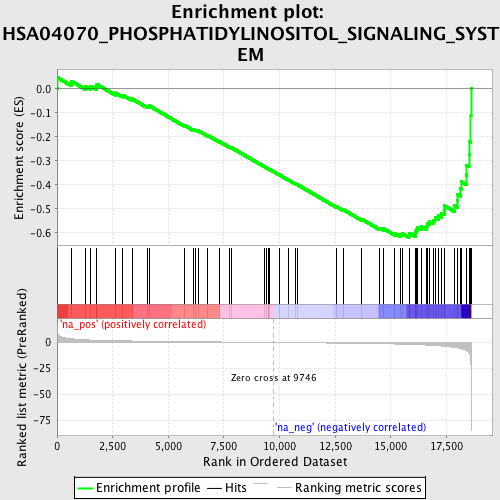
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | set03_absentNotch_versus_normalThy |
| Phenotype | NoPhenotypeAvailable |
| Upregulated in class | na_neg |
| GeneSet | HSA04070_PHOSPHATIDYLINOSITOL_SIGNALING_SYSTEM |
| Enrichment Score (ES) | -0.61927086 |
| Normalized Enrichment Score (NES) | -1.9596571 |
| Nominal p-value | 0.0013297872 |
| FDR q-value | 0.16337404 |
| FWER p-Value | 0.651 |

| PROBE | GENE SYMBOL | GENE_TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|---|
| 1 | INPPL1 | 20 | 8.974 | 0.0464 | No | ||
| 2 | PIK3R3 | 654 | 3.483 | 0.0307 | No | ||
| 3 | ITPR2 | 1283 | 2.557 | 0.0104 | No | ||
| 4 | PIK3R2 | 1505 | 2.342 | 0.0109 | No | ||
| 5 | IMPA2 | 1756 | 2.153 | 0.0088 | No | ||
| 6 | PLCB3 | 1789 | 2.129 | 0.0183 | No | ||
| 7 | PLCG1 | 2622 | 1.699 | -0.0175 | No | ||
| 8 | ITPKA | 2938 | 1.568 | -0.0262 | No | ||
| 9 | PLCD3 | 3381 | 1.407 | -0.0426 | No | ||
| 10 | PIK3C2A | 4080 | 1.185 | -0.0739 | No | ||
| 11 | CALM1 | 4140 | 1.168 | -0.0709 | No | ||
| 12 | CALML3 | 5725 | 0.794 | -0.1521 | No | ||
| 13 | PLCG2 | 6130 | 0.712 | -0.1701 | No | ||
| 14 | PIK3R1 | 6233 | 0.693 | -0.1719 | No | ||
| 15 | ITPR3 | 6364 | 0.662 | -0.1754 | No | ||
| 16 | CDS1 | 6769 | 0.577 | -0.1942 | No | ||
| 17 | PLCB2 | 7296 | 0.473 | -0.2200 | No | ||
| 18 | PIK3C2G | 7766 | 0.384 | -0.2433 | No | ||
| 19 | PLCB1 | 7859 | 0.363 | -0.2463 | No | ||
| 20 | PIK3CB | 9318 | 0.093 | -0.3244 | No | ||
| 21 | PLCB4 | 9419 | 0.074 | -0.3294 | No | ||
| 22 | INPP5E | 9501 | 0.055 | -0.3335 | No | ||
| 23 | PLCD4 | 9521 | 0.052 | -0.3342 | No | ||
| 24 | CALM2 | 9554 | 0.043 | -0.3357 | No | ||
| 25 | PLCZ1 | 10015 | -0.062 | -0.3602 | No | ||
| 26 | PRKCA | 10410 | -0.150 | -0.3806 | No | ||
| 27 | DGKB | 10705 | -0.206 | -0.3954 | No | ||
| 28 | PIP5K3 | 10790 | -0.223 | -0.3987 | No | ||
| 29 | CARKL | 12577 | -0.622 | -0.4917 | No | ||
| 30 | ITGB1BP3 | 12870 | -0.697 | -0.5037 | No | ||
| 31 | PRKCB1 | 13692 | -0.927 | -0.5431 | No | ||
| 32 | PIK3CA | 14498 | -1.205 | -0.5801 | No | ||
| 33 | ITPR1 | 14675 | -1.265 | -0.5829 | No | ||
| 34 | ITPK1 | 15175 | -1.475 | -0.6020 | No | ||
| 35 | PIK3C3 | 15426 | -1.606 | -0.6070 | No | ||
| 36 | PLCE1 | 15502 | -1.645 | -0.6023 | No | ||
| 37 | PLCD1 | 15818 | -1.827 | -0.6096 | Yes | ||
| 38 | PTEN | 15842 | -1.843 | -0.6011 | Yes | ||
| 39 | INPP5A | 16104 | -2.004 | -0.6046 | Yes | ||
| 40 | DGKE | 16108 | -2.013 | -0.5941 | Yes | ||
| 41 | PIK3CG | 16169 | -2.073 | -0.5863 | Yes | ||
| 42 | INPP5B | 16215 | -2.106 | -0.5776 | Yes | ||
| 43 | PIK3CD | 16364 | -2.240 | -0.5737 | Yes | ||
| 44 | SYNJ2 | 16616 | -2.474 | -0.5742 | Yes | ||
| 45 | CDS2 | 16630 | -2.485 | -0.5617 | Yes | ||
| 46 | CALM3 | 16720 | -2.583 | -0.5529 | Yes | ||
| 47 | INPP4A | 16907 | -2.818 | -0.5480 | Yes | ||
| 48 | IMPA1 | 16985 | -2.909 | -0.5367 | Yes | ||
| 49 | CDIPT | 17137 | -3.128 | -0.5283 | Yes | ||
| 50 | FN3K | 17275 | -3.345 | -0.5180 | Yes | ||
| 51 | DGKZ | 17399 | -3.584 | -0.5057 | Yes | ||
| 52 | PIB5PA | 17428 | -3.634 | -0.4880 | Yes | ||
| 53 | PIP5K1C | 17856 | -4.690 | -0.4862 | Yes | ||
| 54 | DGKQ | 17996 | -5.108 | -0.4666 | Yes | ||
| 55 | INPP5D | 18013 | -5.161 | -0.4402 | Yes | ||
| 56 | PIK3R5 | 18141 | -5.704 | -0.4169 | Yes | ||
| 57 | ITPKB | 18188 | -5.966 | -0.3878 | Yes | ||
| 58 | PIP5K1B | 18388 | -7.474 | -0.3589 | Yes | ||
| 59 | INPP1 | 18413 | -7.686 | -0.3196 | Yes | ||
| 60 | DGKA | 18516 | -9.602 | -0.2743 | Yes | ||
| 61 | INPP4B | 18538 | -10.612 | -0.2193 | Yes | ||
| 62 | PIP5K1A | 18602 | -20.953 | -0.1118 | Yes | ||
| 63 | DGKG | 18605 | -21.261 | 0.0006 | Yes |
