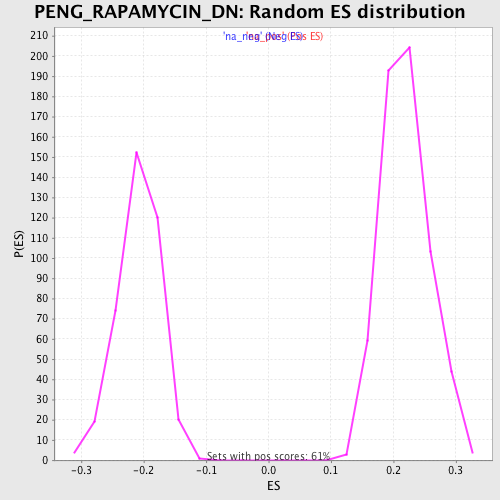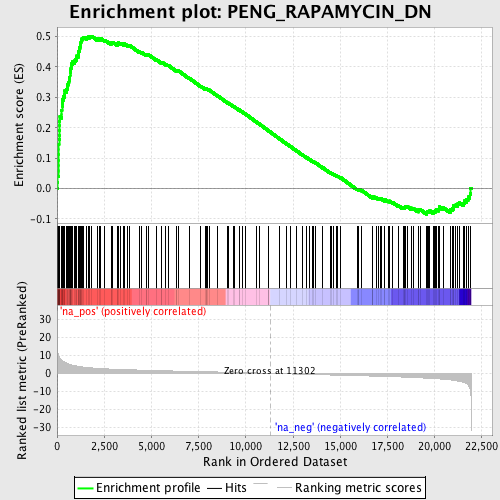
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | set01_ATM_minus_versus_SCID |
| Phenotype | NoPhenotypeAvailable |
| Upregulated in class | na_pos |
| GeneSet | PENG_RAPAMYCIN_DN |
| Enrichment Score (ES) | 0.5015362 |
| Normalized Enrichment Score (NES) | 2.2890663 |
| Nominal p-value | 0.0 |
| FDR q-value | 4.8903807E-4 |
| FWER p-Value | 0.0020 |

| PROBE | GENE SYMBOL | GENE_TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|---|
| 1 | EIF4G2 | 28 | 12.462 | 0.0208 | Yes | ||
| 2 | SYNCRIP | 38 | 11.556 | 0.0410 | Yes | ||
| 3 | PSMD14 | 67 | 10.195 | 0.0578 | Yes | ||
| 4 | AMD1 | 69 | 10.138 | 0.0757 | Yes | ||
| 5 | RABGGTB | 72 | 10.071 | 0.0935 | Yes | ||
| 6 | ETF1 | 77 | 9.911 | 0.1110 | Yes | ||
| 7 | DNAJA1 | 83 | 9.638 | 0.1279 | Yes | ||
| 8 | DDX21 | 85 | 9.586 | 0.1448 | Yes | ||
| 9 | DDX3X | 100 | 9.225 | 0.1606 | Yes | ||
| 10 | PSMC2 | 114 | 9.034 | 0.1760 | Yes | ||
| 11 | GSPT1 | 122 | 8.818 | 0.1914 | Yes | ||
| 12 | HSPA4 | 132 | 8.686 | 0.2064 | Yes | ||
| 13 | KPNA3 | 138 | 8.635 | 0.2215 | Yes | ||
| 14 | SF3A3 | 149 | 8.506 | 0.2361 | Yes | ||
| 15 | CTSC | 245 | 7.244 | 0.2446 | Yes | ||
| 16 | CCT6A | 247 | 7.235 | 0.2575 | Yes | ||
| 17 | CCT2 | 259 | 7.109 | 0.2696 | Yes | ||
| 18 | PITPNB | 267 | 7.084 | 0.2818 | Yes | ||
| 19 | CYCS | 295 | 6.886 | 0.2928 | Yes | ||
| 20 | HCCS | 338 | 6.560 | 0.3025 | Yes | ||
| 21 | CCT5 | 400 | 6.178 | 0.3107 | Yes | ||
| 22 | HSPA5 | 411 | 6.113 | 0.3211 | Yes | ||
| 23 | RARS | 509 | 5.559 | 0.3265 | Yes | ||
| 24 | CHUK | 537 | 5.425 | 0.3349 | Yes | ||
| 25 | ETFA | 567 | 5.262 | 0.3430 | Yes | ||
| 26 | CAPZA2 | 618 | 5.056 | 0.3496 | Yes | ||
| 27 | NMT1 | 630 | 5.008 | 0.3580 | Yes | ||
| 28 | TCP1 | 662 | 4.923 | 0.3653 | Yes | ||
| 29 | UBE2L3 | 682 | 4.865 | 0.3731 | Yes | ||
| 30 | CSE1L | 685 | 4.855 | 0.3816 | Yes | ||
| 31 | NARS | 697 | 4.826 | 0.3897 | Yes | ||
| 32 | IDH3A | 731 | 4.716 | 0.3966 | Yes | ||
| 33 | CCT4 | 750 | 4.659 | 0.4040 | Yes | ||
| 34 | EBNA1BP2 | 779 | 4.608 | 0.4109 | Yes | ||
| 35 | GOT2 | 827 | 4.480 | 0.4167 | Yes | ||
| 36 | MLF2 | 923 | 4.240 | 0.4199 | Yes | ||
| 37 | POLR2B | 966 | 4.154 | 0.4253 | Yes | ||
