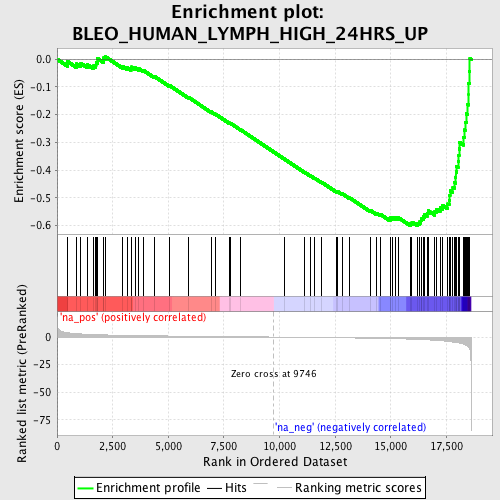
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | set03_absentNotch_versus_normalThy |
| Phenotype | NoPhenotypeAvailable |
| Upregulated in class | na_neg |
| GeneSet | BLEO_HUMAN_LYMPH_HIGH_24HRS_UP |
| Enrichment Score (ES) | -0.60099 |
| Normalized Enrichment Score (NES) | -1.992719 |
| Nominal p-value | 0.0 |
| FDR q-value | 0.23183468 |
| FWER p-Value | 0.528 |

| PROBE | GENE SYMBOL | GENE_TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|---|
| 1 | MYO5A | 473 | 3.908 | -0.0086 | No | ||
| 2 | EI24 | 876 | 3.109 | -0.0168 | No | ||
| 3 | CDKN1A | 1066 | 2.845 | -0.0147 | No | ||
| 4 | RPS6KA1 | 1354 | 2.492 | -0.0194 | No | ||
| 5 | GCH1 | 1625 | 2.251 | -0.0242 | No | ||
| 6 | FDXR | 1739 | 2.169 | -0.0209 | No | ||
| 7 | ZFP36L2 | 1748 | 2.164 | -0.0120 | No | ||
| 8 | ENC1 | 1818 | 2.112 | -0.0066 | No | ||
| 9 | CTSH | 1821 | 2.110 | 0.0025 | No | ||
| 10 | PLEK | 2083 | 1.964 | -0.0031 | No | ||
| 11 | CCL2 | 2098 | 1.953 | 0.0046 | No | ||
| 12 | TANK | 2180 | 1.913 | 0.0085 | No | ||
| 13 | ADAM8 | 2944 | 1.565 | -0.0259 | No | ||
| 14 | ACTA2 | 3144 | 1.495 | -0.0301 | No | ||
| 15 | PTP4A1 | 3323 | 1.429 | -0.0336 | No | ||
| 16 | CCNG1 | 3334 | 1.425 | -0.0279 | No | ||
| 17 | GDF15 | 3504 | 1.364 | -0.0311 | No | ||
| 18 | TNFRSF10B | 3671 | 1.307 | -0.0344 | No | ||
| 19 | ANXA4 | 3870 | 1.247 | -0.0397 | No | ||
| 20 | CFLAR | 4367 | 1.109 | -0.0617 | No | ||
| 21 | DUSP14 | 5073 | 0.942 | -0.0956 | No | ||
| 22 | NCF2 | 5899 | 0.759 | -0.1368 | No | ||
| 23 | GPX1 | 6918 | 0.547 | -0.1894 | No | ||
| 24 | RUNX3 | 7109 | 0.509 | -0.1974 | No | ||
| 25 | CAPN1 | 7738 | 0.387 | -0.2296 | No | ||
| 26 | FUCA1 | 7780 | 0.380 | -0.2302 | No | ||
| 27 | CRYZ | 8260 | 0.290 | -0.2548 | No | ||
| 28 | SH3BP5 | 10208 | -0.103 | -0.3594 | No | ||
| 29 | CCL4 | 11116 | -0.294 | -0.4070 | No | ||
| 30 | MS4A1 | 11396 | -0.350 | -0.4206 | No | ||
| 31 | XPC | 11588 | -0.393 | -0.4292 | No | ||
| 32 | HCLS1 | 11875 | -0.458 | -0.4426 | No | ||
| 33 | PRKAR1A | 12559 | -0.619 | -0.4768 | No | ||
| 34 | AIM1 | 12590 | -0.624 | -0.4757 | No | ||
| 35 | CCR1 | 12814 | -0.680 | -0.4848 | No | ||
| 36 | RAD51C | 13155 | -0.776 | -0.4998 | No | ||
