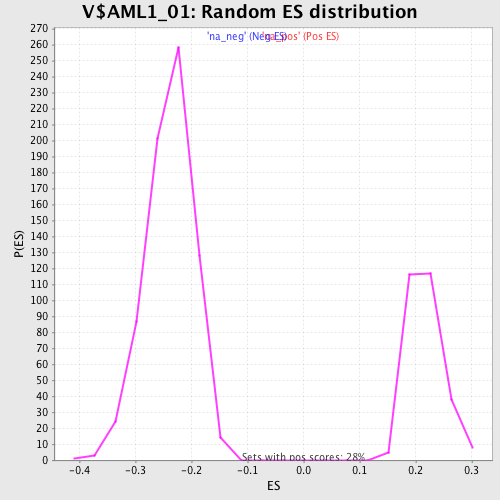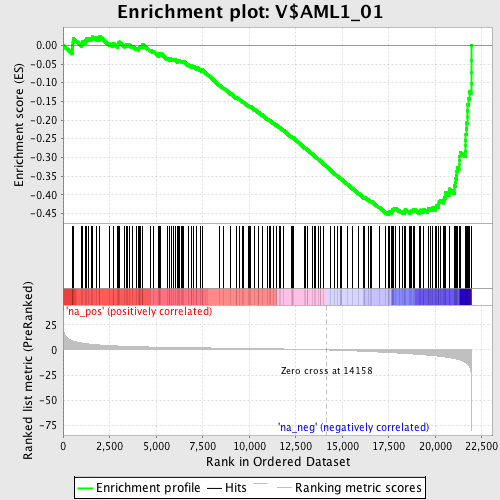
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | set01_ATM_minus_versus_ATM_plus |
| Phenotype | NoPhenotypeAvailable |
| Upregulated in class | na_neg |
| GeneSet | V$AML1_01 |
| Enrichment Score (ES) | -0.45292485 |
| Normalized Enrichment Score (NES) | -1.9074 |
| Nominal p-value | 0.0 |
| FDR q-value | 0.0055881594 |
| FWER p-Value | 0.035 |

| PROBE | GENE SYMBOL | GENE_TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|---|
| 1 | EIF2B3 | 483 | 9.147 | -0.0111 | No | ||
| 2 | HDGF | 511 | 8.966 | -0.0014 | No | ||
| 3 | EIF3S1 | 529 | 8.837 | 0.0086 | No | ||
| 4 | MTX1 | 561 | 8.626 | 0.0177 | No | ||
| 5 | PSMA1 | 1002 | 6.871 | 0.0058 | No | ||
| 6 | JDP2 | 1060 | 6.695 | 0.0114 | No | ||
| 7 | TPM1 | 1198 | 6.300 | 0.0127 | No | ||
| 8 | NDUFA9 | 1233 | 6.204 | 0.0187 | No | ||
| 9 | INPPL1 | 1375 | 5.849 | 0.0194 | No | ||
| 10 | ACSL5 | 1522 | 5.562 | 0.0194 | No | ||
| 11 | FCER1G | 1598 | 5.415 | 0.0226 | No | ||
| 12 | MRPS31 | 1790 | 5.089 | 0.0200 | No | ||
| 13 | ATP2A2 | 1935 | 4.894 | 0.0194 | No | ||
| 14 | CORO1C | 1947 | 4.883 | 0.0248 | No | ||
| 15 | SLC15A3 | 2498 | 4.213 | 0.0047 | No | ||
| 16 | ANK3 | 2682 | 4.033 | 0.0012 | No | ||
| 17 | PAX6 | 2693 | 4.022 | 0.0056 | No | ||
| 18 | RCOR2 | 2916 | 3.860 | 0.0001 | No | ||
| 19 | HOXC6 | 2951 | 3.830 | 0.0032 | No | ||
| 20 | AGTRL1 | 2978 | 3.814 | 0.0067 | No | ||
| 21 | RIN1 | 3026 | 3.784 | 0.0091 | No | ||
| 22 | TBX5 | 3322 | 3.557 | -0.0001 | No | ||
| 23 | PDZK1 | 3385 | 3.506 | 0.0013 | No | ||
| 24 | DAB1 | 3474 | 3.449 | 0.0015 | No | ||
| 25 | DNM3 | 3570 | 3.386 | 0.0012 | No | ||
| 26 | SOX5 | 3742 | 3.284 | -0.0026 | No | ||
| 27 | ANXA8 | 3933 | 3.183 | -0.0075 | No | ||
| 28 | SUPT16H | 4044 | 3.117 | -0.0088 | No | ||
| 29 | VRK1 | 4077 | 3.102 | -0.0064 | No | ||
| 30 | NUCB2 | 4119 | 3.087 | -0.0046 | No | ||
| 31 | GTF2A1 | 4142 | 3.074 | -0.0018 | No | ||
| 32 | AKAP3 | 4247 | 3.020 | -0.0029 | No | ||
| 33 | HLX1 | 4266 | 3.010 | -0.0001 | No | ||
| 34 | SLC6A9 | 4279 | 3.006 | 0.0030 | No | ||
| 35 | PTK9 | 4691 | 2.839 | -0.0124 | No | ||
| 36 | FOXD2 | 4834 | 2.782 | -0.0156 | No | ||
| 37 | CRTAC1 | 5127 | 2.678 | -0.0257 | No | ||
| 38 | POU2F1 | 5182 | 2.660 | -0.0250 | No | ||
| 39 | PRICKLE1 | 5195 | 2.655 | -0.0223 | No | ||
| 40 | CYP17A1 | 5257 | 2.637 | -0.0219 | No | ||
| 41 | NRXN3 | 5617 | 2.522 | -0.0353 | No | ||
| 42 | IGSF4 | 5733 | 2.485 | -0.0375 | No | ||
| 43 | SCEL | 5742 | 2.479 | -0.0349 | No | ||
| 44 | GPD1 | 5846 | 2.447 | -0.0366 | No | ||
