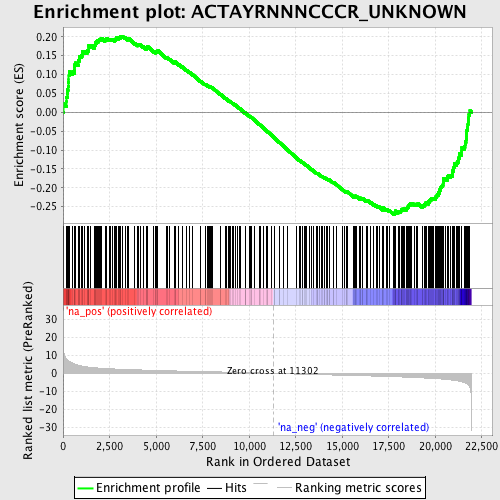
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | set01_ATM_minus_versus_SCID |
| Phenotype | NoPhenotypeAvailable |
| Upregulated in class | na_neg |
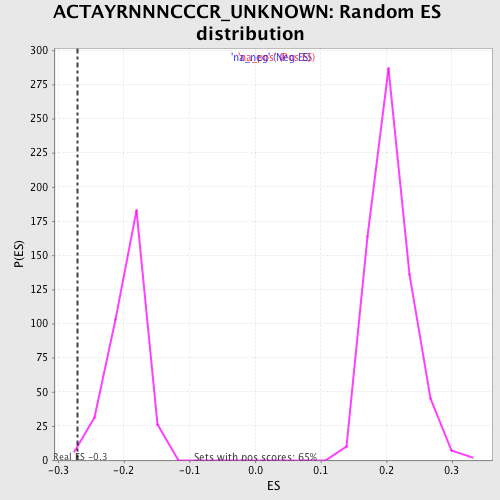
| GeneSet | ACTAYRNNNCCCR_UNKNOWN |
| Enrichment Score (ES) | -0.27058068 |
| Normalized Enrichment Score (NES) | -1.3882871 |
| Nominal p-value | 0.0057306592 |
| FDR q-value | 0.42089424 |
| FWER p-Value | 0.997 |

| PROBE | GENE SYMBOL | GENE_TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|---|
| 1 | SFRS2 | 12 | 15.098 | 0.0225 | No | ||
| 2 | ING3 | 170 | 8.243 | 0.0278 | No | ||
| 3 | PABPC1 | 176 | 8.119 | 0.0400 | No | ||
| 4 | ZNF24 | 222 | 7.510 | 0.0494 | No | ||
| 5 | PPP1R7 | 244 | 7.261 | 0.0595 | No | ||
| 6 | IMMT | 266 | 7.089 | 0.0693 | No | ||
| 7 | CYP39A1 | 298 | 6.839 | 0.0783 | No | ||
| 8 | LARS | 312 | 6.736 | 0.0880 | No | ||
| 9 | DNM1L | 315 | 6.724 | 0.0982 | No | ||
| 10 | PRIM1 | 324 | 6.659 | 0.1080 | No | ||
| 11 | NUP153 | 524 | 5.472 | 0.1071 | No | ||
| 12 | YWHAG | 608 | 5.096 | 0.1111 | No | ||
| 13 | ADSS | 623 | 5.028 | 0.1181 | No | ||
| 14 | NMT1 | 630 | 5.008 | 0.1255 | No | ||
| 15 | CBLL1 | 677 | 4.886 | 0.1308 | No | ||
| 16 | CDC26 | 825 | 4.484 | 0.1309 | No | ||
| 17 | HRSP12 | 826 | 4.484 | 0.1377 | No | ||
| 18 | DNAJC12 | 875 | 4.387 | 0.1422 | No | ||
| 19 | JARID2 | 891 | 4.340 | 0.1481 | No | ||
| 20 | COL4A3BP | 981 | 4.127 | 0.1503 | No | ||
| 21 | MRPL50 | 1019 | 4.046 | 0.1548 | No | ||
| 22 | BRCA1 | 1030 | 4.023 | 0.1605 | No | ||
| 23 | IPO7 | 1127 | 3.845 | 0.1619 | No | ||
| 24 | HSPB6 | 1287 | 3.578 | 0.1600 | No | ||
| 25 | RNF10 | 1326 | 3.522 | 0.1636 | No | ||
| 26 | PRCC | 1375 | 3.454 | 0.1667 | No | ||
| 27 | DMXL1 | 1377 | 3.451 | 0.1719 | No | ||
| 28 | M6PR | 1380 | 3.447 | 0.1771 | No | ||
| 29 | KDELC1 | 1487 | 3.326 | 0.1773 | No | ||
| 30 | LGMN | 1669 | 3.138 | 0.1737 | No | ||
| 31 | BIVM | 1687 | 3.125 | 0.1777 | No | ||
| 32 | TRUB1 | 1728 | 3.089 | 0.1806 | No | ||
| 33 | PA2G4 | 1729 | 3.087 | 0.1853 | No | ||
| 34 | FKBPL | 1802 | 3.027 | 0.1866 | No | ||
| 35 | HIRIP3 | 1842 | 2.988 | 0.1893 | No | ||
| 36 | ETFDH | 1914 | 2.928 | 0.1905 | No | ||
| 37 | UFD1L | 1946 | 2.891 | 0.1935 | No | ||
| 38 | STCH | 2026 | 2.832 | 0.1942 | No | ||
| 39 | TMEM4 | 2072 | 2.789 | 0.1964 | No | ||
