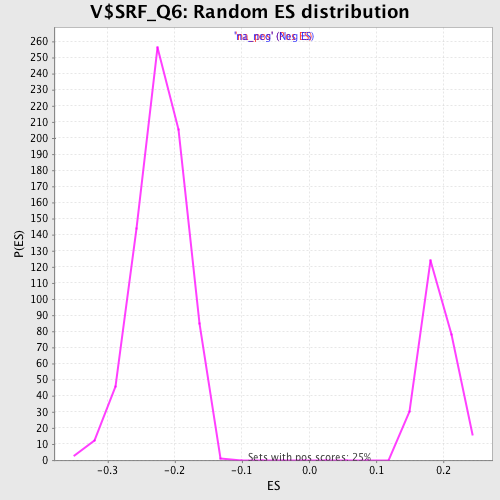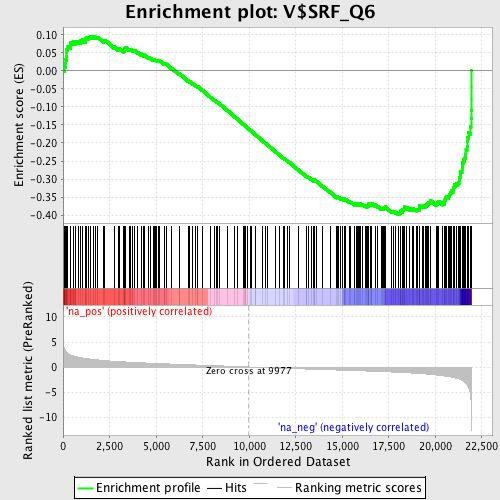
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | set02_ATM_minus_versus_BT_ATM_minus |
| Phenotype | NoPhenotypeAvailable |
| Upregulated in class | na_neg |
| GeneSet | V$SRF_Q6 |
| Enrichment Score (ES) | -0.398249 |
| Normalized Enrichment Score (NES) | -1.7974662 |
| Nominal p-value | 0.0 |
| FDR q-value | 0.016440844 |
| FWER p-Value | 0.133 |

| PROBE | GENE SYMBOL | GENE_TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|---|
| 1 | MATR3 | 55 | 3.775 | 0.0111 | No | ||
| 2 | CHD6 | 116 | 3.310 | 0.0204 | No | ||
| 3 | EML4 | 147 | 3.169 | 0.0305 | No | ||
| 4 | MAP2K6 | 175 | 2.990 | 0.0400 | No | ||
| 5 | ACTB | 198 | 2.933 | 0.0497 | No | ||
| 6 | HERC1 | 202 | 2.907 | 0.0600 | No | ||
| 7 | RHOJ | 259 | 2.699 | 0.0672 | No | ||
| 8 | H3F3A | 403 | 2.420 | 0.0694 | No | ||
| 9 | TCF8 | 411 | 2.409 | 0.0778 | No | ||
| 10 | LPP | 532 | 2.236 | 0.0804 | No | ||
| 11 | ASPH | 674 | 2.084 | 0.0815 | No | ||
| 12 | RASD2 | 823 | 1.939 | 0.0817 | No | ||
| 13 | ROCK2 | 935 | 1.861 | 0.0833 | No | ||
| 14 | ADAMTS12 | 1027 | 1.800 | 0.0857 | No | ||
| 15 | PICALM | 1210 | 1.703 | 0.0835 | No | ||
| 16 | MBNL1 | 1213 | 1.702 | 0.0895 | No | ||
| 17 | TAZ | 1276 | 1.668 | 0.0927 | No | ||
| 18 | FST | 1383 | 1.611 | 0.0937 | No | ||
| 19 | FOXP1 | 1454 | 1.579 | 0.0962 | No | ||
| 20 | ATP1A2 | 1622 | 1.510 | 0.0940 | No | ||
| 21 | CFL2 | 1742 | 1.467 | 0.0938 | No | ||
| 22 | PPAP2B | 1860 | 1.410 | 0.0935 | No | ||
| 23 | EMILIN2 | 2173 | 1.304 | 0.0839 | No | ||
| 24 | DIXDC1 | 2245 | 1.277 | 0.0853 | No | ||
| 25 | SCN3B | 2743 | 1.140 | 0.0665 | No | ||
| 26 | ELAVL4 | 2952 | 1.095 | 0.0609 | No | ||
| 27 | LDB3 | 3018 | 1.077 | 0.0619 | No | ||
| 28 | SYNCRIP | 3238 | 1.028 | 0.0555 | No | ||
| 29 | SCOC | 3287 | 1.015 | 0.0570 | No | ||
| 30 | RARB | 3291 | 1.014 | 0.0605 | No | ||
| 31 | ABL1 | 3313 | 1.010 | 0.0632 | No | ||
| 32 | PPP1R12A | 3345 | 1.003 | 0.0654 | No | ||
| 33 | IGF1 | 3560 | 0.957 | 0.0590 | No | ||
| 34 | PRM1 | 3629 | 0.943 | 0.0593 | No | ||
