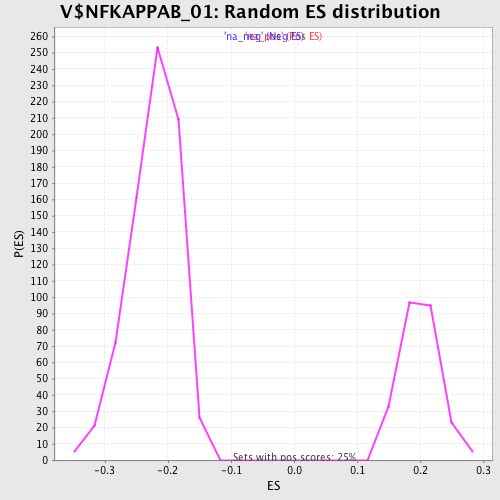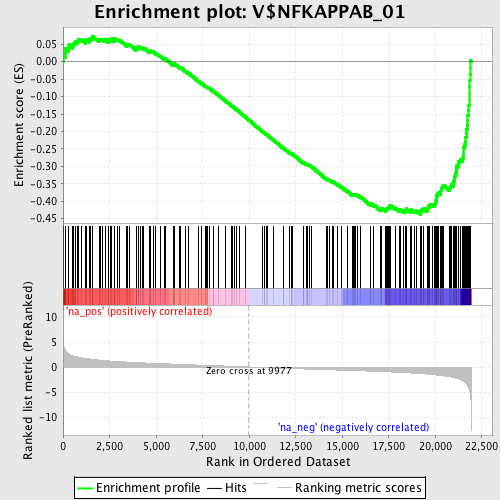
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | set02_ATM_minus_versus_BT_ATM_minus |
| Phenotype | NoPhenotypeAvailable |
| Upregulated in class | na_neg |
| GeneSet | V$NFKAPPAB_01 |
| Enrichment Score (ES) | -0.43795806 |
| Normalized Enrichment Score (NES) | -1.974129 |
| Nominal p-value | 0.0 |
| FDR q-value | 0.0022750648 |
| FWER p-Value | 0.013 |

| PROBE | GENE SYMBOL | GENE_TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|---|
| 1 | IL2RA | 15 | 4.747 | 0.0172 | No | ||
| 2 | CHD6 | 116 | 3.310 | 0.0252 | No | ||
| 3 | CREB1 | 126 | 3.257 | 0.0371 | No | ||
| 4 | SLC12A2 | 297 | 2.620 | 0.0391 | No | ||
| 5 | ASH1L | 314 | 2.583 | 0.0482 | No | ||
| 6 | NLK | 525 | 2.243 | 0.0470 | No | ||
| 7 | CDK6 | 570 | 2.184 | 0.0532 | No | ||
| 8 | RBMS1 | 666 | 2.092 | 0.0568 | No | ||
| 9 | JARID2 | 776 | 1.986 | 0.0593 | No | ||
| 10 | FGF17 | 816 | 1.949 | 0.0648 | No | ||
| 11 | VCAM1 | 984 | 1.831 | 0.0641 | No | ||
| 12 | GNB1 | 1222 | 1.696 | 0.0596 | No | ||
| 13 | TAZ | 1276 | 1.668 | 0.0635 | No | ||
| 14 | GPHN | 1432 | 1.593 | 0.0623 | No | ||
| 15 | CHD4 | 1458 | 1.578 | 0.0672 | No | ||
| 16 | ATP1B1 | 1557 | 1.539 | 0.0685 | No | ||
| 17 | BMF | 1578 | 1.530 | 0.0733 | No | ||
| 18 | PPARBP | 1935 | 1.380 | 0.0622 | No | ||
| 19 | NFAT5 | 1993 | 1.363 | 0.0647 | No | ||
| 20 | P2RY10 | 2098 | 1.330 | 0.0650 | No | ||
| 21 | BMP2K | 2252 | 1.276 | 0.0628 | No | ||
| 22 | ARHGAP5 | 2417 | 1.220 | 0.0598 | No | ||
| 23 | SOX5 | 2456 | 1.209 | 0.0627 | No | ||
| 24 | ITGB4 | 2549 | 1.187 | 0.0629 | No | ||
| 25 | BNC2 | 2586 | 1.180 | 0.0657 | No | ||
| 26 | DDR1 | 2758 | 1.137 | 0.0621 | No | ||
| 27 | LAMA1 | 2771 | 1.133 | 0.0659 | No | ||
| 28 | ALG6 | 2907 | 1.105 | 0.0639 | No | ||
| 29 | CTGF | 3039 | 1.073 | 0.0619 | No | ||
| 30 | GNAO1 | 3428 | 0.985 | 0.0478 | No | ||
| 31 | CXCL9 | 3484 | 0.973 | 0.0489 | No | ||
| 32 | RAP2C | 3553 | 0.957 | 0.0494 | No | ||
| 33 | ERN1 | 3917 | 0.886 | 0.0361 | No | ||
| 34 | FBXL12 | 3928 | 0.885 | 0.0390 | No | ||
| 35 | KCNS3 | 3945 | 0.881 | 0.0416 | No | ||
| 36 | SLC6A12 | 4059 | 0.862 | 0.0396 | No | ||
| 37 | STC2 | 4062 | 0.861 | 0.0428 | No | ||
| 38 | AP1S2 | 4155 | 0.845 | 0.0418 | No | ||
| 39 | SEC14L2 | 4279 | 0.825 | 0.0392 | No | ||
| 40 | NOL4 | 4345 | 0.814 | 0.0393 | No | ||
| 41 | CCL5 | 4636 | 0.767 | 0.0289 | No | ||
| 42 | KRTAP13-1 | 4694 | 0.757 | 0.0291 | No | ||
