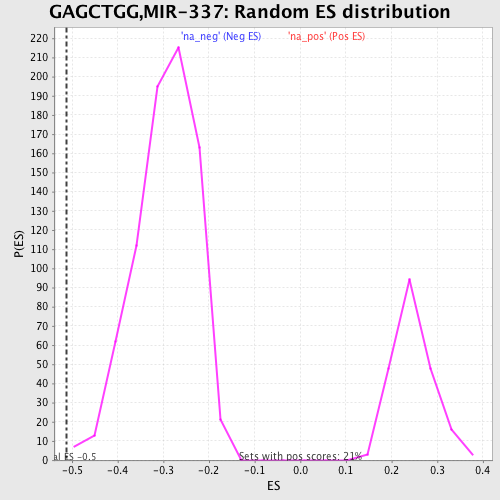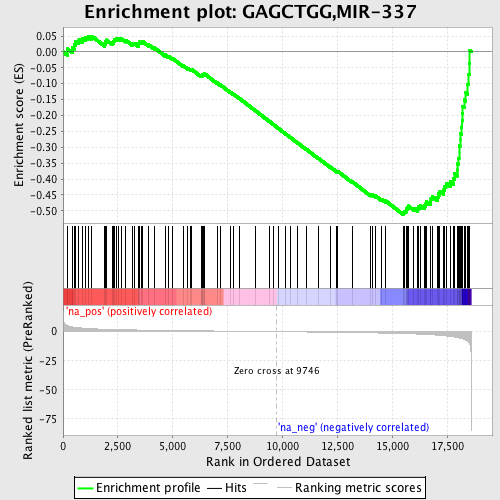
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | set03_absentNotch_versus_normalThy |
| Phenotype | NoPhenotypeAvailable |
| Upregulated in class | na_neg |
| GeneSet | GAGCTGG,MIR-337 |
| Enrichment Score (ES) | -0.5114837 |
| Normalized Enrichment Score (NES) | -1.7337842 |
| Nominal p-value | 0.002538071 |
| FDR q-value | 0.09177021 |
| FWER p-Value | 0.726 |

| PROBE | GENE SYMBOL | GENE_TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|---|
| 1 | ABCF1 | 186 | 5.310 | 0.0101 | No | ||
| 2 | NR1D1 | 421 | 4.076 | 0.0129 | No | ||
| 3 | BCAP29 | 503 | 3.841 | 0.0231 | No | ||
| 4 | THRA | 573 | 3.659 | 0.0333 | No | ||
| 5 | H1F0 | 723 | 3.371 | 0.0380 | No | ||
| 6 | EI24 | 876 | 3.109 | 0.0416 | No | ||
| 7 | MFN1 | 1004 | 2.924 | 0.0458 | No | ||
| 8 | PACSIN2 | 1138 | 2.749 | 0.0491 | No | ||
| 9 | DSCAM | 1305 | 2.544 | 0.0498 | No | ||
| 10 | PDGFRB | 1899 | 2.068 | 0.0256 | No | ||
| 11 | RAB15 | 1924 | 2.050 | 0.0321 | No | ||
| 12 | BNIPL | 1964 | 2.030 | 0.0377 | No | ||
| 13 | CD44 | 2249 | 1.872 | 0.0294 | No | ||
| 14 | EFNB1 | 2301 | 1.846 | 0.0337 | No | ||
| 15 | IER5L | 2343 | 1.830 | 0.0384 | No | ||
| 16 | MCRS1 | 2412 | 1.793 | 0.0415 | No | ||
| 17 | OVOL1 | 2515 | 1.747 | 0.0427 | No | ||
| 18 | NAV1 | 2647 | 1.692 | 0.0420 | No | ||
| 19 | GPD1 | 2848 | 1.606 | 0.0373 | No | ||
| 20 | ACOT7 | 3175 | 1.483 | 0.0253 | No | ||
| 21 | CHD4 | 3237 | 1.460 | 0.0275 | No | ||
| 22 | COMMD3 | 3444 | 1.382 | 0.0217 | No | ||
| 23 | DPH2 | 3445 | 1.382 | 0.0269 | No | ||
| 24 | SLC12A5 | 3459 | 1.379 | 0.0314 | No | ||
| 25 | RNF121 | 3565 | 1.343 | 0.0309 | No | ||
| 26 | ABCA9 | 3623 | 1.320 | 0.0328 | No | ||
| 27 | MAMDC1 | 3899 | 1.238 | 0.0226 | No | ||
| 28 | MRPS16 | 4157 | 1.164 | 0.0132 | No | ||
| 29 | LRRC4 | 4648 | 1.045 | -0.0093 | No | ||
| 30 | RNF19 | 4793 | 1.006 | -0.0133 | No | ||
