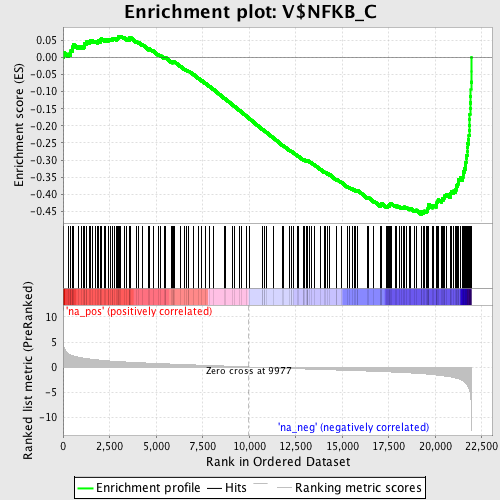
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | set02_ATM_minus_versus_BT_ATM_minus |
| Phenotype | NoPhenotypeAvailable |
| Upregulated in class | na_neg |
| GeneSet | V$NFKB_C |
| Enrichment Score (ES) | -0.45945 |
| Normalized Enrichment Score (NES) | -2.0807068 |
| Nominal p-value | 0.0 |
| FDR q-value | 2.5340138E-4 |
| FWER p-Value | 0.0010 |

| PROBE | GENE SYMBOL | GENE_TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|---|
| 1 | IL2RA | 15 | 4.747 | 0.0153 | No | ||
| 2 | ASH1L | 314 | 2.583 | 0.0103 | No | ||
| 3 | ANKHD1 | 395 | 2.431 | 0.0148 | No | ||
| 4 | SLC43A1 | 420 | 2.395 | 0.0218 | No | ||
| 5 | GABPB2 | 504 | 2.267 | 0.0256 | No | ||
| 6 | NLK | 525 | 2.243 | 0.0323 | No | ||
| 7 | CDK6 | 570 | 2.184 | 0.0376 | No | ||
| 8 | FGF17 | 816 | 1.949 | 0.0329 | No | ||
| 9 | VCAM1 | 984 | 1.831 | 0.0314 | No | ||
| 10 | PTGES | 1114 | 1.751 | 0.0313 | No | ||
| 11 | RAD18 | 1160 | 1.728 | 0.0351 | No | ||
| 12 | PHF6 | 1161 | 1.728 | 0.0409 | No | ||
| 13 | TCTA | 1239 | 1.688 | 0.0431 | No | ||
| 14 | TAZ | 1276 | 1.668 | 0.0470 | No | ||
| 15 | GPHN | 1432 | 1.593 | 0.0453 | No | ||
| 16 | CHD4 | 1458 | 1.578 | 0.0494 | No | ||
| 17 | BMF | 1578 | 1.530 | 0.0491 | No | ||
| 18 | FLI1 | 1719 | 1.474 | 0.0477 | No | ||
| 19 | PTPRJ | 1863 | 1.408 | 0.0458 | No | ||
| 20 | PPP2R5E | 1899 | 1.393 | 0.0489 | No | ||
| 21 | NFAT5 | 1993 | 1.363 | 0.0492 | No | ||
| 22 | SMARCAD1 | 2010 | 1.359 | 0.0531 | No | ||
| 23 | WRN | 2060 | 1.342 | 0.0553 | No | ||
| 24 | BCL2L1 | 2231 | 1.283 | 0.0518 | No | ||
| 25 | SOCS2 | 2303 | 1.259 | 0.0528 | No | ||
| 26 | ARHGAP5 | 2417 | 1.220 | 0.0517 | No | ||
| 27 | SOX5 | 2456 | 1.209 | 0.0541 | No | ||
| 28 | ITGB4 | 2549 | 1.187 | 0.0538 | No | ||
| 29 | GNA13 | 2665 | 1.161 | 0.0525 | No | ||
| 30 | RORC | 2672 | 1.158 | 0.0561 | No | ||
| 31 | DDR1 | 2758 | 1.137 | 0.0560 | No | ||
| 32 | RAB10 | 2870 | 1.112 | 0.0547 | No | ||
| 33 | ALG6 | 2907 | 1.105 | 0.0567 | No | ||
| 34 | CCDC8 | 2958 | 1.093 | 0.0581 | No | ||
| 35 | PLP1 | 2968 | 1.091 | 0.0614 | No | ||
| 36 | CTGF | 3039 | 1.073 | 0.0618 | No | ||
| 37 | TOP1 | 3099 | 1.061 | 0.0626 | No | ||
| 38 | IL18R1 | 3276 | 1.018 | 0.0580 | No | ||
| 39 | GNAO1 | 3428 | 0.985 | 0.0543 | No | ||
| 40 | RAP2C | 3553 | 0.957 | 0.0519 | No | ||
| 41 | PPP3CA | 3559 | 0.957 | 0.0549 | No | ||
| 42 | BCL11A | 3568 | 0.955 | 0.0577 | No | ||
| 43 | SNAP25 | 3593 | 0.949 | 0.0598 | No | ||
| 44 | NUP153 | 3954 | 0.880 | 0.0462 | No | ||
| 45 | SLC6A12 | 4059 | 0.862 | 0.0443 | No | ||
