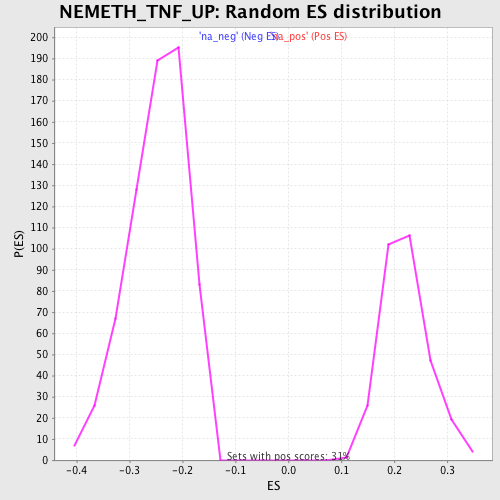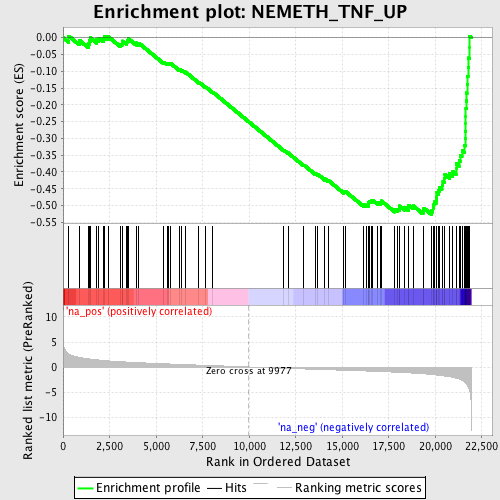
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | set02_ATM_minus_versus_BT_ATM_minus |
| Phenotype | NoPhenotypeAvailable |
| Upregulated in class | na_neg |
| GeneSet | NEMETH_TNF_UP |
| Enrichment Score (ES) | -0.5266431 |
| Normalized Enrichment Score (NES) | -2.1296558 |
| Nominal p-value | 0.0 |
| FDR q-value | 0.0014228467 |
| FWER p-Value | 0.024 |

| PROBE | GENE SYMBOL | GENE_TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|---|
| 1 | APBB1IP | 306 | 2.603 | 0.0045 | No | ||
| 2 | RNF11 | 868 | 1.907 | -0.0076 | No | ||
| 3 | PML | 1363 | 1.624 | -0.0186 | No | ||
| 4 | MYO10 | 1407 | 1.600 | -0.0092 | No | ||
| 5 | KPNA3 | 1465 | 1.575 | -0.0006 | No | ||
| 6 | IFI203 | 1808 | 1.436 | -0.0060 | No | ||
| 7 | IFI202B | 1923 | 1.383 | -0.0013 | No | ||
| 8 | IFRD1 | 2154 | 1.312 | -0.0025 | No | ||
| 9 | BCL2L1 | 2231 | 1.283 | 0.0031 | No | ||
| 10 | ETF1 | 2432 | 1.215 | 0.0026 | No | ||
| 11 | TOP1 | 3099 | 1.061 | -0.0203 | No | ||
| 12 | ACSL4 | 3166 | 1.045 | -0.0159 | No | ||
| 13 | GYK | 3200 | 1.037 | -0.0100 | No | ||
| 14 | CCL3 | 3426 | 0.986 | -0.0133 | No | ||
| 15 | CEBPD | 3462 | 0.977 | -0.0079 | No | ||
| 16 | IER3 | 3521 | 0.965 | -0.0037 | No | ||
| 17 | 2900009I07RIK | 3924 | 0.885 | -0.0158 | No | ||
| 18 | EIF1 | 4065 | 0.861 | -0.0161 | No | ||
| 19 | SLFN4 | 5415 | 0.646 | -0.0732 | No | ||
| 20 | SLC11A1 | 5596 | 0.623 | -0.0770 | No | ||
| 21 | PBEF1 | 5684 | 0.610 | -0.0766 | No | ||
| 22 | SLFN12 | 5771 | 0.596 | -0.0763 | No | ||
| 23 | CDKN1A | 6257 | 0.518 | -0.0948 | No | ||
| 24 | WSB1 | 6377 | 0.502 | -0.0967 | No | ||
| 25 | SOD2 | 6573 | 0.477 | -0.1022 | No | ||
| 26 | MARCKSL1 | 7299 | 0.380 | -0.1327 | No | ||
| 27 | CLEC4N | 7663 | 0.330 | -0.1469 | No | ||
| 28 | EIF2AK2 | 8037 | 0.283 | -0.1620 | No | ||
| 29 | CLN3 | 11867 | -0.258 | -0.3353 | No | ||
| 30 | C3AR1 | 12084 | -0.281 | -0.3432 | No | ||
| 31 | BCL6B | 12936 | -0.383 | -0.3794 | No | ||
| 32 | C5AR1 | 13581 | -0.458 | -0.4056 | No | ||
| 33 | PRG1 | 13691 | -0.472 | -0.4072 | No | ||
| 34 | JUND1 | 14067 | -0.516 | -0.4207 | No | ||
| 35 | GCH1 | 14251 | -0.539 | -0.4253 | No | ||
| 36 | MSR1 | 15091 | -0.638 | -0.4591 | No | ||
| 37 | HMGCR | 15157 | -0.647 | -0.4575 | No | ||
| 38 | TXNRD1 | 16140 | -0.775 | -0.4969 | No | ||
| 39 | CFLAR | 16298 | -0.798 | -0.4984 | No | ||
| 40 | MAPKAPK2 | 16390 | -0.809 | -0.4968 | No | ||
| 41 | NFKBIA | 16391 | -0.809 | -0.4910 | No | ||