| 31 | TWIST1 | 4985 | 0.959 | -0.0200 | No | ||
| 32 | MAT2A | 5468 | 0.847 | -0.0428 | No | ||
| 33 | TOMM34 | 5690 | 0.802 | -0.0517 | No | ||
| 34 | EHMT2 | 5799 | 0.779 | -0.0546 | No | ||
| 35 | KLF11 | 5850 | 0.769 | -0.0544 | No | ||
| 36 | ITGA11 | 6294 | 0.678 | -0.0757 | No | ||
| 37 | KCND1 | 6311 | 0.674 | -0.0740 | No | ||
| 38 | MAP3K9 | 6348 | 0.665 | -0.0735 | No | ||
| 39 | CHRD | 6356 | 0.664 | -0.0713 | No | ||
| 40 | RASGRP4 | 6402 | 0.653 | -0.0713 | No | ||
| 41 | NFASC | 6447 | 0.644 | -0.0712 | No | ||
| 42 | CAMK2G | 6454 | 0.643 | -0.0691 | No | ||
| 43 | AP1S3 | 7016 | 0.527 | -0.0974 | No | ||
| 44 | BNC2 | 7154 | 0.501 | -0.1029 | No | ||
| 45 | JPH1 | 7623 | 0.409 | -0.1266 | No | ||
| 46 | YPEL1 | 7769 | 0.384 | -0.1330 | No | ||
| 47 | SFRS1 | 8028 | 0.331 | -0.1457 | No | ||
| 48 | SLC1A2 | 8754 | 0.195 | -0.1841 | No | ||
| 49 | TYSND1 | 9414 | 0.074 | -0.2194 | No | ||
| 50 | VCP | 9566 | 0.041 | -0.2274 | No | ||
| 51 | B3GAT3 | 9807 | -0.014 | -0.2403 | No | ||
| 52 | STIP1 | 10133 | -0.087 | -0.2576 | No | ||
| 53 | KCNH8 | 10377 | -0.142 | -0.2702 | No | ||
| 54 | GRK6 | 10681 | -0.202 | -0.2858 | No | ||
| 55 | FOS | 11102 | -0.290 | -0.3074 | No | ||
| 56 | TOR2A | 11639 | -0.404 | -0.3348 | No | ||
| 57 | SP7 | 12176 | -0.524 | -0.3618 | No | ||
| 58 | HOXB5 | 12480 | -0.599 | -0.3758 | No | ||
| 59 | SEMA6A | 12520 | -0.609 | -0.3756 | No | ||
| 60 | FOSL1 | 13166 | -0.779 | -0.4075 | No | ||
| 61 | TRIM35 | 14015 | -1.037 | -0.4494 | No | ||
| 62 | B4GALT3 | 14083 | -1.058 | -0.4490 | No | ||
| 63 | SSBP3 | 14229 | -1.112 | -0.4526 | No | ||
| 64 | LBX1 | 14514 | -1.210 | -0.4634 | No | ||
| 65 | TEAD3 | 14698 | -1.277 | -0.4684 | No | ||
| 66 | PARVA | 15496 | -1.642 | -0.5053 | Yes | ||
| 67 | ALDH1A2 | 15539 | -1.660 | -0.5012 | Yes | ||
| 68 | SEC24C | 15631 | -1.713 | -0.4996 | Yes | ||
| 69 | SMARCD2 | 15638 | -1.717 | -0.4934 | Yes | ||
