Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | set01_ATM_minus_versus_SCID |
| Phenotype | NoPhenotypeAvailable |
| Upregulated in class | na_neg |
| GeneSet | CALMODULIN_BINDING |
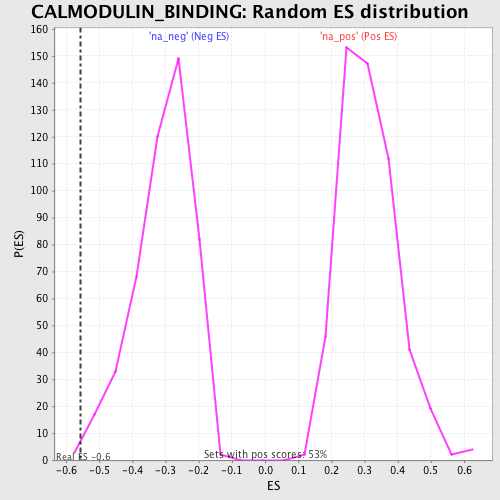
| Enrichment Score (ES) | -0.5560156 |
| Normalized Enrichment Score (NES) | -1.8048432 |
| Nominal p-value | 0.006329114 |
| FDR q-value | 0.9303552 |
| FWER p-Value | 0.895 |

| PROBE | GENE SYMBOL | GENE_TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|---|
| 1 | IQGAP1 | 211 | 7.562 | 0.0953 | No | ||
| 2 | MYO3A | 3302 | 2.181 | -0.0154 | No | ||
| 3 | MARCKS | 4285 | 1.832 | -0.0348 | No | ||
| 4 | TRPV4 | 5152 | 1.558 | -0.0527 | No | ||
| 5 | RIT2 | 5594 | 1.435 | -0.0529 | No | ||
| 6 | CAMKK2 | 5949 | 1.346 | -0.0504 | No | ||
| 7 | MYO6 | 6056 | 1.322 | -0.0369 | No | ||
| 8 | RGS4 | 7073 | 1.081 | -0.0682 | No | ||
| 9 | TRPV6 | 8943 | 0.652 | -0.1444 | No | ||
| 10 | RGS1 | 10104 | 0.353 | -0.1925 | No | ||
| 11 | MYO7A | 10297 | 0.301 | -0.1970 | No | ||
| 12 | NRGN | 10499 | 0.240 | -0.2029 | No | ||
| 13 | SPHK1 | 11395 | -0.030 | -0.2433 | No | ||
| 14 | IQCB1 | 11628 | -0.104 | -0.2524 | No | ||
| 15 | ATPIF1 | 16452 | -1.429 | -0.4526 | No | ||
| 16 | CACNA1C | 18169 | -1.984 | -0.5034 | No | ||
| 17 | TTN | 18906 | -2.297 | -0.5051 | No | ||
| 18 | RGS16 | 20024 | -2.907 | -0.5157 | Yes | ||
| 19 | CALD1 | 20032 | -2.916 | -0.4756 | Yes | ||
| 20 | STRN4 | 20086 | -2.958 | -0.4369 | Yes | ||
| 21 | DAPK2 | 20215 | -3.039 | -0.4006 | Yes | ||
| 22 | RIT1 | 20220 | -3.043 | -0.3586 | Yes | ||
| 23 | RGS2 | 21651 | -5.449 | -0.3482 | Yes | ||
| 24 | MYO9B | 21847 | -7.464 | -0.2536 | Yes | ||
| 25 | DAPK1 | 21946 | -18.608 | 0.0000 | Yes |