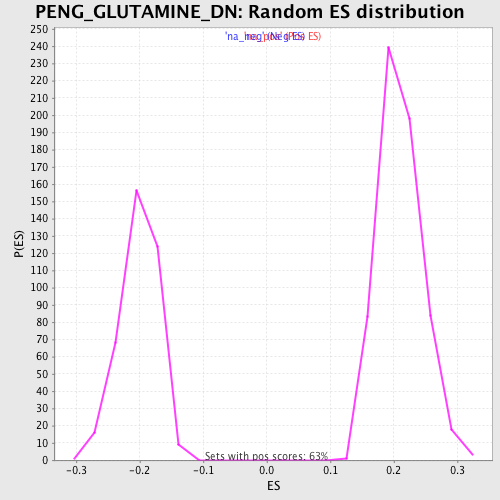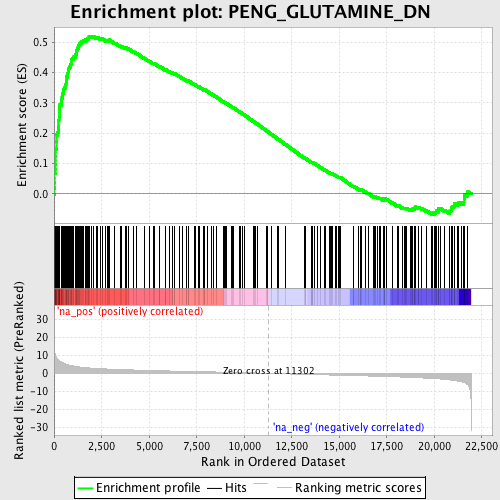
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | set01_ATM_minus_versus_SCID |
| Phenotype | NoPhenotypeAvailable |
| Upregulated in class | na_pos |
| GeneSet | PENG_GLUTAMINE_DN |
| Enrichment Score (ES) | 0.5196777 |
| Normalized Enrichment Score (NES) | 2.47829 |
| Nominal p-value | 0.0 |
| FDR q-value | 6.1129755E-4 |
| FWER p-Value | 0.0010 |

| PROBE | GENE SYMBOL | GENE_TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|---|
| 1 | SFRS2 | 12 | 15.098 | 0.0201 | Yes | ||
| 2 | EIF4G2 | 28 | 12.462 | 0.0365 | Yes | ||
| 3 | NCL | 36 | 11.624 | 0.0520 | Yes | ||
| 4 | SYNCRIP | 38 | 11.556 | 0.0678 | Yes | ||
| 5 | XRCC5 | 53 | 10.783 | 0.0819 | Yes | ||
| 6 | HNRPA2B1 | 59 | 10.466 | 0.0960 | Yes | ||
| 7 | AMD1 | 69 | 10.138 | 0.1094 | Yes | ||
| 8 | RABGGTB | 72 | 10.071 | 0.1231 | Yes | ||
| 9 | DNAJA1 | 83 | 9.638 | 0.1358 | Yes | ||
| 10 | DDX21 | 85 | 9.586 | 0.1489 | Yes | ||
| 11 | PSMD1 | 121 | 8.878 | 0.1595 | Yes | ||
| 12 | GSPT1 | 122 | 8.818 | 0.1715 | Yes | ||
| 13 | HSPA4 | 132 | 8.686 | 0.1830 | Yes | ||
| 14 | PSME4 | 133 | 8.679 | 0.1949 | Yes | ||
| 15 | PSMD12 | 183 | 8.008 | 0.2035 | Yes | ||
| 16 | EIF5 | 212 | 7.562 | 0.2126 | Yes | ||
| 17 | GMPS | 230 | 7.429 | 0.2220 | Yes | ||
| 18 | CTSC | 245 | 7.244 | 0.2312 | Yes | ||
| 19 | CCT6A | 247 | 7.235 | 0.2411 | Yes | ||
| 20 | CCT2 | 259 | 7.109 | 0.2503 | Yes | ||
| 21 | HPRT1 | 264 | 7.095 | 0.2598 | Yes | ||
| 22 | NUP160 | 274 | 7.038 | 0.2690 | Yes | ||
| 23 | SSB | 280 | 6.995 | 0.2784 | Yes | ||
| 24 | CYCS | 295 | 6.886 | 0.2872 | Yes | ||
| 25 | SMARCC1 | 305 | 6.770 | 0.2960 | Yes | ||
| 26 | DHX9 | 379 | 6.306 | 0.3013 | Yes | ||
| 27 | CCT5 | 400 | 6.178 | 0.3088 | Yes | ||
| 28 | HSPA5 | 411 | 6.113 | 0.3167 | Yes | ||
| 29 | PRPS1 | 441 | 5.930 | 0.3235 | Yes | ||
| 30 | DDX1 | 455 | 5.854 | 0.3309 | Yes | ||
| 31 | XPO1 | 482 | 5.700 | 0.3375 | Yes | ||
| 32 | HNRPL | 512 | 5.557 | 0.3438 | Yes | ||
| 33 | ABCF1 | 522 | 5.498 | 0.3509 | Yes | ||
| 34 | NOLC1 | 581 | 5.207 | 0.3553 | Yes | ||
| 35 | HNRPAB | 616 | 5.059 | 0.3607 | Yes | ||
| 36 | NMT1 | 630 | 5.008 | 0.3669 | Yes | ||
| 37 | PAICS | 653 | 4.935 | 0.3727 | Yes | ||
| 38 | TCP1 | 662 | 4.923 | 0.3790 | Yes | ||
| 39 | TCEB1 | 663 | 4.921 | 0.3858 | Yes | ||
| 40 | CSE1L | 685 | 4.855 | 0.3914 | Yes | ||