| 46 | SEC14L2 | 4279 | 0.825 | 0.0370 | No | ||
| 47 | CGGBP1 | 4612 | 0.770 | 0.0244 | No | ||
| 48 | CCL5 | 4636 | 0.767 | 0.0259 | No | ||
| 49 | IL1RAPL1 | 4833 | 0.734 | 0.0194 | No | ||
| 50 | ANP32A | 5102 | 0.695 | 0.0094 | No | ||
| 51 | LRRFIP2 | 5218 | 0.677 | 0.0064 | No | ||
| 52 | RIN2 | 5426 | 0.645 | -0.0010 | No | ||
| 53 | EIF5A | 5477 | 0.638 | -0.0011 | No | ||
| 54 | DNAJA1 | 5505 | 0.633 | -0.0002 | No | ||
| 55 | ECT2 | 5806 | 0.590 | -0.0120 | No | ||
| 56 | PAPPA | 5897 | 0.575 | -0.0142 | No | ||
| 57 | CHX10 | 5933 | 0.569 | -0.0139 | No | ||
| 58 | COL11A2 | 5940 | 0.568 | -0.0123 | No | ||
| 59 | FGF1 | 5996 | 0.558 | -0.0129 | No | ||
| 60 | MOBKL2C | 6294 | 0.513 | -0.0249 | No | ||
| 61 | TSNAXIP1 | 6539 | 0.482 | -0.0345 | No | ||
| 62 | PLA1A | 6620 | 0.470 | -0.0366 | No | ||
| 63 | C1QL1 | 6719 | 0.457 | -0.0395 | No | ||
| 64 | RGL1 | 6759 | 0.451 | -0.0398 | No | ||
| 65 | HOXB6 | 7009 | 0.419 | -0.0498 | No | ||
| 66 | EDG5 | 7274 | 0.384 | -0.0607 | No | ||
| 67 | FUT7 | 7456 | 0.359 | -0.0678 | No | ||
| 68 | OPCML | 7654 | 0.332 | -0.0757 | No | ||
| 69 | BCKDK | 7877 | 0.303 | -0.0849 | No | ||
| 70 | IFNB1 | 8083 | 0.276 | -0.0934 | No | ||
| 71 | KCNQ5 | 8660 | 0.195 | -0.1192 | No | ||
| 72 | IL23A | 8716 | 0.188 | -0.1211 | No | ||
| 73 | HOXB9 | 9119 | 0.131 | -0.1392 | No | ||
| 74 | UACA | 9191 | 0.121 | -0.1420 | No | ||
| 75 | ARHGAP8 | 9477 | 0.078 | -0.1549 | No | ||
| 76 | FOXD2 | 9581 | 0.064 | -0.1594 | No | ||
| 77 | SIX5 | 9872 | 0.016 | -0.1727 | No | ||
| 78 | FOXD3 | 10024 | -0.009 | -0.1796 | No | ||
| 79 | VNN3 | 10699 | -0.107 | -0.2102 | No | ||
| 80 | MSX1 | 10717 | -0.110 | -0.2106 | No | ||
| 81 | EGF | 10831 | -0.129 | -0.2154 | No | ||
| 82 | MSC | 10845 | -0.131 | -0.2156 | No | ||
| 83 | IL1A | 10944 | -0.147 | -0.2196 | No | ||
| 84 | CLCN1 | 11298 | -0.194 | -0.2351 | No | ||
| 85 | CDH5 | 11800 | -0.250 | -0.2573 | No | ||
| 86 | TRIM47 | 11828 | -0.255 | -0.2577 | No | ||
| 87 | ARPC5 | 12171 | -0.292 | -0.2725 | No | ||
| 88 | TSLP | 12250 | -0.301 | -0.2750 | No | ||
| 89 | MAG | 12268 | -0.303 | -0.2748 | No | ||
| 90 | CENTB5 | 12387 | -0.319 | -0.2791 | No | ||
| 91 | LAMB3 | 12570 | -0.341 | -0.2864 | No | ||
| 92 | LIN28 | 12659 | -0.351 | -0.2892 | No | ||
| 93 | TAL1 | 12910 | -0.380 | -0.2994 | No | ||
| 94 | PURG | 12929 | -0.383 | -0.2990 | No | ||
| 95 | SLC34A3 | 12955 | -0.385 | -0.2988 | No | ||
| 96 | EPHB6 | 13096 | -0.402 | -0.3039 | No | ||
| 97 | SMOC1 | 13129 | -0.405 | -0.3040 | No | ||
| 98 | LCAT | 13146 | -0.408 | -0.3034 | No | ||
| 99 | ASCL3 | 13150 | -0.408 | -0.3022 | No | ||