| 40 | QTRTD1 | 2251 | 2.672 | 0.1922 | No | ||
| 41 | RPL15 | 2310 | 2.637 | 0.1936 | No | ||
| 42 | NFKBIB | 2348 | 2.618 | 0.1959 | No | ||
| 43 | RDH12 | 2482 | 2.551 | 0.1936 | No | ||
| 44 | FNTB | 2564 | 2.502 | 0.1937 | No | ||
| 45 | PGM2L1 | 2664 | 2.450 | 0.1929 | No | ||
| 46 | RPP21 | 2764 | 2.404 | 0.1920 | No | ||
| 47 | RPP40 | 2823 | 2.376 | 0.1929 | No | ||
| 48 | COX7C | 2840 | 2.371 | 0.1958 | No | ||
| 49 | KIAA0427 | 2875 | 2.355 | 0.1978 | No | ||
| 50 | TIAL1 | 2987 | 2.307 | 0.1962 | No | ||
| 51 | FTSJ1 | 3017 | 2.291 | 0.1984 | No | ||
| 52 | UBL3 | 3067 | 2.272 | 0.1996 | No | ||
| 53 | MRPL19 | 3105 | 2.258 | 0.2013 | No | ||
| 54 | WRN | 3171 | 2.230 | 0.2017 | No | ||
| 55 | SUFU | 3336 | 2.166 | 0.1975 | No | ||
| 56 | SLC29A2 | 3476 | 2.113 | 0.1943 | No | ||
| 57 | RABAC1 | 3504 | 2.100 | 0.1962 | No | ||
| 58 | GCN5L2 | 3858 | 1.973 | 0.1830 | No | ||
| 59 | ASXL2 | 4020 | 1.918 | 0.1785 | No | ||
| 60 | STXBP5 | 4039 | 1.914 | 0.1806 | No | ||
| 61 | GDAP1 | 4135 | 1.883 | 0.1790 | No | ||
| 62 | SETMAR | 4297 | 1.830 | 0.1744 | No | ||
| 63 | DNAJB12 | 4483 | 1.774 | 0.1686 | No | ||
| 64 | ITSN1 | 4501 | 1.769 | 0.1705 | No | ||
| 65 | FGFR1 | 4520 | 1.764 | 0.1724 | No | ||
| 66 | NAPB | 4544 | 1.753 | 0.1740 | No | ||
| 67 | MAP3K10 | 4869 | 1.647 | 0.1615 | No | ||
| 68 | AAK1 | 4974 | 1.615 | 0.1592 | No | ||
| 69 | CPEB3 | 4982 | 1.611 | 0.1613 | No | ||
| 70 | PSMC4 | 5016 | 1.600 | 0.1623 | No | ||
| 71 | CALR3 | 5082 | 1.579 | 0.1617 | No | ||
| 72 | PLEKHA1 | 5092 | 1.575 | 0.1637 | No | ||
| 73 | ZFP28 | 5541 | 1.449 | 0.1452 | No | ||
| 74 | CDCA3 | 5612 | 1.430 | 0.1441 | No | ||
| 75 | RAB3C | 5726 | 1.401 | 0.1411 | No | ||
| 76 | RALBP1 | 6008 | 1.332 | 0.1301 | No | ||
| 77 | DDX25 | 6012 | 1.331 | 0.1320 | No | ||
| 78 | BCKDK | 6019 | 1.330 | 0.1338 | No | ||
| 79 | UBE2N | 6220 | 1.280 | 0.1265 | No | ||
| 80 | RRAS | 6393 | 1.241 | 0.1205 | No | ||
| 81 | PARK2 | 6628 | 1.190 | 0.1115 | No | ||
| 82 | NUDT12 | 6775 | 1.153 | 0.1065 | No | ||
| 83 | GRWD1 | 6962 | 1.108 | 0.0996 | No | ||
| 84 | MRPL54 | 7378 | 1.009 | 0.0820 | No | ||
| 85 | SLC25A27 | 7406 | 1.004 | 0.0823 | No | ||
| 86 | NOS3 | 7624 | 0.956 | 0.0737 | No | ||
| 87 | SYT3 | 7653 | 0.948 | 0.0739 | No | ||
| 88 | SHMT2 | 7766 | 0.927 | 0.0701 | No | ||
| 89 | HBP1 | 7808 | 0.917 | 0.0697 | No | ||
| 90 | PRPF4 | 7888 | 0.897 | 0.0674 | No | ||