| 41 | PSMF1 | 706 | 4.789 | 0.3971 | Yes | ||
| 42 | IDH3A | 731 | 4.716 | 0.4024 | Yes | ||
| 43 | DNAJB6 | 737 | 4.696 | 0.4086 | Yes | ||
| 44 | EBNA1BP2 | 779 | 4.608 | 0.4130 | Yes | ||
| 45 | DHX15 | 802 | 4.544 | 0.4182 | Yes | ||
| 46 | SFRS1 | 843 | 4.448 | 0.4225 | Yes | ||
| 47 | SRRM1 | 864 | 4.408 | 0.4276 | Yes | ||
| 48 | DDX18 | 922 | 4.243 | 0.4307 | Yes | ||
| 49 | MLF2 | 923 | 4.240 | 0.4365 | Yes | ||
| 50 | BZW1 | 926 | 4.235 | 0.4423 | Yes | ||
| 51 | ATP2A2 | 947 | 4.197 | 0.4471 | Yes | ||
| 52 | SNRPA1 | 1035 | 4.018 | 0.4486 | Yes | ||
| 53 | ADRM1 | 1042 | 4.002 | 0.4538 | Yes | ||
| 54 | NUP98 | 1125 | 3.851 | 0.4552 | Yes | ||
| 55 | IFI30 | 1135 | 3.829 | 0.4601 | Yes | ||
| 56 | BMS1L | 1173 | 3.762 | 0.4635 | Yes | ||
| 57 | SSBP1 | 1186 | 3.734 | 0.4681 | Yes | ||
| 58 | ME2 | 1189 | 3.731 | 0.4731 | Yes | ||
| 59 | TIMM17A | 1214 | 3.698 | 0.4770 | Yes | ||
| 60 | TFRC | 1245 | 3.642 | 0.4806 | Yes | ||
| 61 | EIF4G1 | 1269 | 3.611 | 0.4845 | Yes | ||
| 62 | PPP2R2A | 1272 | 3.605 | 0.4894 | Yes | ||
| 63 | COPS3 | 1315 | 3.533 | 0.4923 | Yes | ||
| 64 | MORF4L2 | 1347 | 3.499 | 0.4956 | Yes | ||
| 65 | SLC29A1 | 1411 | 3.419 | 0.4974 | Yes | ||
| 66 | BAZ1A | 1441 | 3.382 | 0.5007 | Yes | ||
| 67 | TOP1 | 1486 | 3.326 | 0.5032 | Yes | ||
| 68 | CALR | 1541 | 3.273 | 0.5052 | Yes | ||
| 69 | IDI1 | 1642 | 3.162 | 0.5049 | Yes | ||
| 70 | SFPQ | 1655 | 3.149 | 0.5087 | Yes | ||
| 71 | PA2G4 | 1729 | 3.087 | 0.5095 | Yes | ||
| 72 | MPHOSPH10 | 1754 | 3.068 | 0.5126 | Yes | ||
| 73 | SLC5A6 | 1826 | 2.999 | 0.5135 | Yes | ||
| 74 | PSMA6 | 1834 | 2.997 | 0.5172 | Yes | ||
| 75 | SNRPB2 | 1870 | 2.965 | 0.5197 | Yes | ||
| 76 | LAMB3 | 1964 | 2.874 | 0.5193 | No | ||
| 77 | CAD | 2065 | 2.796 | 0.5185 | No | ||
| 78 | CTPS | 2231 | 2.682 | 0.5146 | No | ||
| 79 | PTDSS1 | 2301 | 2.640 | 0.5150 | No | ||
| 80 | TARDBP | 2456 | 2.559 | 0.5115 | No | ||
| 81 | NOL5A | 2528 | 2.524 | 0.5116 | No | ||
| 82 | CHD4 | 2724 | 2.423 | 0.5060 | No | ||
| 83 | DYRK1A | 2783 | 2.396 | 0.5066 | No | ||
| 84 | UMPS | 2852 | 2.367 | 0.5067 | No | ||
| 85 | C1QBP | 2898 | 2.343 | 0.5078 | No | ||
| 86 | BYSL | 3202 | 2.218 | 0.4969 | No | ||
| 87 | SLC29A2 | 3476 | 2.113 | 0.4872 | No | ||
| 88 | PSMD8 | 3556 | 2.078 | 0.4864 | No | ||
| 89 | AK2 | 3741 | 2.013 | 0.4807 | No | ||
| 90 | GRIN2A | 3784 | 1.997 | 0.4815 | No | ||