| 35 | SRD5A2 | 3744 | 0.919 | 0.0574 | No | ||
| 36 | SDPR | 3835 | 0.903 | 0.0566 | No | ||
| 37 | COL1A1 | 4020 | 0.869 | 0.0512 | No | ||
| 38 | ITGB1BP2 | 4185 | 0.841 | 0.0467 | No | ||
| 39 | NOL4 | 4345 | 0.814 | 0.0424 | No | ||
| 40 | PRKAB2 | 4374 | 0.807 | 0.0440 | No | ||
| 41 | PODN | 4576 | 0.774 | 0.0376 | No | ||
| 42 | MRGPRF | 4683 | 0.759 | 0.0354 | No | ||
| 43 | RAB27A | 4851 | 0.732 | 0.0304 | No | ||
| 44 | TNMD | 4889 | 0.727 | 0.0313 | No | ||
| 45 | UBE2D3 | 4979 | 0.711 | 0.0298 | No | ||
| 46 | RIMS1 | 5039 | 0.702 | 0.0297 | No | ||
| 47 | RRM1 | 5103 | 0.695 | 0.0293 | No | ||
| 48 | NPR3 | 5157 | 0.687 | 0.0293 | No | ||
| 49 | TGFB1I1 | 5422 | 0.645 | 0.0195 | No | ||
| 50 | DMPK | 5461 | 0.640 | 0.0201 | No | ||
| 51 | TRIM46 | 5535 | 0.630 | 0.0190 | No | ||
| 52 | HOXB4 | 5849 | 0.581 | 0.0067 | No | ||
| 53 | PDLIM4 | 6233 | 0.522 | -0.0090 | No | ||
| 54 | KCNK3 | 6250 | 0.519 | -0.0078 | No | ||
| 55 | NFATC4 | 6716 | 0.458 | -0.0276 | No | ||
| 56 | PPP2R3A | 6816 | 0.443 | -0.0305 | No | ||
| 57 | AGL | 6973 | 0.423 | -0.0361 | No | ||
| 58 | HOXB5 | 7102 | 0.406 | -0.0406 | No | ||
| 59 | LRRTM4 | 7211 | 0.393 | -0.0441 | No | ||
| 60 | SP7 | 7214 | 0.392 | -0.0428 | No | ||
| 61 | CKM | 7500 | 0.353 | -0.0546 | No | ||
| 62 | FOSB | 7935 | 0.296 | -0.0735 | No | ||
| 63 | DVL3 | 8129 | 0.269 | -0.0814 | No | ||
| 64 | MAP1A | 8260 | 0.253 | -0.0864 | No | ||
| 65 | CTDSP1 | 8301 | 0.247 | -0.0874 | No | ||
| 66 | PPARG | 8381 | 0.235 | -0.0902 | No | ||
| 67 | TNNC1 | 8829 | 0.172 | -0.1101 | No | ||
| 68 | RAP1A | 9206 | 0.119 | -0.1270 | No | ||
| 69 | NPPA | 9379 | 0.092 | -0.1345 | No | ||
| 70 | POU3F4 | 9717 | 0.041 | -0.1499 | No | ||
| 71 | CNN1 | 9765 | 0.034 | -0.1519 | No | ||
| 72 | JMJD1A | 9795 | 0.028 | -0.1531 | No | ||
| 73 | SIX2 | 9813 | 0.025 | -0.1538 | No | ||
| 74 | SEPX1 | 9896 | 0.013 | -0.1576 | No | ||
| 75 | STARD13 | 10083 | -0.016 | -0.1660 | No | ||
| 76 | MYL1 | 10144 | -0.026 | -0.1687 | No | ||
| 77 | GNG8 | 10315 | -0.052 | -0.1763 | No | ||