| 100 | MAP3K8 | 13234 | -0.417 | -0.3046 | No | ||
| 101 | PFN1 | 13354 | -0.432 | -0.3086 | No | ||
| 102 | JPH3 | 13493 | -0.450 | -0.3134 | No | ||
| 103 | SYMPK | 13814 | -0.488 | -0.3265 | No | ||
| 104 | PTHLH | 14038 | -0.513 | -0.3350 | No | ||
| 105 | DUSP22 | 14089 | -0.518 | -0.3356 | No | ||
| 106 | AGC1 | 14232 | -0.536 | -0.3403 | No | ||
| 107 | RND1 | 14328 | -0.548 | -0.3428 | No | ||
| 108 | ZADH2 | 14690 | -0.595 | -0.3574 | No | ||
| 109 | MAP3K13 | 14712 | -0.597 | -0.3564 | No | ||
| 110 | GRIN2D | 14936 | -0.622 | -0.3645 | No | ||
| 111 | IL13 | 15257 | -0.658 | -0.3770 | No | ||
| 112 | SIRT2 | 15387 | -0.675 | -0.3807 | No | ||
| 113 | GATA4 | 15522 | -0.689 | -0.3845 | No | ||
| 114 | SMPD3 | 15637 | -0.706 | -0.3874 | No | ||
| 115 | BAI2 | 15719 | -0.717 | -0.3887 | No | ||
| 116 | MRPS6 | 15817 | -0.731 | -0.3907 | No | ||
| 117 | SESN2 | 15842 | -0.734 | -0.3893 | No | ||
| 118 | AMOTL1 | 16365 | -0.806 | -0.4106 | No | ||
| 119 | NFKBIA | 16391 | -0.809 | -0.4090 | No | ||
| 120 | HSD3B7 | 16699 | -0.847 | -0.4203 | No | ||
| 121 | PTGS2 | 17046 | -0.900 | -0.4331 | No | ||
| 122 | HOXA11 | 17053 | -0.901 | -0.4304 | No | ||
| 123 | CDC14A | 17105 | -0.909 | -0.4297 | No | ||
| 124 | RALGDS | 17121 | -0.911 | -0.4273 | No | ||
| 125 | WNT10A | 17395 | -0.953 | -0.4366 | No | ||
| 126 | SOX10 | 17450 | -0.962 | -0.4359 | No | ||
| 127 | RHOA | 17480 | -0.969 | -0.4339 | No | ||
| 128 | SNX15 | 17529 | -0.977 | -0.4329 | No | ||
| 129 | MAML2 | 17538 | -0.979 | -0.4299 | No | ||
| 130 | CUEDC1 | 17576 | -0.986 | -0.4283 | No | ||
| 131 | SOX3 | 17626 | -0.995 | -0.4272 | No | ||
| 132 | LRCH1 | 17840 | -1.029 | -0.4335 | No | ||
| 133 | ITGB8 | 17931 | -1.048 | -0.4341 | No | ||
| 134 | IL1RN | 18058 | -1.069 | -0.4363 | No | ||
| 135 | FOXA3 | 18182 | -1.089 | -0.4383 | No | ||
| 136 | UNC84B | 18292 | -1.111 | -0.4396 | No | ||
| 137 | LYPLA2 | 18325 | -1.120 | -0.4373 | No | ||
| 138 | EFNA3 | 18430 | -1.141 | -0.4382 | No | ||
| 139 | DDX17 | 18608 | -1.175 | -0.4424 | No | ||
| 140 | BLR1 | 18680 | -1.189 | -0.4417 | No | ||
| 141 | NFKBIB | 18902 | -1.241 | -0.4476 | No | ||
| 142 | CYLD | 18965 | -1.255 | -0.4463 | No | ||
| 143 | DCAMKL1 | 19253 | -1.325 | -0.4550 | Yes | ||
| 144 | RHOG | 19258 | -1.326 | -0.4507 | Yes | ||
| 145 | TNIP1 | 19385 | -1.362 | -0.4519 | Yes | ||
| 146 | SCRT2 | 19427 | -1.373 | -0.4492 | Yes | ||
| 147 | CDC37 | 19551 | -1.410 | -0.4501 | Yes | ||
| 148 | RANBP10 | 19574 | -1.417 | -0.4463 | Yes | ||
| 149 | TCEA2 | 19600 | -1.424 | -0.4427 | Yes | ||
| 150 | S100A10 | 19616 | -1.430 | -0.4385 | Yes | ||
