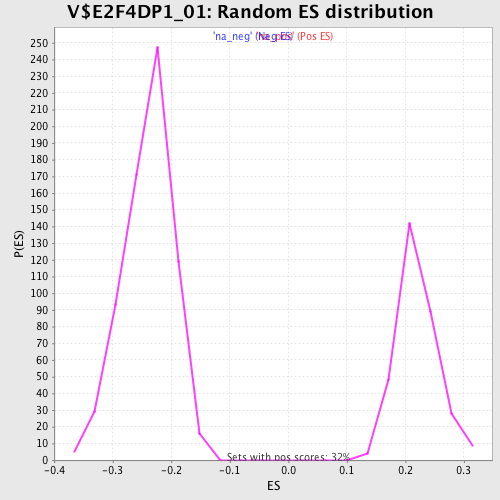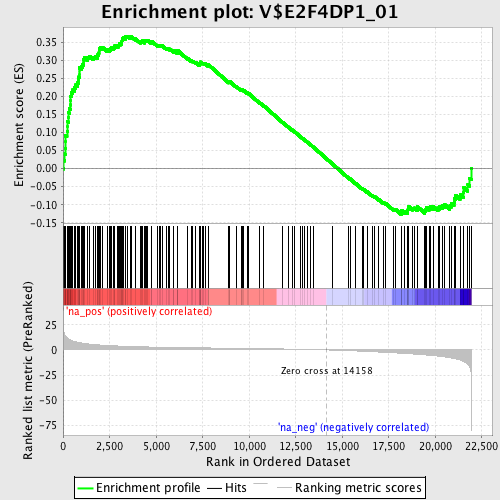
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | set01_ATM_minus_versus_ATM_plus |
| Phenotype | NoPhenotypeAvailable |
| Upregulated in class | na_pos |
| GeneSet | V$E2F4DP1_01 |
| Enrichment Score (ES) | 0.3671343 |
| Normalized Enrichment Score (NES) | 1.6693636 |
| Nominal p-value | 0.0 |
| FDR q-value | 0.017264955 |
| FWER p-Value | 0.289 |

| PROBE | GENE SYMBOL | GENE_TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|---|
| 1 | MCM6 | 26 | 19.246 | 0.0230 | Yes | ||
| 2 | RBBP7 | 70 | 15.794 | 0.0409 | Yes | ||
| 3 | RANBP1 | 110 | 14.273 | 0.0571 | Yes | ||
| 4 | SFRS7 | 119 | 13.984 | 0.0743 | Yes | ||
| 5 | NCL | 125 | 13.890 | 0.0915 | Yes | ||
| 6 | GMNN | 209 | 11.840 | 0.1026 | Yes | ||
| 7 | NOLC1 | 225 | 11.590 | 0.1165 | Yes | ||
| 8 | STAG1 | 231 | 11.525 | 0.1307 | Yes | ||
| 9 | RBL1 | 284 | 10.907 | 0.1421 | Yes | ||
| 10 | MCM4 | 293 | 10.788 | 0.1553 | Yes | ||
| 11 | RQCD1 | 333 | 10.476 | 0.1666 | Yes | ||
| 12 | DNAJC9 | 391 | 9.910 | 0.1765 | Yes | ||
| 13 | MCM7 | 398 | 9.851 | 0.1886 | Yes | ||
| 14 | ACBD6 | 406 | 9.712 | 0.2005 | Yes | ||
| 15 | SNRPD1 | 448 | 9.347 | 0.2104 | Yes | ||
| 16 | MRPL40 | 524 | 8.847 | 0.2180 | Yes | ||
| 17 | NASP | 620 | 8.327 | 0.2241 | Yes | ||
| 18 | TOPBP1 | 674 | 8.102 | 0.2319 | Yes | ||
| 19 | PRPS1 | 761 | 7.714 | 0.2377 | Yes | ||
| 20 | FKBP5 | 808 | 7.543 | 0.2450 | Yes | ||
| 21 | LIG1 | 820 | 7.498 | 0.2540 | Yes | ||
| 22 | MSH5 | 865 | 7.362 | 0.2612 | Yes | ||
| 23 | MYC | 869 | 7.330 | 0.2703 | Yes | ||
| 24 | SYNCRIP | 870 | 7.330 | 0.2795 | Yes | ||
| 25 | ATF5 | 998 | 6.881 | 0.2823 | Yes | ||
| 26 | HMGA1 | 1049 | 6.736 | 0.2885 | Yes | ||
| 27 | UNG | 1088 | 6.592 | 0.2950 | Yes | ||
| 28 | PEG3 | 1091 | 6.573 | 0.3032 | Yes | ||
| 29 | AP4M1 | 1145 | 6.445 | 0.3089 | Yes | ||
| 30 | E2F3 | 1330 | 5.964 | 0.3079 | Yes | ||
| 31 | E2F1 | 1414 | 5.788 | 0.3114 | Yes | ||
| 32 | CDCA7 | 1636 | 5.336 | 0.3080 | Yes | ||
| 33 | AK2 | 1717 | 5.217 | 0.3108 | Yes | ||
| 34 | MCM3 | 1864 | 4.991 | 0.3104 | Yes | ||
| 35 | ILF3 | 1868 | 4.988 | 0.3165 | Yes | ||