| 78 | MYL9 | 10325 | -0.053 | -0.1766 | No | ||
| 79 | MSX1 | 10717 | -0.110 | -0.1941 | No | ||
| 80 | TAGLN | 10864 | -0.134 | -0.2004 | No | ||
| 81 | MUS81 | 10991 | -0.154 | -0.2056 | No | ||
| 82 | EFHD1 | 11389 | -0.204 | -0.2231 | No | ||
| 83 | AQP2 | 11624 | -0.231 | -0.2330 | No | ||
| 84 | PRDM1 | 11841 | -0.256 | -0.2420 | No | ||
| 85 | ACTR3 | 11907 | -0.263 | -0.2441 | No | ||
| 86 | IGF2AS | 12059 | -0.278 | -0.2500 | No | ||
| 87 | BLOC1S1 | 12177 | -0.292 | -0.2543 | No | ||
| 88 | LRRFIP1 | 12643 | -0.349 | -0.2745 | No | ||
| 89 | PLN | 13078 | -0.400 | -0.2930 | No | ||
| 90 | H2AFX | 13090 | -0.401 | -0.2920 | No | ||
| 91 | RAI2 | 13177 | -0.410 | -0.2945 | No | ||
| 92 | PFN1 | 13354 | -0.432 | -0.3010 | No | ||
| 93 | EPHB1 | 13469 | -0.447 | -0.3046 | No | ||
| 94 | NKX2-2 | 13482 | -0.449 | -0.3036 | No | ||
| 95 | KCNA1 | 13486 | -0.449 | -0.3021 | No | ||
| 96 | MYL7 | 13606 | -0.461 | -0.3059 | No | ||
| 97 | MYO18B | 13939 | -0.503 | -0.3193 | No | ||
| 98 | DGKG | 14341 | -0.550 | -0.3358 | No | ||
| 99 | HOXA1 | 14710 | -0.597 | -0.3505 | No | ||
| 100 | FOXE3 | 14721 | -0.598 | -0.3488 | No | ||
| 101 | HIST1H4I | 14803 | -0.607 | -0.3503 | No | ||
| 102 | FGF8 | 14922 | -0.621 | -0.3535 | No | ||
| 103 | PHF12 | 15006 | -0.629 | -0.3551 | No | ||
| 104 | TADA3L | 15092 | -0.638 | -0.3567 | No | ||
| 105 | HOXA3 | 15093 | -0.638 | -0.3544 | No | ||
| 106 | CSMD3 | 15169 | -0.649 | -0.3554 | No | ||
| 107 | MYH11 | 15394 | -0.677 | -0.3633 | No | ||
| 108 | IL17B | 15461 | -0.684 | -0.3639 | No | ||
| 109 | MYLK | 15649 | -0.708 | -0.3699 | No | ||
| 110 | ACAA2 | 15666 | -0.711 | -0.3680 | No | ||
| 111 | MYO1E | 15779 | -0.725 | -0.3706 | No | ||
| 112 | HOXD13 | 15811 | -0.729 | -0.3694 | No | ||
| 113 | KRTCAP2 | 15846 | -0.734 | -0.3683 | No | ||
| 114 | GPR158 | 15941 | -0.747 | -0.3699 | No | ||
| 115 | ACTN1 | 15953 | -0.750 | -0.3677 | No | ||
| 116 | CACNA1B | 16098 | -0.771 | -0.3715 | No | ||
| 117 | NKX6-1 | 16226 | -0.787 | -0.3745 | No | ||
| 118 | FOSL1 | 16325 | -0.800 | -0.3761 | No | ||