| 70 | SBK1 | 15689 | -1.743 | -0.4895 | Yes | ||
| 71 | SLC25A22 | 15723 | -1.770 | -0.4846 | Yes | ||
| 72 | PPP1R3F | 15980 | -1.924 | -0.4911 | Yes | ||
| 73 | SEPT6 | 16162 | -2.069 | -0.4931 | Yes | ||
| 74 | FRAG1 | 16197 | -2.095 | -0.4869 | Yes | ||
| 75 | BTG2 | 16278 | -2.162 | -0.4831 | Yes | ||
| 76 | ZMYND11 | 16465 | -2.332 | -0.4843 | Yes | ||
| 77 | ROM1 | 16523 | -2.385 | -0.4783 | Yes | ||
| 78 | SORT1 | 16557 | -2.417 | -0.4709 | Yes | ||
| 79 | FBXL17 | 16743 | -2.610 | -0.4710 | Yes | ||
| 80 | KLF12 | 16764 | -2.638 | -0.4621 | Yes | ||
| 81 | ZFAND3 | 16817 | -2.705 | -0.4546 | Yes | ||
| 82 | LIMK2 | 17076 | -3.030 | -0.4570 | Yes | ||
| 83 | DNAJB5 | 17098 | -3.061 | -0.4466 | Yes | ||
| 84 | POGZ | 17173 | -3.185 | -0.4385 | Yes | ||
| 85 | PHF13 | 17340 | -3.474 | -0.4343 | Yes | ||
| 86 | DGKZ | 17399 | -3.584 | -0.4238 | Yes | ||
| 87 | BCL2L1 | 17452 | -3.694 | -0.4126 | Yes | ||
| 88 | TXNL4B | 17652 | -4.135 | -0.4076 | Yes | ||
| 89 | RANBP10 | 17810 | -4.553 | -0.3988 | Yes | ||
| 90 | JUNB | 17839 | -4.638 | -0.3828 | Yes | ||
| 91 | PLAGL2 | 17986 | -5.060 | -0.3714 | Yes | ||
| 92 | KLF13 | 17992 | -5.096 | -0.3524 | Yes | ||
| 93 | MAP3K3 | 18023 | -5.201 | -0.3343 | Yes | ||
| 94 | GPRASP2 | 18045 | -5.292 | -0.3153 | Yes | ||
| 95 | XBP1 | 18046 | -5.294 | -0.2952 | Yes | ||
| 96 | LPHN1 | 18092 | -5.472 | -0.2769 | Yes | ||
| 97 | BRD2 | 18095 | -5.480 | -0.2562 | Yes | ||
| 98 | UHRF2 | 18159 | -5.804 | -0.2376 | Yes | ||
| 99 | GLCCI1 | 18180 | -5.895 | -0.2163 | Yes | ||
| 100 | RNF44 | 18193 | -5.987 | -0.1942 | Yes | ||
| 101 | UBAP1 | 18210 | -6.076 | -0.1720 | Yes | ||
| 102 | PARP6 | 18278 | -6.517 | -0.1509 | Yes | ||
| 103 | PPP1R9B | 18353 | -7.203 | -0.1276 | Yes | ||
| 104 | RARA | 18447 | -8.209 | -0.1015 | Yes | ||
| 105 | RGS16 | 18492 | -8.976 | -0.0698 | Yes | ||
| 106 | VAMP1 | 18512 | -9.484 | -0.0348 | Yes | ||
| 107 | PHF2 | 18539 | -10.641 | 0.0042 | Yes |