| 91 | RPS11 | 7903 | 0.894 | 0.0681 | No | ||
| 92 | RNASEH2A | 7983 | 0.875 | 0.0658 | No | ||
| 93 | MRPL34 | 8052 | 0.858 | 0.0640 | No | ||
| 94 | ARID5A | 8432 | 0.773 | 0.0476 | No | ||
| 95 | ARV1 | 8452 | 0.770 | 0.0479 | No | ||
| 96 | RAB8B | 8718 | 0.709 | 0.0368 | No | ||
| 97 | RDH10 | 8761 | 0.696 | 0.0359 | No | ||
| 98 | SDCCAG8 | 8911 | 0.658 | 0.0301 | No | ||
| 99 | BET1 | 8952 | 0.649 | 0.0292 | No | ||
| 100 | RPIA | 8988 | 0.641 | 0.0286 | No | ||
| 101 | VTI1B | 9109 | 0.613 | 0.0240 | No | ||
| 102 | RAB30 | 9131 | 0.607 | 0.0239 | No | ||
| 103 | ABLIM2 | 9245 | 0.583 | 0.0196 | No | ||
| 104 | MTR | 9343 | 0.561 | 0.0160 | No | ||
| 105 | CFL1 | 9464 | 0.533 | 0.0113 | No | ||
| 106 | CMAS | 9545 | 0.509 | 0.0083 | No | ||
| 107 | RABL3 | 9776 | 0.445 | -0.0016 | No | ||
| 108 | NR1D1 | 9787 | 0.441 | -0.0014 | No | ||
| 109 | HSPC171 | 9803 | 0.437 | -0.0014 | No | ||
| 110 | EIF2B4 | 9997 | 0.382 | -0.0097 | No | ||
| 111 | RNF121 | 10069 | 0.364 | -0.0124 | No | ||
| 112 | SFXN5 | 10096 | 0.355 | -0.0131 | No | ||
| 113 | ZFP1 | 10134 | 0.345 | -0.0143 | No | ||
| 114 | KCNN3 | 10298 | 0.301 | -0.0213 | No | ||
| 115 | ZFYVE19 | 10532 | 0.233 | -0.0317 | No | ||
| 116 | DCTN2 | 10568 | 0.223 | -0.0330 | No | ||
| 117 | LCMT1 | 10615 | 0.207 | -0.0348 | No | ||
| 118 | RPL6 | 10773 | 0.162 | -0.0418 | No | ||
| 119 | UPF3B | 10940 | 0.106 | -0.0493 | No | ||
| 120 | APBA3 | 11005 | 0.087 | -0.0521 | No | ||
| 121 | TBX6 | 11008 | 0.086 | -0.0521 | No | ||
| 122 | STMN1 | 11211 | 0.027 | -0.0614 | No | ||
| 123 | CYB561D2 | 11345 | -0.014 | -0.0675 | No | ||
| 124 | HSPB9 | 11349 | -0.015 | -0.0676 | No | ||
| 125 | RDH11 | 11621 | -0.103 | -0.0800 | No | ||
| 126 | SIRT2 | 11642 | -0.107 | -0.0807 | No | ||
| 127 | NTHL1 | 11842 | -0.170 | -0.0896 | No | ||
| 128 | TBN | 12045 | -0.224 | -0.0986 | No | ||
| 129 | FDPS | 12555 | -0.380 | -0.1215 | No | ||
| 130 | EXO1 | 12709 | -0.425 | -0.1279 | No | ||
| 131 | TRIM23 | 12741 | -0.434 | -0.1287 | No | ||
| 132 | PAFAH2 | 12842 | -0.462 | -0.1326 | No | ||
| 133 | TRIP10 | 12868 | -0.467 | -0.1331 | No | ||
| 134 | GSK3A | 12974 | -0.496 | -0.1371 | No | ||
| 135 | PACRG | 13009 | -0.507 | -0.1379 | No | ||
| 136 | USP5 | 13075 | -0.525 | -0.1401 | No | ||
| 137 | RAB1B | 13233 | -0.569 | -0.1465 | No | ||
| 138 | DPYSL5 | 13371 | -0.612 | -0.1519 | No | ||