| 42 | HCK | 16436 | -0.815 | -0.4872 | No | ||
| 43 | TPST1 | 16545 | -0.828 | -0.4863 | No | ||
| 44 | SAP30 | 16637 | -0.840 | -0.4845 | No | ||
| 45 | CASP4 | 16907 | -0.877 | -0.4905 | No | ||
| 46 | PTGS2 | 17046 | -0.900 | -0.4904 | No | ||
| 47 | NP | 17099 | -0.908 | -0.4863 | No | ||
| 48 | CCL9 | 17821 | -1.025 | -0.5120 | No | ||
| 49 | IL1RL1 | 17977 | -1.054 | -0.5116 | No | ||
| 50 | IL1RN | 18058 | -1.069 | -0.5077 | No | ||
| 51 | DAB2 | 18085 | -1.074 | -0.5012 | No | ||
| 52 | RCN1 | 18354 | -1.125 | -0.5055 | No | ||
| 53 | SLPI | 18560 | -1.165 | -0.5065 | No | ||
| 54 | TRIM25 | 18565 | -1.166 | -0.4984 | No | ||
| 55 | RHOC | 18812 | -1.221 | -0.5010 | No | ||
| 56 | TNF | 19341 | -1.351 | -0.5155 | No | ||
| 57 | TNIP1 | 19385 | -1.362 | -0.5078 | No | ||
| 58 | SQSTM1 | 19798 | -1.497 | -0.5160 | Yes | ||
| 59 | FURIN | 19875 | -1.519 | -0.5086 | Yes | ||
| 60 | MLLT11 | 19884 | -1.522 | -0.4982 | Yes | ||
| 61 | HCLS1 | 19929 | -1.540 | -0.4892 | Yes | ||
| 62 | LTB | 20040 | -1.586 | -0.4830 | Yes | ||
| 63 | ATF3 | 20071 | -1.597 | -0.4730 | Yes | ||
| 64 | TAP1 | 20082 | -1.603 | -0.4620 | Yes | ||
| 65 | STAT1 | 20172 | -1.640 | -0.4544 | Yes | ||
| 66 | ETS2 | 20232 | -1.668 | -0.4452 | Yes | ||
| 67 | TNFSF9 | 20359 | -1.724 | -0.4387 | Yes | ||
| 68 | LCP2 | 20409 | -1.744 | -0.4285 | Yes | ||
| 69 | MAP2K1 | 20473 | -1.779 | -0.4188 | Yes | ||
| 70 | PRKCD | 20514 | -1.805 | -0.4077 | Yes | ||
| 71 | DEGS1 | 20757 | -1.942 | -0.4050 | Yes | ||
| 72 | NFKB1 | 20944 | -2.065 | -0.3988 | Yes | ||
| 73 | ICAM1 | 21114 | -2.204 | -0.3908 | Yes | ||
| 74 | SLC2A1 | 21119 | -2.209 | -0.3753 | Yes | ||
| 75 | SAT1 | 21277 | -2.377 | -0.3655 | Yes | ||
| 76 | CD44 | 21333 | -2.464 | -0.3505 | Yes | ||
| 77 | CITED2 | 21435 | -2.655 | -0.3362 | Yes | ||
| 78 | TGFB1 | 21573 | -2.995 | -0.3212 | Yes | ||
| 79 | IGTP | 21608 | -3.115 | -0.3005 | Yes | ||
| 80 | SLC20A1 | 21618 | -3.142 | -0.2786 | Yes | ||
| 81 | MYADM | 21637 | -3.193 | -0.2567 | Yes | ||
| 82 | GLRX | 21641 | -3.213 | -0.2339 | Yes | ||
| 83 | IIGP2 | 21645 | -3.224 | -0.2111 | Yes | ||
| 84 | SLFN2 | 21669 | -3.350 | -0.1883 | Yes | ||
| 85 | FCGR2B | 21689 | -3.450 | -0.1646 | Yes | ||
| 86 | RGS16 | 21704 | -3.548 | -0.1400 | Yes | ||
| 87 | ADAR | 21719 | -3.627 | -0.1148 | Yes | ||
| 88 | STK10 | 21759 | -3.915 | -0.0887 | Yes | ||
| 89 | H2-D1 | 21787 | -4.139 | -0.0605 | Yes | ||
| 90 | IER2 | 21848 | -4.749 | -0.0294 | Yes | ||
| 91 | TNFRSF1B | 21851 | -4.763 | 0.0044 | Yes |