| 36 | FANCD2 | 1920 | 4.913 | 0.3204 | Yes | ||
| 37 | CLSPN | 1930 | 4.901 | 0.3261 | Yes | ||
| 38 | CDC6 | 1934 | 4.895 | 0.3322 | Yes | ||
| 39 | EZH2 | 2009 | 4.793 | 0.3348 | Yes | ||
| 40 | PHF5A | 2139 | 4.624 | 0.3347 | Yes | ||
| 41 | CDC25A | 2359 | 4.348 | 0.3301 | Yes | ||
| 42 | PCNA | 2502 | 4.211 | 0.3289 | Yes | ||
| 43 | IPO7 | 2553 | 4.153 | 0.3318 | Yes | ||
| 44 | PKMYT1 | 2581 | 4.124 | 0.3357 | Yes | ||
| 45 | PAX6 | 2693 | 4.022 | 0.3357 | Yes | ||
| 46 | POLD1 | 2748 | 3.984 | 0.3382 | Yes | ||
| 47 | NPR3 | 2783 | 3.965 | 0.3416 | Yes | ||
| 48 | POLD3 | 2921 | 3.858 | 0.3402 | Yes | ||
| 49 | SUV39H1 | 2996 | 3.804 | 0.3416 | Yes | ||
| 50 | NUP62 | 3002 | 3.797 | 0.3461 | Yes | ||
| 51 | NELL2 | 3085 | 3.745 | 0.3471 | Yes | ||
| 52 | HS6ST3 | 3148 | 3.695 | 0.3489 | Yes | ||
| 53 | MCM2 | 3157 | 3.689 | 0.3532 | Yes | ||
| 54 | EIF4A1 | 3164 | 3.687 | 0.3575 | Yes | ||
| 55 | CDC5L | 3204 | 3.652 | 0.3603 | Yes | ||
| 56 | HNRPA2B1 | 3240 | 3.624 | 0.3633 | Yes | ||
| 57 | POLE2 | 3339 | 3.542 | 0.3632 | Yes | ||
| 58 | SLC9A5 | 3367 | 3.520 | 0.3664 | Yes | ||
| 59 | DNMT1 | 3447 | 3.462 | 0.3671 | Yes | ||
| 60 | SFRS1 | 3608 | 3.355 | 0.3640 | No | ||
| 61 | SEZ6 | 3670 | 3.323 | 0.3654 | No | ||
| 62 | TYRO3 | 3877 | 3.216 | 0.3600 | No | ||
| 63 | PHF15 | 4144 | 3.073 | 0.3516 | No | ||
| 64 | ATAD2 | 4214 | 3.035 | 0.3523 | No | ||
| 65 | PAQR4 | 4246 | 3.021 | 0.3546 | No | ||
| 66 | CSPG2 | 4395 | 2.954 | 0.3516 | No | ||
| 67 | ASXL2 | 4402 | 2.952 | 0.3550 | No | ||
| 68 | HTF9C | 4461 | 2.927 | 0.3560 | No | ||
| 69 | TMPO | 4551 | 2.891 | 0.3556 | No | ||
| 70 | OLFML3 | 4725 | 2.825 | 0.3512 | No | ||
| 71 | MYH10 | 4767 | 2.808 | 0.3528 | No | ||
| 72 | FANCC | 5078 | 2.693 | 0.3420 | No | ||
| 73 | RET | 5159 | 2.665 | 0.3417 | No | ||
| 74 | SYNGR4 | 5243 | 2.641 | 0.3412 | No | ||
| 75 | AP1S1 | 5325 | 2.616 | 0.3407 | No | ||
| 76 | PCSK1 | 5578 | 2.532 | 0.3324 | No | ||
| 77 | STMN1 | 5657 | 2.509 | 0.3319 | No | ||
| 78 | MDGA1 | 5734 | 2.485 | 0.3316 | No | ||
| 79 | MAPT | 5946 | 2.415 | 0.3249 | No | ||
| 80 | EFNA5 | 5947 | 2.415 | 0.3280 | No | ||