| 43 | KCNH3 | 4721 | 0.753 | 0.0308 | No | ||
| 44 | IL1RAPL1 | 4833 | 0.734 | 0.0284 | No | ||
| 45 | UBE2D3 | 4979 | 0.711 | 0.0245 | No | ||
| 46 | RIMS2 | 5217 | 0.677 | 0.0161 | No | ||
| 47 | RIN2 | 5426 | 0.645 | 0.0090 | No | ||
| 48 | EIF5A | 5477 | 0.638 | 0.0091 | No | ||
| 49 | CHX10 | 5933 | 0.569 | -0.0096 | No | ||
| 50 | COL11A2 | 5940 | 0.568 | -0.0078 | No | ||
| 51 | ZBTB11 | 5985 | 0.561 | -0.0077 | No | ||
| 52 | FGF1 | 5996 | 0.558 | -0.0060 | No | ||
| 53 | AGPAT1 | 6276 | 0.515 | -0.0169 | No | ||
| 54 | MOBKL2C | 6294 | 0.513 | -0.0157 | No | ||
| 55 | PLAU | 6575 | 0.477 | -0.0268 | No | ||
| 56 | C1QL1 | 6719 | 0.457 | -0.0317 | No | ||
| 57 | EDG5 | 7274 | 0.384 | -0.0557 | No | ||
| 58 | FUT7 | 7456 | 0.359 | -0.0626 | No | ||
| 59 | OPCML | 7654 | 0.332 | -0.0704 | No | ||
| 60 | ATOH1 | 7690 | 0.328 | -0.0708 | No | ||
| 61 | BFSP1 | 7760 | 0.320 | -0.0728 | No | ||
| 62 | BCKDK | 7877 | 0.303 | -0.0769 | No | ||
| 63 | KLK9 | 8062 | 0.279 | -0.0843 | No | ||
| 64 | IFNB1 | 8083 | 0.276 | -0.0842 | No | ||
| 65 | FLRT1 | 8357 | 0.238 | -0.0959 | No | ||
| 66 | IL23A | 8716 | 0.188 | -0.1116 | No | ||
| 67 | SALL1 | 9026 | 0.143 | -0.1253 | No | ||
| 68 | HOXB9 | 9119 | 0.131 | -0.1290 | No | ||
| 69 | UACA | 9191 | 0.121 | -0.1318 | No | ||
| 70 | SAMSN1 | 9342 | 0.099 | -0.1383 | No | ||
| 71 | ARHGAP8 | 9477 | 0.078 | -0.1442 | No | ||
| 72 | PRX | 9775 | 0.032 | -0.1577 | No | ||
| 73 | MSX1 | 10717 | -0.110 | -0.2006 | No | ||
| 74 | EGF | 10831 | -0.129 | -0.2053 | No | ||
| 75 | MSC | 10845 | -0.131 | -0.2054 | No | ||
| 76 | IL1A | 10944 | -0.147 | -0.2093 | No | ||
| 77 | WNT10B | 10979 | -0.152 | -0.2103 | No | ||
| 78 | CLCN1 | 11298 | -0.194 | -0.2242 | No | ||
| 79 | ACTN3 | 11825 | -0.254 | -0.2474 | No | ||
| 80 | TRIM47 | 11828 | -0.255 | -0.2465 | No | ||
| 81 | COL16A1 | 12170 | -0.292 | -0.2611 | No | ||
| 82 | TSLP | 12250 | -0.301 | -0.2636 | No | ||
| 83 | MAG | 12268 | -0.303 | -0.2632 | No | ||
| 84 | GREM1 | 12314 | -0.311 | -0.2641 | No | ||
| 85 | TAL1 | 12910 | -0.380 | -0.2900 | No | ||
| 86 | BCL6B | 12936 | -0.383 | -0.2897 | No | ||
| 87 | CXCL5 | 13063 | -0.399 | -0.2940 | No | ||
| 88 | SMOC1 | 13129 | -0.405 | -0.2955 | No | ||
| 89 | ASCL3 | 13150 | -0.408 | -0.2948 | No | ||
| 90 | MAP3K8 | 13234 | -0.417 | -0.2971 | No | ||
| 91 | PFN1 | 13354 | -0.432 | -0.3009 | No | ||
| 92 | ZBTB5 | 14161 | -0.528 | -0.3360 | No | ||
| 93 | AGC1 | 14232 | -0.536 | -0.3372 | No | ||
