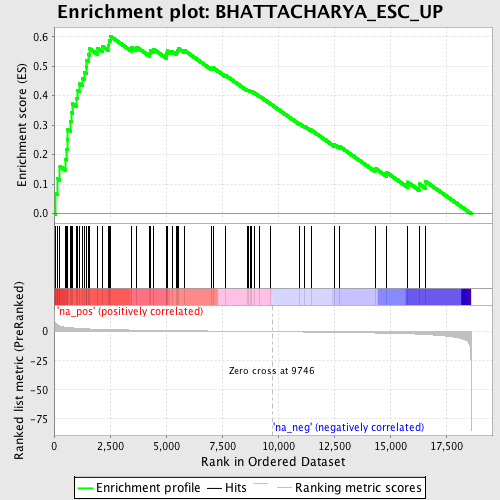
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | set03_absentNotch_versus_normalThy |
| Phenotype | NoPhenotypeAvailable |
| Upregulated in class | na_pos |
| GeneSet | BHATTACHARYA_ESC_UP |
| Enrichment Score (ES) | 0.60207117 |
| Normalized Enrichment Score (NES) | 2.2176452 |
| Nominal p-value | 0.0 |
| FDR q-value | 1.353349E-4 |
| FWER p-Value | 0.0020 |

| PROBE | GENE SYMBOL | GENE_TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|---|
| 1 | NME2 | 51 | 7.284 | 0.0693 | Yes | ||
| 2 | PPAT | 158 | 5.554 | 0.1185 | Yes | ||
| 3 | IMPDH2 | 260 | 4.782 | 0.1604 | Yes | ||
| 4 | HSPA4 | 512 | 3.809 | 0.1846 | Yes | ||
| 5 | DDX21 | 564 | 3.690 | 0.2183 | Yes | ||
| 6 | RPL4 | 590 | 3.612 | 0.2527 | Yes | ||
| 7 | SERPINH1 | 612 | 3.566 | 0.2868 | Yes | ||
| 8 | TNNT1 | 743 | 3.339 | 0.3129 | Yes | ||
| 9 | EIF4A1 | 769 | 3.290 | 0.3441 | Yes | ||
| 10 | SNRPF | 831 | 3.189 | 0.3723 | Yes | ||
| 11 | CCT8 | 1014 | 2.909 | 0.3913 | Yes | ||
| 12 | MTHFD1 | 1052 | 2.871 | 0.4177 | Yes | ||
| 13 | SMS | 1141 | 2.742 | 0.4401 | Yes | ||
| 14 | NASP | 1272 | 2.565 | 0.4584 | Yes | ||
| 15 | RPS24 | 1352 | 2.493 | 0.4788 | Yes | ||
| 16 | CALU | 1446 | 2.405 | 0.4976 | Yes | ||
| 17 | ELOVL6 | 1455 | 2.393 | 0.5209 | Yes | ||
| 18 | PSIP1 | 1536 | 2.314 | 0.5394 | Yes | ||
| 19 | RPL24 | 1579 | 2.279 | 0.5597 | Yes | ||
| 20 | PSMA3 | 1928 | 2.048 | 0.5612 | Yes | ||
| 21 | CCNC | 2168 | 1.918 | 0.5673 | Yes | ||
| 22 | KIF4A | 2427 | 1.786 | 0.5711 | Yes | ||
| 23 | GDF3 | 2450 | 1.775 | 0.5875 | Yes | ||
| 24 | EPRS | 2502 | 1.755 | 0.6021 | Yes | ||
| 25 | TK1 | 3446 | 1.381 | 0.5649 | No | ||
| 26 | MAD2L2 | 3688 | 1.302 | 0.5648 | No | ||
| 27 | TDGF1 | 4277 | 1.137 | 0.5444 | No | ||
| 28 | PSMA2 | 4304 | 1.128 | 0.5541 | No | ||
| 29 | HDAC2 | 4438 | 1.091 | 0.5578 | No | ||
| 30 | PODXL | 5003 | 0.957 | 0.5368 | No | ||
| 31 | LAPTM4B | 5029 | 0.952 | 0.5449 | No | ||
| 32 | GSH1 | 5071 | 0.943 | 0.5520 | No | ||
| 33 | SSB | 5264 | 0.893 | 0.5505 | No | ||
| 34 | MTHFD2 | 5477 | 0.845 | 0.5475 | No | ||
| 35 | SFRP2 | 5499 | 0.839 | 0.5546 | No | ||
| 36 | GJA1 | 5530 | 0.833 | 0.5613 | No | ||
| 37 | CRABP1 | 5832 | 0.774 | 0.5527 | No | ||
| 38 | SLC16A1 | 7026 | 0.525 | 0.4936 | No | ||
| 39 | RPL7 | 7103 | 0.510 | 0.4946 | No | ||
| 40 | FABP5 | 7656 | 0.404 | 0.4688 | No | ||
| 41 | GAL | 8632 | 0.216 | 0.4184 | No | ||
| 42 | KPNA2 | 8668 | 0.210 | 0.4186 | No | ||
| 43 | CCNB1 | 8775 | 0.192 | 0.4148 | No | ||
| 44 | HMGB2 | 8789 | 0.189 | 0.4160 | No | ||
| 45 | NPM1 | 8937 | 0.166 | 0.4097 | No | ||
| 46 | CRABP2 | 9173 | 0.121 | 0.3982 | No | ||
| 47 | RPL6 | 9665 | 0.017 | 0.3719 | No | ||
| 48 | SET | 10961 | -0.259 | 0.3047 | No | ||
| 49 | CYP26A1 | 11171 | -0.306 | 0.2965 | No | ||
| 50 | HNRPAB | 11474 | -0.367 | 0.2838 | No | ||
| 51 | SEMA6A | 12520 | -0.609 | 0.2336 | No | ||
| 52 | PTTG1 | 12731 | -0.661 | 0.2288 | No | ||
| 53 | LRRN1 | 14325 | -1.144 | 0.1543 | No | ||
| 54 | PITX2 | 14855 | -1.340 | 0.1390 | No | ||
| 55 | IDH1 | 15773 | -1.800 | 0.1074 | No | ||
| 56 | MGST1 | 16295 | -2.180 | 0.1009 | No | ||
| 57 | DSG2 | 16571 | -2.433 | 0.1102 | No |