| 45 | PGF | 5944 | 2.416 | -0.0382 | No | ||
| 46 | MOAP1 | 6014 | 2.391 | -0.0384 | No | ||
| 47 | BCAR3 | 6163 | 2.347 | -0.0424 | No | ||
| 48 | MYOD1 | 6186 | 2.340 | -0.0405 | No | ||
| 49 | IFNG | 6243 | 2.320 | -0.0403 | No | ||
| 50 | MAMDC1 | 6348 | 2.292 | -0.0423 | No | ||
| 51 | FGF4 | 6434 | 2.267 | -0.0434 | No | ||
| 52 | BTG4 | 6482 | 2.257 | -0.0428 | No | ||
| 53 | SLC2A3 | 6711 | 2.195 | -0.0506 | No | ||
| 54 | HOXC4 | 6893 | 2.143 | -0.0563 | No | ||
| 55 | ADAMTS4 | 6897 | 2.142 | -0.0539 | No | ||
| 56 | RAB39 | 7013 | 2.116 | -0.0566 | No | ||
| 57 | TNNT2 | 7176 | 2.071 | -0.0615 | No | ||
| 58 | RAB30 | 7183 | 2.069 | -0.0592 | No | ||
| 59 | ANKRD1 | 7387 | 2.014 | -0.0661 | No | ||
| 60 | PDE6H | 7465 | 1.995 | -0.0672 | No | ||
| 61 | KCNJ1 | 7476 | 1.993 | -0.0653 | No | ||
| 62 | LMO2 | 8415 | 1.764 | -0.1062 | No | ||
| 63 | THBS3 | 8639 | 1.707 | -0.1144 | No | ||
| 64 | PDZRN4 | 9008 | 1.623 | -0.1293 | No | ||
| 65 | COL9A2 | 9301 | 1.551 | -0.1408 | No | ||
| 66 | ARHGAP21 | 9330 | 1.544 | -0.1402 | No | ||
| 67 | TITF1 | 9457 | 1.515 | -0.1442 | No | ||
| 68 | PXN | 9661 | 1.471 | -0.1517 | No | ||
| 69 | KCTD4 | 9700 | 1.462 | -0.1517 | No | ||
| 70 | SCN3B | 9954 | 1.394 | -0.1616 | No | ||
| 71 | ATP6V0B | 10031 | 1.377 | -0.1634 | No | ||
| 72 | FGD4 | 10058 | 1.368 | -0.1630 | No | ||
| 73 | NGB | 10288 | 1.314 | -0.1719 | No | ||
| 74 | COL4A2 | 10494 | 1.263 | -0.1798 | No | ||
| 75 | ENTPD1 | 10518 | 1.255 | -0.1793 | No | ||
| 76 | SLITRK5 | 10686 | 1.218 | -0.1855 | No | ||
| 77 | TLL2 | 10997 | 1.137 | -0.1983 | No | ||
| 78 | PDE3B | 11073 | 1.116 | -0.2004 | No | ||
| 79 | RDH5 | 11117 | 1.104 | -0.2010 | No | ||
| 80 | SCN8A | 11312 | 1.051 | -0.2087 | No | ||
| 81 | ADAMTS8 | 11444 | 1.009 | -0.2135 | No | ||
| 82 | WNT8B | 11451 | 1.007 | -0.2125 | No | ||
| 83 | ZBTB8 | 11637 | 0.955 | -0.2199 | No | ||
| 84 | STRC | 11668 | 0.946 | -0.2201 | No | ||
| 85 | COL4A1 | 11838 | 0.893 | -0.2268 | No | ||
| 86 | ABCD4 | 12253 | 0.767 | -0.2448 | No | ||
| 87 | RGS18 | 12314 | 0.750 | -0.2467 | No | ||
| 88 | TNFSF4 | 12334 | 0.745 | -0.2467 | No | ||
| 89 | LY9 | 12378 | 0.730 | -0.2477 | No | ||
| 90 | CTSK | 12946 | 0.535 | -0.2731 | No | ||
| 91 | VIM | 13001 | 0.512 | -0.2750 | No | ||
| 92 | MMP13 | 13115 | 0.472 | -0.2796 | No | ||
| 93 | RASAL2 | 13152 | 0.456 | -0.2807 | No | ||
| 94 | NAV3 | 13380 | 0.365 | -0.2907 | No | ||
| 95 | MATN1 | 13529 | 0.304 | -0.2971 | No | ||
| 96 | IL23A | 13545 | 0.297 | -0.2975 | No | ||
| 97 | CLDN10 | 13708 | 0.223 | -0.3046 | No | ||
| 98 | MKRN3 | 13808 | 0.168 | -0.3090 | No | ||