| 119 | PLCB3 | 16358 | -0.805 | -0.3747 | No | ||
| 120 | MAPKAPK2 | 16390 | -0.809 | -0.3731 | No | ||
| 121 | FGFRL1 | 16396 | -0.810 | -0.3704 | No | ||
| 122 | PCDH7 | 16430 | -0.813 | -0.3690 | No | ||
| 123 | RBBP7 | 16538 | -0.828 | -0.3709 | No | ||
| 124 | HOXD10 | 16566 | -0.831 | -0.3692 | No | ||
| 125 | NAV1 | 16592 | -0.834 | -0.3673 | No | ||
| 126 | ITGA7 | 16758 | -0.855 | -0.3718 | No | ||
| 127 | HOXC5 | 16896 | -0.874 | -0.3749 | No | ||
| 128 | WNT5A | 17118 | -0.911 | -0.3818 | No | ||
| 129 | GPR20 | 17162 | -0.917 | -0.3804 | No | ||
| 130 | ARPC4 | 17232 | -0.928 | -0.3803 | No | ||
| 131 | DHH | 17280 | -0.934 | -0.3790 | No | ||
| 132 | JPH2 | 17302 | -0.938 | -0.3766 | No | ||
| 133 | ACTG2 | 17644 | -0.997 | -0.3887 | No | ||
| 134 | SLC4A3 | 17725 | -1.011 | -0.3887 | No | ||
| 135 | RSU1 | 17862 | -1.035 | -0.3912 | No | ||
| 136 | HOXA5 | 18015 | -1.061 | -0.3943 | No | ||
| 137 | CABP1 | 18101 | -1.076 | -0.3944 | Yes | ||
| 138 | ATF4 | 18125 | -1.079 | -0.3915 | Yes | ||
| 139 | CDKN1B | 18144 | -1.082 | -0.3884 | Yes | ||
| 140 | VCL | 18239 | -1.100 | -0.3888 | Yes | ||
| 141 | CSNK1E | 18249 | -1.101 | -0.3852 | Yes | ||
| 142 | UNC84B | 18292 | -1.111 | -0.3831 | Yes | ||
| 143 | CFL1 | 18320 | -1.119 | -0.3803 | Yes | ||
| 144 | CALD1 | 18341 | -1.123 | -0.3771 | Yes | ||
| 145 | DUSP5 | 18475 | -1.149 | -0.3791 | Yes | ||
| 146 | PDLIM7 | 18630 | -1.180 | -0.3819 | Yes | ||
| 147 | TPM2 | 18762 | -1.208 | -0.3835 | Yes | ||
| 148 | PDLIM3 | 18840 | -1.226 | -0.3826 | Yes | ||
| 149 | LDHA | 18972 | -1.257 | -0.3841 | Yes | ||
| 150 | COL1A2 | 19064 | -1.274 | -0.3837 | Yes | ||
| 151 | GRK6 | 19126 | -1.289 | -0.3818 | Yes | ||
| 152 | SVIL | 19147 | -1.295 | -0.3781 | Yes | ||
| 153 | SLC25A4 | 19161 | -1.300 | -0.3739 | Yes | ||
| 154 | RUNDC1 | 19296 | -1.339 | -0.3753 | Yes | ||
| 155 | MYL6 | 19370 | -1.358 | -0.3737 | Yes | ||
| 156 | ABR | 19483 | -1.389 | -0.3738 | Yes | ||
| 157 | NPAS2 | 19533 | -1.405 | -0.3710 | Yes | ||
| 158 | TITF1 | 19597 | -1.423 | -0.3687 | Yes | ||
| 159 | NR2F1 | 19622 | -1.432 | -0.3646 | Yes | ||