| 81 | RRM2 | 6132 | 2.358 | 0.3225 | No | ||
| 82 | SEMA6A | 6142 | 2.353 | 0.3250 | No | ||
| 83 | PRPS2 | 6148 | 2.353 | 0.3277 | No | ||
| 84 | MCM8 | 6691 | 2.199 | 0.3056 | No | ||
| 85 | KCNS2 | 6899 | 2.140 | 0.2988 | No | ||
| 86 | MAP3K7 | 6975 | 2.124 | 0.2980 | No | ||
| 87 | MSH2 | 7101 | 2.090 | 0.2949 | No | ||
| 88 | ING3 | 7332 | 2.028 | 0.2869 | No | ||
| 89 | RPS6KA5 | 7337 | 2.027 | 0.2893 | No | ||
| 90 | RTBDN | 7360 | 2.021 | 0.2908 | No | ||
| 91 | KLF4 | 7361 | 2.021 | 0.2934 | No | ||
| 92 | ARHGAP6 | 7368 | 2.020 | 0.2956 | No | ||
| 93 | STAG2 | 7493 | 1.988 | 0.2924 | No | ||
| 94 | ADAMTS2 | 7554 | 1.973 | 0.2922 | No | ||
| 95 | GRIA4 | 7645 | 1.949 | 0.2905 | No | ||
| 96 | MAT2A | 7787 | 1.908 | 0.2864 | No | ||
| 97 | GABRB3 | 7813 | 1.903 | 0.2876 | No | ||
| 98 | CORT | 8912 | 1.645 | 0.2393 | No | ||
| 99 | HIRA | 8916 | 1.644 | 0.2412 | No | ||
| 100 | ARHGAP11A | 8940 | 1.640 | 0.2422 | No | ||
| 101 | ZIM2 | 9293 | 1.553 | 0.2280 | No | ||
| 102 | GATA1 | 9563 | 1.490 | 0.2176 | No | ||
| 103 | CBX3 | 9582 | 1.486 | 0.2186 | No | ||
| 104 | ONECUT1 | 9623 | 1.480 | 0.2186 | No | ||
| 105 | POU4F1 | 9704 | 1.460 | 0.2168 | No | ||
| 106 | RPS20 | 9885 | 1.414 | 0.2103 | No | ||
| 107 | HNRPD | 9966 | 1.392 | 0.2084 | No | ||
| 108 | DMD | 10568 | 1.245 | 0.1823 | No | ||
| 109 | LHX5 | 10793 | 1.188 | 0.1736 | No | ||
| 110 | NRK | 11809 | 0.902 | 0.1281 | No | ||
| 111 | YBX2 | 12113 | 0.808 | 0.1152 | No | ||
| 112 | KCNA6 | 12307 | 0.751 | 0.1073 | No | ||
| 113 | EPHB1 | 12428 | 0.716 | 0.1027 | No | ||
| 114 | HOXC10 | 12756 | 0.605 | 0.0884 | No | ||
| 115 | FHOD1 | 12880 | 0.559 | 0.0835 | No | ||
| 116 | FBXO5 | 12981 | 0.520 | 0.0795 | No | ||
| 117 | RASAL2 | 13152 | 0.456 | 0.0723 | No | ||
| 118 | PODN | 13271 | 0.409 | 0.0674 | No | ||
| 119 | PIM1 | 13451 | 0.338 | 0.0596 | No | ||
| 120 | FANCG | 14489 | -0.216 | 0.0123 | No | ||
| 121 | CASP8AP2 | 15356 | -0.775 | -0.0265 | No | ||
| 122 | MXD3 | 15419 | -0.824 | -0.0283 | No | ||
| 123 | EED | 15730 | -1.033 | -0.0413 | No | ||
| 124 | FMO4 | 16061 | -1.291 | -0.0548 | No | ||
| 125 | STK35 | 16140 | -1.357 | -0.0567 | No | ||