| 91 | ALDOA | 3888 | 1.961 | 0.4794 | No | ||
| 92 | VDAC1 | 4182 | 1.868 | 0.4685 | No | ||
| 93 | METTL1 | 4334 | 1.816 | 0.4640 | No | ||
| 94 | ATP5G3 | 4754 | 1.681 | 0.4470 | No | ||
| 95 | NUDC | 5010 | 1.602 | 0.4375 | No | ||
| 96 | ZMPSTE24 | 5213 | 1.538 | 0.4303 | No | ||
| 97 | SCD | 5271 | 1.524 | 0.4297 | No | ||
| 98 | ATP1B3 | 5534 | 1.451 | 0.4197 | No | ||
| 99 | ARMET | 5556 | 1.444 | 0.4207 | No | ||
| 100 | ADAM19 | 5859 | 1.368 | 0.4086 | No | ||
| 101 | BTK | 5881 | 1.361 | 0.4095 | No | ||
| 102 | GJB1 | 6071 | 1.319 | 0.4026 | No | ||
| 103 | SLC19A1 | 6211 | 1.281 | 0.3980 | No | ||
| 104 | UBE2N | 6220 | 1.280 | 0.3994 | No | ||
| 105 | SRM | 6325 | 1.256 | 0.3963 | No | ||
| 106 | PPIF | 6348 | 1.250 | 0.3970 | No | ||
| 107 | PTPN6 | 6588 | 1.199 | 0.3876 | No | ||
| 108 | RNF126 | 6771 | 1.154 | 0.3808 | No | ||
| 109 | INHBC | 6969 | 1.107 | 0.3733 | No | ||
| 110 | SYNGR2 | 6987 | 1.103 | 0.3740 | No | ||
| 111 | LDHA | 7048 | 1.089 | 0.3727 | No | ||
| 112 | CCNB1 | 7091 | 1.075 | 0.3722 | No | ||
| 113 | EIF5A | 7362 | 1.015 | 0.3612 | No | ||
| 114 | PCBP1 | 7434 | 0.998 | 0.3593 | No | ||
| 115 | MRPS12 | 7608 | 0.960 | 0.3526 | No | ||
| 116 | IFRD2 | 7629 | 0.954 | 0.3530 | No | ||
| 117 | MRPL12 | 7870 | 0.902 | 0.3432 | No | ||
| 118 | PRPF4 | 7888 | 0.897 | 0.3437 | No | ||
| 119 | PRDX1 | 7900 | 0.895 | 0.3444 | No | ||
| 120 | AHSA1 | 8059 | 0.856 | 0.3383 | No | ||
| 121 | PSMD3 | 8256 | 0.811 | 0.3303 | No | ||
| 122 | PIK3R2 | 8374 | 0.789 | 0.3260 | No | ||
| 123 | NP | 8522 | 0.755 | 0.3203 | No | ||
| 124 | RUVBL2 | 8897 | 0.663 | 0.3040 | No | ||
| 125 | SGTA | 8969 | 0.646 | 0.3016 | No | ||
| 126 | PIK3R3 | 9002 | 0.639 | 0.3010 | No | ||
| 127 | RNU3IP2 | 9073 | 0.621 | 0.2986 | No | ||
| 128 | NOL1 | 9335 | 0.562 | 0.2874 | No | ||
| 129 | ALG3 | 9384 | 0.551 | 0.2859 | No | ||
| 130 | CELSR2 | 9425 | 0.542 | 0.2848 | No | ||
| 131 | MTHFD1 | 9742 | 0.456 | 0.2709 | No | ||
| 132 | ERH | 9802 | 0.437 | 0.2688 | No | ||
| 133 | MYD88 | 9889 | 0.412 | 0.2654 | No | ||
| 134 | EIF2B4 | 9997 | 0.382 | 0.2610 | No | ||
| 135 | PSMB3 | 10496 | 0.241 | 0.2384 | No | ||
| 136 | RQCD1 | 10542 | 0.231 | 0.2366 | No | ||
| 137 | RPS26 | 10576 | 0.220 | 0.2354 | No | ||
| 138 | DOK4 | 10715 | 0.182 | 0.2293 | No | ||
| 139 | EBP | 11179 | 0.036 | 0.2080 | No | ||
| 140 | SQLE | 11220 | 0.023 | 0.2062 | No | ||
