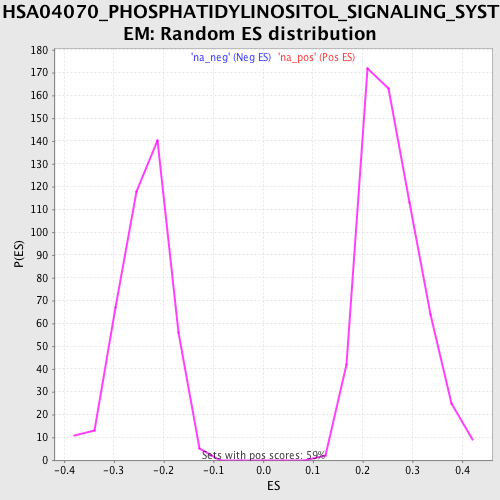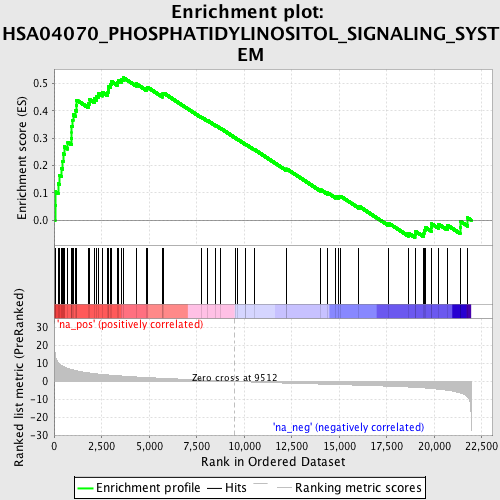
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | set02_BT_ATM_minus_versus_ATM_plus |
| Phenotype | NoPhenotypeAvailable |
| Upregulated in class | na_pos |
| GeneSet | HSA04070_PHOSPHATIDYLINOSITOL_SIGNALING_SYSTEM |
| Enrichment Score (ES) | 0.5212375 |
| Normalized Enrichment Score (NES) | 2.0205977 |
| Nominal p-value | 0.0 |
| FDR q-value | 0.010512049 |
| FWER p-Value | 0.139 |

| PROBE | GENE SYMBOL | GENE_TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|---|
| 1 | ITPKB | 48 | 15.738 | 0.0526 | Yes | ||
| 2 | DGKD | 60 | 14.906 | 0.1041 | Yes | ||
| 3 | PIK3CD | 211 | 10.436 | 0.1336 | Yes | ||
| 4 | INPP1 | 286 | 9.630 | 0.1637 | Yes | ||
| 5 | PIK3R5 | 401 | 8.752 | 0.1890 | Yes | ||
| 6 | PIP5K3 | 458 | 8.393 | 0.2157 | Yes | ||
| 7 | CDS2 | 495 | 8.171 | 0.2425 | Yes | ||
| 8 | PLCG1 | 539 | 7.963 | 0.2683 | Yes | ||
| 9 | INPP5D | 724 | 7.159 | 0.2848 | Yes | ||
| 10 | DGKG | 903 | 6.584 | 0.2996 | Yes | ||
| 11 | INPP5E | 924 | 6.529 | 0.3214 | Yes | ||
| 12 | ITPR2 | 927 | 6.521 | 0.3441 | Yes | ||
| 13 | PRKCB1 | 961 | 6.424 | 0.3649 | Yes | ||
| 14 | DGKQ | 996 | 6.351 | 0.3855 | Yes | ||
| 15 | SYNJ2 | 1128 | 6.030 | 0.4005 | Yes | ||
| 16 | DGKZ | 1177 | 5.913 | 0.4189 | Yes | ||
| 17 | PTEN | 1189 | 5.873 | 0.4389 | Yes | ||
| 18 | FN3K | 1810 | 4.738 | 0.4270 | Yes | ||
| 19 | OCRL | 1872 | 4.653 | 0.4405 | Yes | ||
| 20 | ITPR1 | 2124 | 4.312 | 0.4440 | Yes | ||
| 21 | INPP5A | 2244 | 4.176 | 0.4531 | Yes | ||
| 22 | PIP5K1A | 2320 | 4.092 | 0.4639 | Yes | ||
| 23 | PIP5K1B | 2542 | 3.858 | 0.4673 | Yes | ||
| 24 | DGKE | 2827 | 3.589 | 0.4668 | Yes | ||
| 25 | INPP4B | 2844 | 3.573 | 0.4785 | Yes | ||
| 26 | PIK3CA | 2870 | 3.541 | 0.4897 | Yes | ||
| 27 | INPP5B | 2973 | 3.457 | 0.4971 | Yes | ||
| 28 | CALM3 | 3006 | 3.431 | 0.5076 | Yes | ||
| 29 | PLCB2 | 3318 | 3.167 | 0.5044 | Yes | ||