| 38 | SNRPA1 | 1035 | 4.018 | 0.4294 | Yes | ||
| 39 | ADRM1 | 1042 | 4.002 | 0.4362 | Yes | ||
| 40 | ATP6V0C | 1121 | 3.858 | 0.4395 | Yes | ||
| 41 | NUP98 | 1125 | 3.851 | 0.4462 | Yes | ||
| 42 | IFI30 | 1135 | 3.829 | 0.4526 | Yes | ||
| 43 | SSBP1 | 1186 | 3.734 | 0.4569 | Yes | ||
| 44 | PSMD5 | 1197 | 3.724 | 0.4630 | Yes | ||
| 45 | TIMM17A | 1214 | 3.698 | 0.4689 | Yes | ||
| 46 | TFRC | 1245 | 3.642 | 0.4740 | Yes | ||
| 47 | DAP3 | 1259 | 3.622 | 0.4798 | Yes | ||
| 48 | EIF4G1 | 1269 | 3.611 | 0.4858 | Yes | ||
| 49 | PPP2R2A | 1272 | 3.605 | 0.4921 | Yes | ||
| 50 | MORF4L2 | 1347 | 3.499 | 0.4949 | Yes | ||
| 51 | SLC29A1 | 1411 | 3.419 | 0.4981 | Yes | ||
| 52 | GORASP2 | 1561 | 3.250 | 0.4970 | Yes | ||
| 53 | IDI1 | 1642 | 3.162 | 0.4990 | Yes | ||
| 54 | PA2G4 | 1729 | 3.087 | 0.5005 | Yes | ||
| 55 | PTRF | 1824 | 3.001 | 0.5015 | Yes | ||
| 56 | TARS | 2116 | 2.760 | 0.4931 | No | ||
| 57 | CTPS | 2231 | 2.682 | 0.4926 | No | ||
| 58 | PTDSS1 | 2301 | 2.640 | 0.4941 | No | ||
| 59 | NOL5A | 2528 | 2.524 | 0.4882 | No | ||
| 60 | CHERP | 2858 | 2.364 | 0.4773 | No | ||
| 61 | C1QBP | 2898 | 2.343 | 0.4797 | No | ||
| 62 | COPB2 | 2945 | 2.324 | 0.4817 | No | ||
| 63 | BYSL | 3202 | 2.218 | 0.4739 | No | ||
| 64 | EIF3S9 | 3206 | 2.218 | 0.4777 | No | ||
| 65 | HYOU1 | 3243 | 2.203 | 0.4799 | No | ||
| 66 | SLC39A14 | 3367 | 2.153 | 0.4781 | No | ||
| 67 | MAPKAPK3 | 3529 | 2.090 | 0.4744 | No | ||
| 68 | PSMD8 | 3556 | 2.078 | 0.4769 | No | ||
| 69 | AK2 | 3741 | 2.013 | 0.4721 | No | ||
| 70 | GCN5L2 | 3858 | 1.973 | 0.4702 | No | ||
| 71 | CCNC | 4378 | 1.805 | 0.4496 | No | ||
| 72 | CDK4 | 4473 | 1.777 | 0.4485 | No | ||
| 73 | EIF2B2 | 4732 | 1.688 | 0.4396 | No | ||
| 74 | ATP5G3 | 4754 | 1.681 | 0.4416 | No | ||
| 75 | UCHL1 | 4845 | 1.656 | 0.4404 | No | ||
| 76 | SCD | 5271 | 1.524 | 0.4236 | No | ||
| 77 | ATP1B3 | 5534 | 1.451 | 0.4142 | No | ||
| 78 | ARMET | 5556 | 1.444 | 0.4158 | No | ||
| 79 | RTCD1 | 5765 | 1.390 | 0.4087 | No | ||
| 80 | BTK | 5881 | 1.361 | 0.4058 | No | ||
| 81 | SRM | 6325 | 1.256 | 0.3877 | No | ||
| 82 | PPIF | 6348 | 1.250 | 0.3889 | No | ||
| 83 | MYCBP | 6407 | 1.239 | 0.3885 | No | ||
| 84 | P2RX5 | 6997 | 1.101 | 0.3634 | No | ||