| 94 | RND1 | 14328 | -0.548 | -0.3395 | No | ||
| 95 | PUNC | 14461 | -0.564 | -0.3434 | No | ||
| 96 | ZFP36L2 | 14528 | -0.574 | -0.3443 | No | ||
| 97 | ARPC2 | 14768 | -0.602 | -0.3530 | No | ||
| 98 | GRIN2D | 14936 | -0.622 | -0.3583 | No | ||
| 99 | IL13 | 15257 | -0.658 | -0.3705 | No | ||
| 100 | GATA4 | 15522 | -0.689 | -0.3800 | No | ||
| 101 | LIG1 | 15585 | -0.699 | -0.3802 | No | ||
| 102 | SMPD3 | 15637 | -0.706 | -0.3799 | No | ||
| 103 | BAI2 | 15719 | -0.717 | -0.3809 | No | ||
| 104 | SESN2 | 15842 | -0.734 | -0.3838 | No | ||
| 105 | ACTN1 | 15953 | -0.750 | -0.3860 | No | ||
| 106 | UBD | 16505 | -0.824 | -0.4082 | No | ||
| 107 | CXCL11 | 16542 | -0.828 | -0.4067 | No | ||
| 108 | HSD3B7 | 16699 | -0.847 | -0.4107 | No | ||
| 109 | HOXA11 | 17053 | -0.901 | -0.4235 | No | ||
| 110 | CDC14A | 17105 | -0.909 | -0.4224 | No | ||
| 111 | RALGDS | 17121 | -0.911 | -0.4197 | No | ||
| 112 | BIRC3 | 17339 | -0.944 | -0.4261 | No | ||
| 113 | RRAS | 17359 | -0.947 | -0.4234 | No | ||
| 114 | WNT10A | 17395 | -0.953 | -0.4214 | No | ||
| 115 | SOX10 | 17450 | -0.962 | -0.4202 | No | ||
| 116 | SDC4 | 17459 | -0.965 | -0.4169 | No | ||
| 117 | MAML2 | 17538 | -0.979 | -0.4168 | No | ||
| 118 | ABHD8 | 17544 | -0.980 | -0.4134 | No | ||
| 119 | CXCL2 | 17607 | -0.991 | -0.4125 | No | ||
| 120 | LRCH1 | 17840 | -1.029 | -0.4192 | No | ||
| 121 | IL1RN | 18058 | -1.069 | -0.4252 | No | ||
| 122 | RPS6KA4 | 18117 | -1.078 | -0.4238 | No | ||
| 123 | UNC84B | 18292 | -1.111 | -0.4276 | No | ||
| 124 | UBE2H | 18421 | -1.140 | -0.4291 | No | ||
| 125 | TLX1 | 18435 | -1.142 | -0.4254 | No | ||
| 126 | FKHL18 | 18450 | -1.144 | -0.4217 | No | ||
| 127 | BLR1 | 18680 | -1.189 | -0.4278 | No | ||
| 128 | NTN1 | 18700 | -1.194 | -0.4241 | No | ||
| 129 | NFKBIB | 18902 | -1.241 | -0.4287 | No | ||
| 130 | CYLD | 18965 | -1.255 | -0.4268 | No | ||
| 131 | CPD | 19209 | -1.313 | -0.4330 | Yes | ||
| 132 | KCNN3 | 19213 | -1.314 | -0.4282 | Yes | ||
| 133 | DCAMKL1 | 19253 | -1.325 | -0.4250 | Yes | ||
| 134 | TNF | 19341 | -1.351 | -0.4238 | Yes | ||
| 135 | TNIP1 | 19385 | -1.362 | -0.4207 | Yes | ||
| 136 | MADCAM1 | 19568 | -1.416 | -0.4237 | Yes | ||
| 137 | TCEA2 | 19600 | -1.424 | -0.4197 | Yes | ||
| 138 | DAP3 | 19633 | -1.434 | -0.4158 | Yes | ||
| 139 | TP53 | 19687 | -1.455 | -0.4127 | Yes | ||
| 140 | MSN | 19710 | -1.464 | -0.4082 | Yes | ||
| 141 | CXCL10 | 19873 | -1.519 | -0.4099 | Yes | ||