| 139 | BANP | 13442 | -0.632 | -0.1542 | No | ||
| 140 | PPRC1 | 13636 | -0.681 | -0.1620 | No | ||
| 141 | FBXO15 | 13644 | -0.682 | -0.1613 | No | ||
| 142 | IPO13 | 13658 | -0.686 | -0.1609 | No | ||
| 143 | ADCK1 | 13757 | -0.713 | -0.1643 | No | ||
| 144 | ADAM22 | 13886 | -0.747 | -0.1691 | No | ||
| 145 | VKORC1 | 13935 | -0.761 | -0.1701 | No | ||
| 146 | ZCCHC8 | 14040 | -0.789 | -0.1737 | No | ||
| 147 | AAMP | 14068 | -0.796 | -0.1737 | No | ||
| 148 | ZNF8 | 14137 | -0.814 | -0.1756 | No | ||
| 149 | TLE4 | 14190 | -0.826 | -0.1768 | No | ||
| 150 | BTBD5 | 14217 | -0.830 | -0.1767 | No | ||
| 151 | MLLT7 | 14309 | -0.856 | -0.1796 | No | ||
| 152 | MAX | 14312 | -0.856 | -0.1784 | No | ||
| 153 | PPP2R5B | 14506 | -0.909 | -0.1859 | No | ||
| 154 | CTF1 | 14517 | -0.913 | -0.1850 | No | ||
| 155 | PPP6C | 14680 | -0.954 | -0.1910 | No | ||
| 156 | FANCG | 14990 | -1.045 | -0.2037 | No | ||
| 157 | WDFY2 | 15130 | -1.082 | -0.2084 | No | ||
| 158 | HDAC8 | 15218 | -1.106 | -0.2107 | No | ||
| 159 | PDE6D | 15234 | -1.111 | -0.2097 | No | ||
| 160 | RAB5C | 15288 | -1.125 | -0.2105 | No | ||
| 161 | POLK | 15603 | -1.208 | -0.2231 | No | ||
| 162 | DQX1 | 15680 | -1.229 | -0.2247 | No | ||
| 163 | ATP6V1F | 15684 | -1.230 | -0.2230 | No | ||
| 164 | ZNF235 | 15691 | -1.230 | -0.2214 | No | ||
| 165 | RUTBC3 | 15762 | -1.248 | -0.2227 | No | ||
| 166 | PET112L | 15941 | -1.296 | -0.2290 | No | ||
| 167 | PASK | 15978 | -1.305 | -0.2286 | No | ||
| 168 | CRYZL1 | 15983 | -1.307 | -0.2268 | No | ||
| 169 | MUS81 | 16064 | -1.326 | -0.2285 | No | ||
| 170 | HSD17B4 | 16098 | -1.333 | -0.2280 | No | ||
| 171 | SESN2 | 16304 | -1.390 | -0.2353 | No | ||
| 172 | RPL7 | 16326 | -1.397 | -0.2342 | No | ||
| 173 | SH3BGRL2 | 16354 | -1.405 | -0.2333 | No | ||
| 174 | SMARCAL1 | 16538 | -1.452 | -0.2395 | No | ||
| 175 | WASF1 | 16697 | -1.494 | -0.2445 | No | ||
| 176 | KCNE4 | 16816 | -1.530 | -0.2476 | No | ||
| 177 | ZFYVE1 | 16902 | -1.553 | -0.2492 | No | ||
| 178 | RDH14 | 16984 | -1.580 | -0.2505 | No | ||
| 179 | USP48 | 17178 | -1.641 | -0.2569 | No | ||
| 180 | WBP2 | 17180 | -1.642 | -0.2544 | No | ||
| 181 | FBXO9 | 17196 | -1.646 | -0.2526 | No | ||
| 182 | TRPC4AP | 17353 | -1.703 | -0.2572 | No | ||
| 183 | MITF | 17421 | -1.727 | -0.2577 | No | ||
| 184 | MDH1 | 17511 | -1.754 | -0.2591 | No | ||