| 151 | DAP3 | 19633 | -1.434 | -0.4344 | Yes | ||
| 152 | DACH2 | 19644 | -1.437 | -0.4300 | Yes | ||
| 153 | CXCL10 | 19873 | -1.519 | -0.4354 | Yes | ||
| 154 | SEMA3B | 19915 | -1.535 | -0.4321 | Yes | ||
| 155 | LTB | 20040 | -1.586 | -0.4325 | Yes | ||
| 156 | GNG4 | 20044 | -1.587 | -0.4273 | Yes | ||
| 157 | PCSK2 | 20085 | -1.604 | -0.4237 | Yes | ||
| 158 | TNFSF7 | 20122 | -1.618 | -0.4199 | Yes | ||
| 159 | EHF | 20162 | -1.636 | -0.4162 | Yes | ||
| 160 | TYRO3 | 20318 | -1.705 | -0.4176 | Yes | ||
| 161 | SLAMF8 | 20358 | -1.724 | -0.4136 | Yes | ||
| 162 | TRIB2 | 20441 | -1.762 | -0.4114 | Yes | ||
| 163 | MNT | 20472 | -1.778 | -0.4068 | Yes | ||
| 164 | PRKCD | 20514 | -1.805 | -0.4026 | Yes | ||
| 165 | LCN2 | 20596 | -1.844 | -0.4001 | Yes | ||
| 166 | FOXJ2 | 20816 | -1.975 | -0.4035 | Yes | ||
| 167 | BAHD1 | 20822 | -1.978 | -0.3971 | Yes | ||
| 168 | SIN3A | 20857 | -2.001 | -0.3919 | Yes | ||
| 169 | HCFC1 | 20984 | -2.097 | -0.3906 | Yes | ||
| 170 | USP52 | 21073 | -2.166 | -0.3874 | Yes | ||
| 171 | ICAM1 | 21114 | -2.204 | -0.3818 | Yes | ||
| 172 | UBE2I | 21139 | -2.226 | -0.3754 | Yes | ||
| 173 | PIGC | 21197 | -2.289 | -0.3703 | Yes | ||
| 174 | NDRG2 | 21223 | -2.314 | -0.3636 | Yes | ||
| 175 | GRK5 | 21250 | -2.344 | -0.3569 | Yes | ||
| 176 | BHLHB2 | 21329 | -2.459 | -0.3522 | Yes | ||
| 177 | SP6 | 21482 | -2.751 | -0.3500 | Yes | ||
| 178 | BDNF | 21491 | -2.758 | -0.3410 | Yes | ||
| 179 | STAT6 | 21519 | -2.830 | -0.3327 | Yes | ||
| 180 | JAK3 | 21578 | -3.015 | -0.3252 | Yes | ||
| 181 | NFKB2 | 21615 | -3.137 | -0.3163 | Yes | ||
| 182 | IRF1 | 21636 | -3.191 | -0.3065 | Yes | ||
| 183 | ZHX2 | 21654 | -3.267 | -0.2963 | Yes | ||
| 184 | IER5 | 21661 | -3.308 | -0.2854 | Yes | ||
| 185 | ENO3 | 21711 | -3.589 | -0.2755 | Yes | ||
| 186 | CD74 | 21718 | -3.626 | -0.2636 | Yes | ||
| 187 | XPO6 | 21734 | -3.713 | -0.2518 | Yes | ||
| 188 | ARHGEF2 | 21769 | -3.994 | -0.2399 | Yes | ||
| 189 | GADD45B | 21806 | -4.294 | -0.2271 | Yes | ||
| 190 | LASP1 | 21823 | -4.444 | -0.2128 | Yes | ||
| 191 | ID2 | 21838 | -4.620 | -0.1979 | Yes | ||
| 192 | IL4I1 | 21849 | -4.754 | -0.1824 | Yes | ||
| 193 | PCBP4 | 21856 | -4.849 | -0.1663 | Yes | ||
| 194 | BCL3 | 21865 | -4.977 | -0.1499 | Yes | ||
| 195 | IL27RA | 21874 | -5.232 | -0.1326 | Yes | ||
| 196 | CCL22 | 21893 | -5.744 | -0.1141 | Yes | ||
| 197 | PPP1R13B | 21915 | -6.417 | -0.0934 | Yes | ||
| 198 | FOS | 21917 | -6.478 | -0.0717 | Yes | ||
| 199 | EGR3 | 21943 | -9.744 | -0.0400 | Yes | ||
| 200 | EGR2 | 21944 | -11.902 | 0.0001 | Yes |