| 160 | KCNMB1 | 19719 | -1.468 | -0.3637 | Yes | ||
| 161 | TRDN | 19735 | -1.473 | -0.3591 | Yes | ||
| 162 | ACTA1 | 20070 | -1.597 | -0.3687 | Yes | ||
| 163 | FOXP2 | 20114 | -1.616 | -0.3648 | Yes | ||
| 164 | TPM3 | 20175 | -1.642 | -0.3616 | Yes | ||
| 165 | CX3CL1 | 20401 | -1.741 | -0.3657 | Yes | ||
| 166 | DMD | 20507 | -1.799 | -0.3640 | Yes | ||
| 167 | GADD45G | 20517 | -1.805 | -0.3578 | Yes | ||
| 168 | SRF | 20535 | -1.816 | -0.3520 | Yes | ||
| 169 | TLE3 | 20574 | -1.836 | -0.3471 | Yes | ||
| 170 | DUSP10 | 20732 | -1.927 | -0.3474 | Yes | ||
| 171 | LSP1 | 20764 | -1.943 | -0.3418 | Yes | ||
| 172 | NNAT | 20797 | -1.968 | -0.3361 | Yes | ||
| 173 | SIN3A | 20857 | -2.001 | -0.3316 | Yes | ||
| 174 | PRELP | 20955 | -2.071 | -0.3285 | Yes | ||
| 175 | KIF1B | 20997 | -2.115 | -0.3228 | Yes | ||
| 176 | NR2F2 | 21012 | -2.123 | -0.3157 | Yes | ||
| 177 | SLC7A1 | 21110 | -2.202 | -0.3122 | Yes | ||
| 178 | STAT5B | 21219 | -2.310 | -0.3088 | Yes | ||
| 179 | INSM1 | 21275 | -2.374 | -0.3028 | Yes | ||
| 180 | FOXP3 | 21314 | -2.435 | -0.2957 | Yes | ||
| 181 | HIVEP3 | 21332 | -2.463 | -0.2875 | Yes | ||
| 182 | SIPA1 | 21344 | -2.478 | -0.2791 | Yes | ||
| 183 | CITED2 | 21435 | -2.655 | -0.2736 | Yes | ||
| 184 | JUNB | 21440 | -2.670 | -0.2641 | Yes | ||
| 185 | RASSF2 | 21452 | -2.692 | -0.2549 | Yes | ||
| 186 | BDNF | 21491 | -2.758 | -0.2466 | Yes | ||
| 187 | FLNA | 21593 | -3.079 | -0.2401 | Yes | ||
| 188 | MYADM | 21637 | -3.193 | -0.2305 | Yes | ||
| 189 | TES | 21647 | -3.234 | -0.2192 | Yes | ||
| 190 | CORO1C | 21703 | -3.548 | -0.2089 | Yes | ||
| 191 | RERE | 21714 | -3.605 | -0.1963 | Yes | ||
| 192 | DUSP2 | 21727 | -3.672 | -0.1836 | Yes | ||
| 193 | CNN2 | 21797 | -4.241 | -0.1714 | Yes | ||
| 194 | ANXA6 | 21887 | -5.560 | -0.1554 | Yes | ||
| 195 | FOS | 21917 | -6.478 | -0.1332 | Yes | ||
| 196 | NR4A1 | 21923 | -6.633 | -0.1095 | Yes | ||
| 197 | EGR1 | 21942 | -8.847 | -0.0782 | Yes | ||
| 198 | EGR3 | 21943 | -9.744 | -0.0430 | Yes | ||
| 199 | EGR2 | 21944 | -11.902 | 0.0001 | Yes |