| 185 | ACBD5 | 17761 | -1.840 | -0.2678 | Yes | ||
| 186 | UBE2H | 17821 | -1.857 | -0.2677 | Yes | ||
| 187 | SERTAD3 | 17846 | -1.868 | -0.2659 | Yes | ||
| 188 | THAP11 | 17859 | -1.872 | -0.2636 | Yes | ||
| 189 | MAF1 | 17868 | -1.877 | -0.2611 | Yes | ||
| 190 | TOP3A | 17995 | -1.918 | -0.2640 | Yes | ||
| 191 | CREBL1 | 18013 | -1.924 | -0.2619 | Yes | ||
| 192 | PPARGC1A | 18084 | -1.952 | -0.2621 | Yes | ||
| 193 | ABCC5 | 18155 | -1.979 | -0.2623 | Yes | ||
| 194 | COMMD9 | 18158 | -1.980 | -0.2594 | Yes | ||
| 195 | HCFC1R1 | 18195 | -1.994 | -0.2580 | Yes | ||
| 196 | OPHN1 | 18218 | -2.002 | -0.2560 | Yes | ||
| 197 | DPM3 | 18304 | -2.040 | -0.2568 | Yes | ||
| 198 | FKBP4 | 18329 | -2.051 | -0.2548 | Yes | ||
| 199 | CDC25A | 18430 | -2.090 | -0.2562 | Yes | ||
| 200 | GRK4 | 18459 | -2.101 | -0.2543 | Yes | ||
| 201 | POP1 | 18490 | -2.115 | -0.2524 | Yes | ||
| 202 | COVA1 | 18524 | -2.130 | -0.2507 | Yes | ||
| 203 | BRMS1L | 18546 | -2.138 | -0.2484 | Yes | ||
| 204 | INVS | 18576 | -2.147 | -0.2465 | Yes | ||
| 205 | PHF13 | 18585 | -2.149 | -0.2435 | Yes | ||
| 206 | ITPR1 | 18648 | -2.183 | -0.2431 | Yes | ||
| 207 | ESPL1 | 18665 | -2.189 | -0.2405 | Yes | ||
| 208 | SNX17 | 18745 | -2.223 | -0.2407 | Yes | ||
| 209 | CDC45L | 18866 | -2.277 | -0.2428 | Yes | ||
| 210 | PTDSS2 | 18974 | -2.325 | -0.2442 | Yes | ||
| 211 | ORMDL3 | 19023 | -2.349 | -0.2428 | Yes | ||
| 212 | VPS45A | 19065 | -2.373 | -0.2411 | Yes | ||
| 213 | ACTR1A | 19311 | -2.504 | -0.2486 | Yes | ||
| 214 | SKP2 | 19330 | -2.512 | -0.2456 | Yes | ||
| 215 | PANK3 | 19397 | -2.548 | -0.2447 | Yes | ||
| 216 | RPS6KB2 | 19448 | -2.572 | -0.2431 | Yes | ||
| 217 | FEN1 | 19469 | -2.588 | -0.2401 | Yes | ||
| 218 | BTRC | 19514 | -2.615 | -0.2381 | Yes | ||
| 219 | CDC6 | 19624 | -2.669 | -0.2391 | Yes | ||
| 220 | SNX16 | 19644 | -2.678 | -0.2359 | Yes | ||
| 221 | NKIRAS1 | 19691 | -2.704 | -0.2339 | Yes | ||
| 222 | SLC35B4 | 19715 | -2.719 | -0.2308 | Yes | ||
| 223 | PCM1 | 19770 | -2.751 | -0.2291 | Yes | ||
| 224 | NTAN1 | 19839 | -2.791 | -0.2279 | Yes | ||
| 225 | BUB1B | 19923 | -2.844 | -0.2274 | Yes | ||
| 226 | DTX2 | 20022 | -2.906 | -0.2275 | Yes | ||
| 227 | RAD51L1 | 20033 | -2.916 | -0.2235 | Yes | ||
| 228 | RANGAP1 | 20085 | -2.958 | -0.2214 | Yes | ||
| 229 | ZW10 | 20093 | -2.962 | -0.2172 | Yes | ||