| 85 | MRPS12 | 7608 | 0.960 | 0.3370 | No | ||
| 86 | MRPL12 | 7870 | 0.902 | 0.3267 | No | ||
| 87 | NDUFS3 | 7890 | 0.897 | 0.3274 | No | ||
| 88 | JTB | 7895 | 0.895 | 0.3288 | No | ||
| 89 | S100A4 | 7981 | 0.876 | 0.3264 | No | ||
| 90 | AHSA1 | 8059 | 0.856 | 0.3244 | No | ||
| 91 | NP | 8522 | 0.755 | 0.3045 | No | ||
| 92 | ATF5 | 9046 | 0.628 | 0.2816 | No | ||
| 93 | RNU3IP2 | 9073 | 0.621 | 0.2816 | No | ||
| 94 | NOL1 | 9335 | 0.562 | 0.2706 | No | ||
| 95 | YARS | 9400 | 0.548 | 0.2686 | No | ||
| 96 | MARS | 9639 | 0.484 | 0.2585 | No | ||
| 97 | WBSCR22 | 9641 | 0.484 | 0.2593 | No | ||
| 98 | NXF1 | 9812 | 0.435 | 0.2523 | No | ||
| 99 | EIF2B4 | 9997 | 0.382 | 0.2445 | No | ||
| 100 | RPS26 | 10576 | 0.220 | 0.2184 | No | ||
| 101 | GTF2A2 | 10711 | 0.183 | 0.2125 | No | ||
| 102 | EBP | 11179 | 0.036 | 0.1912 | No | ||
| 103 | SQLE | 11220 | 0.023 | 0.1894 | No | ||
| 104 | TUBG1 | 11222 | 0.021 | 0.1894 | No | ||
| 105 | FASN | 11773 | -0.148 | 0.1644 | No | ||
| 106 | CYC1 | 12180 | -0.268 | 0.1462 | No | ||
| 107 | ASNS | 12348 | -0.318 | 0.1391 | No | ||
| 108 | IDH3B | 12680 | -0.415 | 0.1246 | No | ||
| 109 | PFKM | 13013 | -0.507 | 0.1103 | No | ||
| 110 | LDHB | 13230 | -0.568 | 0.1014 | No | ||
| 111 | POLR2F | 13365 | -0.610 | 0.0963 | No | ||
| 112 | PSMB5 | 13523 | -0.657 | 0.0903 | No | ||
| 113 | SLC7A5 | 13601 | -0.673 | 0.0879 | No | ||
| 114 | MMP15 | 13690 | -0.695 | 0.0851 | No | ||
| 115 | PIP5K1A | 14085 | -0.800 | 0.0684 | No | ||
| 116 | USP10 | 14462 | -0.896 | 0.0528 | No | ||
| 117 | RNASEH1 | 14538 | -0.916 | 0.0509 | No | ||
| 118 | NME1 | 14665 | -0.949 | 0.0468 | No | ||
| 119 | UBE2M | 14802 | -0.987 | 0.0423 | No | ||
| 120 | ATP6V0E | 14865 | -1.005 | 0.0413 | No | ||
| 121 | ACVRL1 | 15021 | -1.052 | 0.0360 | No | ||
| 122 | ARF1 | 15903 | -1.285 | -0.0021 | No | ||
| 123 | EXTL3 | 15987 | -1.308 | -0.0036 | No | ||
| 124 | PRDX3 | 15992 | -1.309 | -0.0015 | No | ||
| 125 | ACAT2 | 16138 | -1.344 | -0.0058 | No | ||
| 126 | ICT1 | 16706 | -1.499 | -0.0291 | No | ||
| 127 | WNT10B | 16710 | -1.500 | -0.0266 | No | ||
| 128 | COX11 | 16733 | -1.507 | -0.0250 | No | ||
| 129 | EEF1E1 | 16910 | -1.555 | -0.0303 | No | ||
| 130 | GCDH | 17033 | -1.595 | -0.0330 | No | ||