| 142 | SLC11A2 | 19972 | -1.555 | -0.4085 | Yes | ||
| 143 | EDN2 | 19996 | -1.569 | -0.4037 | Yes | ||
| 144 | HIVEP1 | 20031 | -1.583 | -0.3992 | Yes | ||
| 145 | LTB | 20040 | -1.586 | -0.3936 | Yes | ||
| 146 | GNG4 | 20044 | -1.587 | -0.3878 | Yes | ||
| 147 | PCSK2 | 20085 | -1.604 | -0.3835 | Yes | ||
| 148 | TNFSF7 | 20122 | -1.618 | -0.3791 | Yes | ||
| 149 | EHF | 20162 | -1.636 | -0.3747 | Yes | ||
| 150 | HCST | 20297 | -1.698 | -0.3744 | Yes | ||
| 151 | UBE4B | 20303 | -1.700 | -0.3682 | Yes | ||
| 152 | C1ORF24 | 20322 | -1.708 | -0.3626 | Yes | ||
| 153 | SLAMF8 | 20358 | -1.724 | -0.3577 | Yes | ||
| 154 | TRIB2 | 20441 | -1.762 | -0.3548 | Yes | ||
| 155 | LTA | 20785 | -1.958 | -0.3632 | Yes | ||
| 156 | FOXJ2 | 20816 | -1.975 | -0.3571 | Yes | ||
| 157 | SIN3A | 20857 | -2.001 | -0.3514 | Yes | ||
| 158 | HCFC1 | 20984 | -2.097 | -0.3492 | Yes | ||
| 159 | TRAF4 | 21000 | -2.116 | -0.3419 | Yes | ||
| 160 | NR2F2 | 21012 | -2.123 | -0.3344 | Yes | ||
| 161 | EBI3 | 21047 | -2.149 | -0.3279 | Yes | ||
| 162 | USP52 | 21073 | -2.166 | -0.3208 | Yes | ||
| 163 | ICAM1 | 21114 | -2.204 | -0.3143 | Yes | ||
| 164 | UBE2I | 21139 | -2.226 | -0.3070 | Yes | ||
| 165 | MAP4K2 | 21147 | -2.232 | -0.2989 | Yes | ||
| 166 | ETV6 | 21235 | -2.325 | -0.2941 | Yes | ||
| 167 | GRK5 | 21250 | -2.344 | -0.2859 | Yes | ||
| 168 | SHOX2 | 21345 | -2.479 | -0.2809 | Yes | ||
| 169 | SP6 | 21482 | -2.751 | -0.2767 | Yes | ||
| 170 | BDNF | 21491 | -2.758 | -0.2667 | Yes | ||
| 171 | STAT6 | 21519 | -2.830 | -0.2572 | Yes | ||
| 172 | ZBTB9 | 21520 | -2.835 | -0.2465 | Yes | ||
| 173 | JAK3 | 21578 | -3.015 | -0.2377 | Yes | ||
| 174 | NFKB2 | 21615 | -3.137 | -0.2275 | Yes | ||
| 175 | IRF1 | 21636 | -3.191 | -0.2164 | Yes | ||
| 176 | IER5 | 21661 | -3.308 | -0.2050 | Yes | ||
| 177 | MMP9 | 21679 | -3.409 | -0.1929 | Yes | ||
| 178 | ENO3 | 21711 | -3.589 | -0.1807 | Yes | ||
| 179 | ZMYND15 | 21743 | -3.778 | -0.1679 | Yes | ||
| 180 | RELB | 21749 | -3.826 | -0.1537 | Yes | ||
| 181 | TNFRSF9 | 21782 | -4.110 | -0.1396 | Yes | ||
| 182 | GADD45B | 21806 | -4.294 | -0.1244 | Yes | ||
| 183 | IL4I1 | 21849 | -4.754 | -0.1084 | Yes | ||
| 184 | TNFRSF1B | 21851 | -4.763 | -0.0905 | Yes | ||
| 185 | CXCL16 | 21853 | -4.800 | -0.0724 | Yes | ||
| 186 | PCBP4 | 21856 | -4.849 | -0.0541 | Yes | ||
| 187 | BCL3 | 21865 | -4.977 | -0.0357 | Yes | ||
| 188 | CD83 | 21872 | -5.185 | -0.0164 | Yes | ||
| 189 | IL27RA | 21874 | -5.232 | 0.0034 | Yes |