| 141 | TUBG1 | 11222 | 0.021 | 0.2062 | No | ||
| 142 | PSMB6 | 11428 | -0.044 | 0.1968 | No | ||
| 143 | FASN | 11773 | -0.148 | 0.1812 | No | ||
| 144 | TUBA1 | 11825 | -0.163 | 0.1790 | No | ||
| 145 | CYC1 | 12180 | -0.268 | 0.1631 | No | ||
| 146 | SNRPF | 13159 | -0.548 | 0.1188 | No | ||
| 147 | SEC61B | 13223 | -0.566 | 0.1167 | No | ||
| 148 | LDHB | 13230 | -0.568 | 0.1172 | No | ||
| 149 | DDX48 | 13541 | -0.661 | 0.1038 | No | ||
| 150 | PDCD4 | 13563 | -0.666 | 0.1037 | No | ||
| 151 | CCND1 | 13583 | -0.671 | 0.1038 | No | ||
| 152 | PFN1 | 13606 | -0.674 | 0.1037 | No | ||
| 153 | MMP15 | 13690 | -0.695 | 0.1008 | No | ||
| 154 | HS6ST1 | 13873 | -0.744 | 0.0935 | No | ||
| 155 | POLR2C | 14008 | -0.781 | 0.0884 | No | ||
| 156 | RRS1 | 14233 | -0.835 | 0.0792 | No | ||
| 157 | EIF2B5 | 14284 | -0.848 | 0.0780 | No | ||
| 158 | IMPDH1 | 14480 | -0.900 | 0.0703 | No | ||
| 159 | RNASEH1 | 14538 | -0.916 | 0.0689 | No | ||
| 160 | LMNB1 | 14609 | -0.933 | 0.0670 | No | ||
| 161 | NME1 | 14665 | -0.949 | 0.0657 | No | ||
| 162 | UBE2M | 14802 | -0.987 | 0.0608 | No | ||
| 163 | IDH1 | 14882 | -1.010 | 0.0586 | No | ||
| 164 | SC4MOL | 14974 | -1.040 | 0.0558 | No | ||
| 165 | OGFR | 15023 | -1.052 | 0.0550 | No | ||
| 166 | ERF | 15068 | -1.064 | 0.0545 | No | ||
| 167 | CDC20 | 15732 | -1.240 | 0.0256 | No | ||
| 168 | TXNIP | 15765 | -1.249 | 0.0259 | No | ||
| 169 | BOP1 | 16009 | -1.314 | 0.0165 | No | ||
| 170 | GOLGA1 | 16104 | -1.334 | 0.0140 | No | ||
| 171 | ACAT2 | 16138 | -1.344 | 0.0143 | No | ||
| 172 | PWP2H | 16168 | -1.352 | 0.0148 | No | ||
| 173 | SRPK1 | 16374 | -1.408 | 0.0073 | No | ||
| 174 | RPLP2 | 16519 | -1.445 | 0.0026 | No | ||
| 175 | UBE2G2 | 16826 | -1.533 | -0.0094 | No | ||
| 176 | ILF2 | 16883 | -1.549 | -0.0098 | No | ||
| 177 | EEF1E1 | 16910 | -1.555 | -0.0089 | No | ||
| 178 | GCDH | 17033 | -1.595 | -0.0123 | No | ||
| 179 | HRAS | 17104 | -1.619 | -0.0133 | No | ||
| 180 | MRPL28 | 17150 | -1.632 | -0.0132 | No | ||
| 181 | MYST1 | 17322 | -1.690 | -0.0188 | No | ||
| 182 | EMP3 | 17364 | -1.707 | -0.0183 | No | ||
| 183 | PRDX4 | 17370 | -1.708 | -0.0162 | No | ||
| 184 | TGFB1 | 17374 | -1.712 | -0.0140 | No | ||
| 185 | ACLY | 17482 | -1.745 | -0.0165 | No | ||
| 186 | FDFT1 | 17798 | -1.850 | -0.0285 | No | ||
| 187 | ALG8 | 18069 | -1.949 | -0.0383 | No | ||