| 37 | LRMP | 14085 | -1.059 | -0.5453 | No | ||
| 38 | MARCKS | 14370 | -1.156 | -0.5556 | No | ||
| 39 | HIF1A | 14529 | -1.213 | -0.5589 | No | ||
| 40 | LAPTM5 | 14969 | -1.387 | -0.5766 | No | ||
| 41 | PPM1D | 14995 | -1.396 | -0.5719 | No | ||
| 42 | SOX4 | 15077 | -1.432 | -0.5700 | No | ||
| 43 | SMTN | 15204 | -1.486 | -0.5704 | No | ||
| 44 | BIRC3 | 15345 | -1.560 | -0.5712 | No | ||
| 45 | ICAM1 | 15892 | -1.876 | -0.5925 | Yes | ||
| 46 | PMAIP1 | 15949 | -1.907 | -0.5873 | Yes | ||
| 47 | BASP1 | 16204 | -2.098 | -0.5919 | Yes | ||
| 48 | BTG2 | 16278 | -2.162 | -0.5865 | Yes | ||
| 49 | TNFSF9 | 16356 | -2.231 | -0.5810 | Yes | ||
| 50 | TUSC2 | 16393 | -2.265 | -0.5731 | Yes | ||
| 51 | VASP | 16467 | -2.333 | -0.5670 | Yes | ||
| 52 | CD40 | 16527 | -2.391 | -0.5598 | Yes | ||
| 53 | LPXN | 16631 | -2.486 | -0.5546 | Yes | ||
| 54 | NFKBIA | 16685 | -2.540 | -0.5464 | Yes | ||
| 55 | NFKB2 | 16979 | -2.902 | -0.5497 | Yes | ||
| 56 | DUSP2 | 17039 | -2.987 | -0.5399 | Yes | ||
| 57 | CD83 | 17215 | -3.254 | -0.5353 | Yes | ||
| 58 | PKIG | 17318 | -3.425 | -0.5260 | Yes | ||
| 59 | CCND2 | 17556 | -3.932 | -0.5217 | Yes | ||
| 60 | MRPL49 | 17621 | -4.049 | -0.5076 | Yes | ||
| 61 | NCOA3 | 17650 | -4.131 | -0.4913 | Yes | ||
| 62 | PRKAB1 | 17662 | -4.161 | -0.4738 | Yes | ||
| 63 | PIM2 | 17790 | -4.508 | -0.4612 | Yes | ||
| 64 | LTA | 17854 | -4.683 | -0.4443 | Yes | ||
| 65 | STAT3 | 17907 | -4.850 | -0.4261 | Yes | ||
| 66 | PLD3 | 17929 | -4.908 | -0.4060 | Yes | ||
| 67 | TNFRSF9 | 17957 | -4.990 | -0.3858 | Yes | ||
| 68 | TAX1BP3 | 18026 | -5.208 | -0.3670 | Yes | ||
| 69 | FAS | 18055 | -5.334 | -0.3454 | Yes | ||
| 70 | WDR1 | 18069 | -5.373 | -0.3228 | Yes | ||
| 71 | BRD2 | 18095 | -5.480 | -0.3004 | Yes | ||
| 72 | SORL1 | 18263 | -6.369 | -0.2819 | Yes | ||
| 73 | DDB2 | 18301 | -6.705 | -0.2548 | Yes | ||
| 74 | TNFAIP3 | 18342 | -7.081 | -0.2263 | Yes | ||
| 75 | CYFIP2 | 18398 | -7.597 | -0.1964 | Yes | ||
| 76 | TRAF1 | 18458 | -8.368 | -0.1634 | Yes | ||
| 77 | RGS16 | 18492 | -8.976 | -0.1263 | Yes | ||
| 78 | GADD45A | 18496 | -9.058 | -0.0872 | Yes | ||
| 79 | BTG1 | 18531 | -10.306 | -0.0445 | Yes | ||
| 80 | CCL5 | 18555 | -11.313 | 0.0033 | Yes |
