBE3C | 17742 | -4.355 | -0.3318 | Yes | ||
| 201 | AEBP2 | 17820 | -4.582 | -0.3258 | Yes | ||
| 202 | KLF7 | 17865 | -4.734 | -0.3177 | Yes | ||
| 203 | PAFAH1B1 | 17867 | -4.737 | -0.3073 | Yes | ||
| 204 | PCMT1 | 17876 | -4.760 | -0.2972 | Yes | ||
| 205 | MAP1LC3B | 17969 | -5.015 | -0.2911 | Yes | ||
| 206 | KLF13 | 17992 | -5.096 | -0.2810 | Yes | ||
| 207 | ZNRF1 | 18044 | -5.291 | -0.2720 | Yes | ||
| 208 | XBP1 | 18046 | -5.294 | -0.2603 | Yes | ||
| 209 | CSNK1E | 18048 | -5.309 | -0.2486 | Yes | ||
| 210 | ZFP36L1 | 18112 | -5.581 | -0.2397 | Yes | ||
| 211 | RNF44 | 18193 | -5.987 | -0.2307 | Yes | ||
| 212 | SMAD1 | 18200 | -6.017 | -0.2177 | Yes | ||
| 213 | SH2D1A | 18225 | -6.148 | -0.2054 | Yes | ||
| 214 | SS18L1 | 18232 | -6.212 | -0.1920 | Yes | ||
| 215 | DYRK1A | 18283 | -6.537 | -0.1802 | Yes | ||
| 216 | IHPK1 | 18319 | -6.870 | -0.1669 | Yes | ||
| 217 | FLOT1 | 18322 | -6.883 | -0.1517 | Yes | ||
| 218 | BCL11A | 18381 | -7.407 | -0.1385 | Yes | ||
| 219 | GIT2 | 18409 | -7.660 | -0.1230 | Yes | ||
| 220 | WDR40A | 18449 | -8.226 | -0.1068 | Yes | ||
| 221 | PBX2 | 18469 | -8.589 | -0.0888 | Yes | ||
| 222 | BCL2 | 18536 | -10.511 | -0.0691 | Yes | ||
| 223 | ZFP36 | 18563 | -12.225 | -0.0434 | Yes | ||
| 224 | BACH2 | 18601 | -20.879 | 0.0008 | Yes |