| 230 | NUMA1 | 20161 | -3.004 | -0.2157 | Yes | ||
| 231 | TMOD3 | 20178 | -3.015 | -0.2118 | Yes | ||
| 232 | ZCCHC7 | 20235 | -3.050 | -0.2097 | Yes | ||
| 233 | PSMD11 | 20244 | -3.055 | -0.2054 | Yes | ||
| 234 | IRF2BP1 | 20284 | -3.087 | -0.2025 | Yes | ||
| 235 | PPP1R12B | 20296 | -3.097 | -0.1983 | Yes | ||
| 236 | SERPINI1 | 20341 | -3.130 | -0.1956 | Yes | ||
| 237 | TSC2 | 20379 | -3.156 | -0.1925 | Yes | ||
| 238 | TTC13 | 20415 | -3.182 | -0.1892 | Yes | ||
| 239 | BACE1 | 20416 | -3.182 | -0.1844 | Yes | ||
| 240 | LIG1 | 20430 | -3.201 | -0.1801 | Yes | ||
| 241 | PDCD10 | 20445 | -3.216 | -0.1758 | Yes | ||
| 242 | MAP2K6 | 20569 | -3.327 | -0.1764 | Yes | ||
| 243 | CDK9 | 20639 | -3.389 | -0.1744 | Yes | ||
| 244 | WDTC1 | 20643 | -3.392 | -0.1694 | Yes | ||
| 245 | HMG20A | 20718 | -3.467 | -0.1675 | Yes | ||
| 246 | SNX1 | 20823 | -3.591 | -0.1668 | Yes | ||
| 247 | TUSC4 | 20917 | -3.706 | -0.1655 | Yes | ||
| 248 | SOLH | 20928 | -3.715 | -0.1603 | Yes | ||
| 249 | TTLL4 | 20940 | -3.740 | -0.1551 | Yes | ||
| 250 | DFFA | 20981 | -3.791 | -0.1511 | Yes | ||
| 251 | IQGAP3 | 20994 | -3.812 | -0.1459 | Yes | ||
| 252 | SESN1 | 21012 | -3.838 | -0.1408 | Yes | ||
| 253 | ASXL1 | 21029 | -3.860 | -0.1356 | Yes | ||
| 254 | FBXW11 | 21133 | -4.021 | -0.1342 | Yes | ||
| 255 | PLEKHB1 | 21191 | -4.125 | -0.1306 | Yes | ||
| 256 | SMUG1 | 21228 | -4.195 | -0.1258 | Yes | ||
| 257 | AP2M1 | 21262 | -4.285 | -0.1208 | Yes | ||
| 258 | CDC40 | 21294 | -4.345 | -0.1156 | Yes | ||
| 259 | HACE1 | 21300 | -4.361 | -0.1092 | Yes | ||
| 260 | DHX30 | 21418 | -4.645 | -0.1075 | Yes | ||
| 261 | ZDHHC14 | 21422 | -4.658 | -0.1005 | Yes | ||
| 262 | SMCR8 | 21427 | -4.681 | -0.0936 | Yes | ||
| 263 | ITCH | 21548 | -5.062 | -0.0914 | Yes | ||
| 264 | SCAMP3 | 21595 | -5.212 | -0.0856 | Yes | ||
| 265 | PHF12 | 21607 | -5.257 | -0.0780 | Yes | ||
| 266 | VPS41 | 21650 | -5.441 | -0.0717 | Yes | ||
| 267 | TP53 | 21670 | -5.532 | -0.0641 | Yes | ||
| 268 | MAPBPIP | 21680 | -5.566 | -0.0560 | Yes | ||
| 269 | PURG | 21693 | -5.647 | -0.0480 | Yes | ||
| 270 | BUB3 | 21716 | -5.775 | -0.0402 | Yes | ||
| 271 | NUDT13 | 21751 | -6.095 | -0.0324 | Yes | ||
| 272 | AP2S1 | 21760 | -6.131 | -0.0235 | Yes | ||
| 273 | TNPO2 | 21787 | -6.441 | -0.0148 | Yes | ||
| 274 | MFN2 | 21806 | -6.718 | -0.0054 | Yes | ||
| 275 | NCOA3 | 21861 | -7.771 | 0.0040 | Yes |