| 126 | PRKDC | 16339 | -1.517 | -0.0638 | No | ||
| 127 | BRMS1L | 16635 | -1.787 | -0.0751 | No | ||
| 128 | DNAJC5G | 16745 | -1.879 | -0.0778 | No | ||
| 129 | HCN3 | 16961 | -2.085 | -0.0850 | No | ||
| 130 | PHC1 | 17194 | -2.296 | -0.0928 | No | ||
| 131 | GPRC5B | 17315 | -2.402 | -0.0953 | No | ||
| 132 | USP37 | 17763 | -2.848 | -0.1122 | No | ||
| 133 | SOAT1 | 17864 | -2.959 | -0.1131 | No | ||
| 134 | IL4I1 | 18167 | -3.291 | -0.1228 | No | ||
| 135 | ALDH6A1 | 18168 | -3.291 | -0.1187 | No | ||
| 136 | ARID4A | 18192 | -3.324 | -0.1156 | No | ||
| 137 | SLC25A14 | 18364 | -3.495 | -0.1190 | No | ||
| 138 | RAVER1 | 18505 | -3.665 | -0.1208 | No | ||
| 139 | POLE4 | 18516 | -3.673 | -0.1167 | No | ||
| 140 | HNRPUL1 | 18530 | -3.691 | -0.1126 | No | ||
| 141 | CCNT1 | 18554 | -3.710 | -0.1090 | No | ||
| 142 | MAZ | 18575 | -3.730 | -0.1053 | No | ||
| 143 | PRPF4B | 18784 | -3.983 | -0.1098 | No | ||
| 144 | TBX6 | 18878 | -4.125 | -0.1089 | No | ||
| 145 | H2AFV | 19027 | -4.322 | -0.1102 | No | ||
| 146 | E2F7 | 19032 | -4.324 | -0.1050 | No | ||
| 147 | UGCGL1 | 19443 | -4.963 | -0.1176 | No | ||
| 148 | TLE3 | 19490 | -5.019 | -0.1134 | No | ||
| 149 | H2AFZ | 19510 | -5.048 | -0.1079 | No | ||
| 150 | ACO2 | 19662 | -5.276 | -0.1082 | No | ||
| 151 | HNRPR | 19749 | -5.403 | -0.1053 | No | ||
| 152 | ZCCHC8 | 19907 | -5.676 | -0.1054 | No | ||
| 153 | PPM1D | 20148 | -6.160 | -0.1087 | No | ||
| 154 | SUMO1 | 20226 | -6.338 | -0.1043 | No | ||
| 155 | PAPOLG | 20374 | -6.646 | -0.1026 | No | ||
| 156 | SP3 | 20510 | -6.981 | -0.1001 | No | ||
| 157 | POLR2A | 20787 | -7.778 | -0.1030 | No | ||
| 158 | KBTBD7 | 20863 | -8.031 | -0.0963 | No | ||
| 159 | MAPK6 | 21006 | -8.592 | -0.0920 | No | ||
| 160 | ID3 | 21036 | -8.725 | -0.0824 | No | ||
| 161 | USP52 | 21092 | -8.896 | -0.0737 | No | ||
| 162 | SFRS10 | 21328 | -10.041 | -0.0719 | No | ||
| 163 | PB1 | 21513 | -11.385 | -0.0660 | No | ||
| 164 | GSPT1 | 21530 | -11.532 | -0.0522 | No | ||
| 165 | SLCO3A1 | 21749 | -14.577 | -0.0439 | No | ||
| 166 | ZBTB4 | 21848 | -17.473 | -0.0265 | No | ||
| 167 | DCK | 21929 | -24.615 | 0.0008 | No |