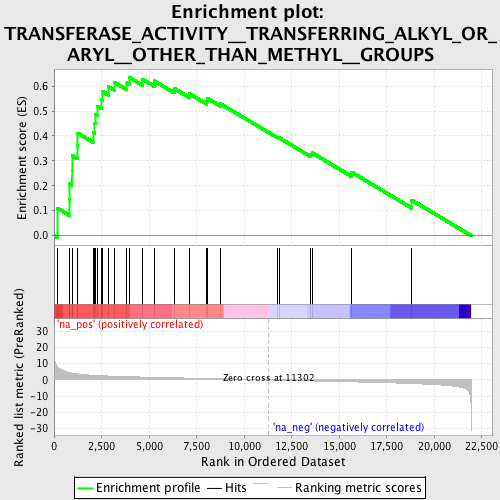
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | set01_ATM_minus_versus_SCID |
| Phenotype | NoPhenotypeAvailable |
| Upregulated in class | na_pos |
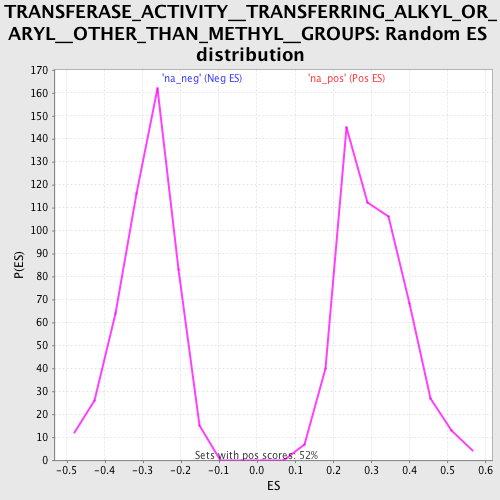
| GeneSet | TRANSFERASE_ACTIVITY__TRANSFERRING_ALKYL_OR_ARYL__OTHER_THAN_METHYL__GROUPS |
| Enrichment Score (ES) | 0.63443685 |
| Normalized Enrichment Score (NES) | 2.0705757 |
| Nominal p-value | 0.0 |
| FDR q-value | 0.008733807 |
| FWER p-Value | 0.076 |

| PROBE | GENE SYMBOL | GENE_TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|---|
| 1 | GGPS1 | 162 | 8.324 | 0.1104 | Yes | ||
| 2 | SMS | 794 | 4.561 | 0.1462 | Yes | ||
| 3 | GSTA4 | 829 | 4.474 | 0.2080 | Yes | ||
| 4 | PDSS1 | 941 | 4.211 | 0.2625 | Yes | ||
| 5 | GSTM1 | 943 | 4.207 | 0.3220 | Yes | ||
| 6 | GSTT2 | 1240 | 3.658 | 0.3603 | Yes | ||
| 7 | MGST1 | 1249 | 3.639 | 0.4115 | Yes | ||
| 8 | GSTM3 | 2048 | 2.811 | 0.4149 | Yes | ||
| 9 | GSTM2 | 2143 | 2.738 | 0.4493 | Yes | ||
| 10 | CHM | 2163 | 2.725 | 0.4870 | Yes | ||
| 11 | GSTT1 | 2263 | 2.665 | 0.5202 | Yes | ||
| 12 | PGGT1B | 2495 | 2.540 | 0.5457 | Yes | ||
| 13 | FNTB | 2564 | 2.502 | 0.5780 | Yes | ||
| 14 | RABGGTA | 2873 | 2.357 | 0.5973 | Yes | ||
| 15 | MAT2A | 3156 | 2.238 | 0.6161 | Yes | ||
| 16 | HMBS | 3833 | 1.980 | 0.6133 | Yes | ||
| 17 | TRIT1 | 3972 | 1.938 | 0.6344 | Yes | ||
| 18 | GSTM4 | 4661 | 1.711 | 0.6273 | No | ||
| 19 | GSTA3 | 5274 | 1.523 | 0.6209 | No | ||
| 20 | SRM | 6325 | 1.256 | 0.5908 | No | ||
| 21 | GSTM5 | 7110 | 1.071 | 0.5702 | No | ||
| 22 | LTC4S | 8034 | 0.862 | 0.5403 | No | ||
| 23 | AGPS | 8073 | 0.852 | 0.5506 | No | ||
| 24 | MGST3 | 8755 | 0.697 | 0.5294 | No | ||
| 25 | MGST2 | 11734 | -0.137 | 0.3955 | No | ||
| 26 | GSTA2 | 11839 | -0.168 | 0.3931 | No | ||
| 27 | FNTA | 13475 | -0.640 | 0.3276 | No | ||
| 28 | MAT1A | 13574 | -0.669 | 0.3326 | No | ||
| 29 | GSTZ1 | 15673 | -1.227 | 0.2542 | No | ||
| 30 | PDSS2 | 18828 | -2.259 | 0.1423 | No |