| 131 | SLC3A2 | 17131 | -1.624 | -0.0346 | No | ||
| 132 | BCS1L | 17163 | -1.635 | -0.0331 | No | ||
| 133 | EMP3 | 17364 | -1.707 | -0.0393 | No | ||
| 134 | GLO1 | 17371 | -1.709 | -0.0365 | No | ||
| 135 | SAFB | 17541 | -1.765 | -0.0412 | No | ||
| 136 | SDHB | 17604 | -1.785 | -0.0408 | No | ||
| 137 | BMP1 | 17784 | -1.846 | -0.0458 | No | ||
| 138 | GM2A | 18081 | -1.952 | -0.0559 | No | ||
| 139 | FKBP4 | 18329 | -2.051 | -0.0636 | No | ||
| 140 | ID2 | 18403 | -2.081 | -0.0633 | No | ||
| 141 | CDC25A | 18430 | -2.090 | -0.0607 | No | ||
| 142 | WDR18 | 18461 | -2.102 | -0.0584 | No | ||
| 143 | PPIB | 18556 | -2.141 | -0.0589 | No | ||
| 144 | BCAP31 | 18752 | -2.226 | -0.0639 | No | ||
| 145 | RBBP6 | 18884 | -2.286 | -0.0659 | No | ||
| 146 | SREBF1 | 19131 | -2.408 | -0.0729 | No | ||
| 147 | PTPN1 | 19166 | -2.425 | -0.0701 | No | ||
| 148 | HADH2 | 19245 | -2.472 | -0.0693 | No | ||
| 149 | TRIP13 | 19594 | -2.655 | -0.0806 | No | ||
| 150 | INPP5E | 19600 | -2.659 | -0.0761 | No | ||
| 151 | TIMM44 | 19667 | -2.694 | -0.0743 | No | ||
| 152 | XPO6 | 19753 | -2.740 | -0.0734 | No | ||
| 153 | PTBP1 | 19927 | -2.846 | -0.0763 | No | ||
| 154 | LIMK2 | 20017 | -2.904 | -0.0752 | No | ||
| 155 | GTF3C1 | 20066 | -2.936 | -0.0722 | No | ||
| 156 | DHCR24 | 20099 | -2.965 | -0.0684 | No | ||
| 157 | XBP1 | 20223 | -3.044 | -0.0686 | No | ||
| 158 | PSMD11 | 20244 | -3.055 | -0.0641 | No | ||
| 159 | NDUFA9 | 20264 | -3.066 | -0.0596 | No | ||
| 160 | PICALM | 20475 | -3.247 | -0.0634 | No | ||
| 161 | TOMM40 | 20829 | -3.600 | -0.0732 | No | ||
| 162 | AATF | 20859 | -3.643 | -0.0681 | No | ||
| 163 | CALU | 20942 | -3.741 | -0.0652 | No | ||
| 164 | RTN4 | 20996 | -3.813 | -0.0609 | No | ||
| 165 | RAB14 | 21014 | -3.839 | -0.0549 | No | ||
| 166 | RAE1 | 21136 | -4.030 | -0.0533 | No | ||
| 167 | RAB1A | 21240 | -4.235 | -0.0505 | No | ||
| 168 | HSPE1 | 21303 | -4.365 | -0.0456 | No | ||
| 169 | PAPOLA | 21551 | -5.071 | -0.0479 | No | ||
| 170 | PHB | 21598 | -5.236 | -0.0407 | No | ||
| 171 | TGDS | 21699 | -5.663 | -0.0352 | No | ||
| 172 | ALDH4A1 | 21776 | -6.315 | -0.0275 | No | ||
| 173 | SLC1A5 | 21879 | -8.578 | -0.0170 | No | ||
| 174 | CD47 | 21918 | -11.277 | 0.0013 | No |