| 30 | SYNJ1 | 3382 | 3.115 | 0.5124 | Yes | ||
| 31 | DGKA | 3551 | 2.979 | 0.5151 | Yes | ||
| 32 | IMPA1 | 3639 | 2.914 | 0.5212 | Yes | ||
| 33 | CDIPT | 4319 | 2.428 | 0.4987 | No | ||
| 34 | PIK3CG | 4859 | 2.117 | 0.4814 | No | ||
| 35 | PIK3C2G | 4903 | 2.096 | 0.4867 | No | ||
| 36 | PIK3C3 | 5706 | 1.681 | 0.4559 | No | ||
| 37 | PIK3CB | 5715 | 1.678 | 0.4614 | No | ||
| 38 | PIK3R1 | 5770 | 1.654 | 0.4647 | No | ||
| 39 | ITPR3 | 7763 | 0.754 | 0.3763 | No | ||
| 40 | ITPK1 | 8072 | 0.633 | 0.3644 | No | ||
| 41 | ITPKA | 8518 | 0.445 | 0.3456 | No | ||
| 42 | PLCD4 | 8737 | 0.346 | 0.3369 | No | ||
| 43 | PIP5K1C | 9535 | -0.007 | 0.3005 | No | ||
| 44 | ITGB1BP3 | 9662 | -0.071 | 0.2950 | No | ||
| 45 | PLCE1 | 10051 | -0.243 | 0.2781 | No | ||
| 46 | CARKL | 10065 | -0.249 | 0.2783 | No | ||
| 47 | CALML3 | 10546 | -0.420 | 0.2579 | No | ||
| 48 | PLCB1 | 12231 | -0.985 | 0.1843 | No | ||
| 49 | PLCD1 | 12249 | -0.991 | 0.1870 | No | ||
| 50 | INPP4A | 14043 | -1.520 | 0.1103 | No | ||
| 51 | PLCB3 | 14401 | -1.621 | 0.0997 | No | ||
| 52 | CALM2 | 14820 | -1.748 | 0.0867 | No | ||
| 53 | PLCD3 | 14970 | -1.790 | 0.0861 | No | ||
| 54 | PLCZ1 | 15068 | -1.820 | 0.0880 | No | ||
| 55 | PIK3C2A | 16040 | -2.134 | 0.0510 | No | ||
| 56 | PIB5PA | 17614 | -2.687 | -0.0115 | No | ||
| 57 | DGKB | 18644 | -3.148 | -0.0476 | No | ||
| 58 | CALM1 | 18993 | -3.349 | -0.0518 | No | ||
| 59 | PIK3R2 | 19008 | -3.358 | -0.0407 | No | ||
| 60 | CDS1 | 19459 | -3.655 | -0.0486 | No | ||
| 61 | DGKI | 19471 | -3.663 | -0.0363 | No | ||
| 62 | PIK3R3 | 19528 | -3.708 | -0.0260 | No | ||
| 63 | INPPL1 | 19844 | -3.988 | -0.0265 | No | ||
| 64 | PLCG2 | 19865 | -4.009 | -0.0134 | No | ||
| 65 | DGKH | 20223 | -4.384 | -0.0145 | No | ||
| 66 | IMPA2 | 20692 | -5.010 | -0.0184 | No | ||
| 67 | PLCB4 | 21366 | -6.490 | -0.0265 | No | ||
| 68 | PRKCA | 21377 | -6.512 | -0.0043 | No | ||
| 69 | PTPMT1 | 21764 | -8.705 | 0.0084 | No |