| 188 | SLC4A7 | 18116 | -1.964 | -0.0377 | No | ||
| 189 | FKBP4 | 18329 | -2.051 | -0.0447 | No | ||
| 190 | CDC25A | 18430 | -2.090 | -0.0464 | No | ||
| 191 | PPP1R10 | 18497 | -2.120 | -0.0466 | No | ||
| 192 | PPIB | 18556 | -2.141 | -0.0463 | No | ||
| 193 | BCAP31 | 18752 | -2.226 | -0.0522 | No | ||
| 194 | PFKP | 18804 | -2.250 | -0.0515 | No | ||
| 195 | EWSR1 | 18870 | -2.279 | -0.0514 | No | ||
| 196 | RBBP6 | 18884 | -2.286 | -0.0489 | No | ||
| 197 | ATR | 18966 | -2.321 | -0.0494 | No | ||
| 198 | ADCY3 | 18986 | -2.335 | -0.0471 | No | ||
| 199 | ARS2 | 18989 | -2.337 | -0.0440 | No | ||
| 200 | MRPL3 | 19010 | -2.344 | -0.0417 | No | ||
| 201 | MAPRE2 | 19161 | -2.422 | -0.0453 | No | ||
| 202 | FBL | 19202 | -2.447 | -0.0438 | No | ||
| 203 | SF1 | 19340 | -2.515 | -0.0467 | No | ||
| 204 | TRIP13 | 19594 | -2.655 | -0.0547 | No | ||
| 205 | FUS | 19872 | -2.810 | -0.0636 | No | ||
| 206 | TAP1 | 19931 | -2.847 | -0.0624 | No | ||
| 207 | LIMK2 | 20017 | -2.904 | -0.0623 | No | ||
| 208 | RANGAP1 | 20085 | -2.958 | -0.0613 | No | ||
| 209 | DHCR24 | 20099 | -2.965 | -0.0579 | No | ||
| 210 | STAT1 | 20111 | -2.972 | -0.0543 | No | ||
| 211 | XBP1 | 20223 | -3.044 | -0.0553 | No | ||
| 212 | PABPN1 | 20239 | -3.051 | -0.0518 | No | ||
| 213 | PSMD11 | 20244 | -3.055 | -0.0478 | No | ||
| 214 | SNRP70 | 20333 | -3.122 | -0.0476 | No | ||
| 215 | FKBP2 | 20548 | -3.305 | -0.0529 | No | ||
| 216 | YPEL1 | 20806 | -3.576 | -0.0599 | No | ||
| 217 | TOMM40 | 20829 | -3.600 | -0.0559 | No | ||
| 218 | RBM14 | 20888 | -3.671 | -0.0536 | No | ||
| 219 | DHCR7 | 20894 | -3.680 | -0.0488 | No | ||
| 220 | MRPL23 | 20923 | -3.710 | -0.0450 | No | ||
| 221 | PES1 | 20968 | -3.769 | -0.0419 | No | ||
| 222 | MAP3K11 | 21046 | -3.888 | -0.0401 | No | ||
| 223 | SACS | 21079 | -3.937 | -0.0362 | No | ||
| 224 | TBL3 | 21092 | -3.953 | -0.0313 | No | ||
| 225 | DNAJB1 | 21242 | -4.240 | -0.0324 | No | ||
| 226 | HSPE1 | 21303 | -4.365 | -0.0292 | No | ||
| 227 | NDUFS6 | 21424 | -4.666 | -0.0283 | No | ||
| 228 | CLIC1 | 21553 | -5.073 | -0.0273 | No | ||
| 229 | SURF5 | 21572 | -5.147 | -0.0211 | No | ||
| 230 | COG2 | 21573 | -5.147 | -0.0140 | No | ||
| 231 | PRKCBP1 | 21590 | -5.204 | -0.0077 | No | ||
| 232 | PHB | 21598 | -5.236 | -0.0008 | No | ||
| 233 | MAZ | 21733 | -5.947 | 0.0011 | No | ||
| 234 | KHSRP | 21778 | -6.332 | 0.0078 | No |