| 99 | GZMB | 13821 | 0.160 | -0.3093 | No | ||
| 100 | RGS1 | 14005 | 0.079 | -0.3176 | No | ||
| 101 | LMO3 | 14365 | -0.133 | -0.3340 | No | ||
| 102 | ITGA10 | 14575 | -0.261 | -0.3433 | No | ||
| 103 | BATF | 14761 | -0.391 | -0.3513 | No | ||
| 104 | MPL | 14900 | -0.475 | -0.3570 | No | ||
| 105 | SCNN1A | 14906 | -0.481 | -0.3567 | No | ||
| 106 | BLOC1S2 | 14947 | -0.500 | -0.3579 | No | ||
| 107 | HIVEP3 | 14971 | -0.509 | -0.3584 | No | ||
| 108 | TACSTD2 | 15295 | -0.734 | -0.3723 | No | ||
| 109 | DKK1 | 15575 | -0.930 | -0.3840 | No | ||
| 110 | PLXNC1 | 15885 | -1.144 | -0.3968 | No | ||
| 111 | ARHGAP12 | 16162 | -1.379 | -0.4078 | No | ||
| 112 | LUZP1 | 16195 | -1.404 | -0.4076 | No | ||
| 113 | DDX23 | 16204 | -1.409 | -0.4062 | No | ||
| 114 | GRK5 | 16384 | -1.558 | -0.4125 | No | ||
| 115 | IBRDC3 | 16526 | -1.679 | -0.4170 | No | ||
| 116 | UBL3 | 16588 | -1.746 | -0.4176 | No | ||
| 117 | DIABLO | 16978 | -2.104 | -0.4330 | No | ||
| 118 | AP1G2 | 17340 | -2.429 | -0.4466 | No | ||
| 119 | RICS | 17479 | -2.581 | -0.4498 | Yes | ||
| 120 | BMF | 17491 | -2.595 | -0.4471 | Yes | ||
| 121 | RASGEF1A | 17514 | -2.616 | -0.4450 | Yes | ||
| 122 | ITGB7 | 17644 | -2.731 | -0.4476 | Yes | ||
| 123 | SLC37A4 | 17669 | -2.752 | -0.4453 | Yes | ||
| 124 | RHOG | 17692 | -2.775 | -0.4429 | Yes | ||
| 125 | MR1 | 17713 | -2.796 | -0.4405 | Yes | ||
| 126 | GRASP | 17775 | -2.863 | -0.4398 | Yes | ||
| 127 | TACC2 | 17778 | -2.868 | -0.4364 | Yes | ||
| 128 | PTPN22 | 17874 | -2.968 | -0.4371 | Yes | ||
| 129 | SIRT1 | 18076 | -3.186 | -0.4425 | Yes | ||
| 130 | GPATC2 | 18254 | -3.387 | -0.4465 | Yes | ||
| 131 | CD6 | 18354 | -3.478 | -0.4468 | Yes | ||
| 132 | ACIN1 | 18367 | -3.498 | -0.4431 | Yes | ||
| 133 | BLOC1S1 | 18395 | -3.528 | -0.4400 | Yes | ||
| 134 | S100A9 | 18637 | -3.794 | -0.4465 | Yes | ||
| 135 | NFRKB | 18682 | -3.857 | -0.4438 | Yes | ||
| 136 | TBK1 | 18737 | -3.926 | -0.4415 | Yes | ||
| 137 | SFRS4 | 18800 | -3.999 | -0.4395 | Yes | ||
| 138 | XCL1 | 18906 | -4.154 | -0.4393 | Yes | ||
| 139 | SNX1 | 19172 | -4.536 | -0.4459 | Yes | ||
| 140 | PCF11 | 19184 | -4.553 | -0.4409 | Yes | ||
| 141 | MAP3K11 | 19341 | -4.796 | -0.4422 | Yes | ||
| 142 | NR4A1 | 19386 | -4.872 | -0.4383 | Yes | ||
| 143 | MIZF | 19606 | -5.199 | -0.4420 | Yes | ||
| 144 | EIF2C1 | 19610 | -5.207 | -0.4358 | Yes | ||
| 145 | NHLRC2 | 19760 | -5.421 | -0.4361 | Yes | ||
| 146 | ARHGEF2 | 19872 | -5.614 | -0.4343 | Yes | ||
| 147 | CREM | 20025 | -5.922 | -0.4341 | Yes | ||
| 148 | KIF1B | 20048 | -5.972 | -0.4278 | Yes | ||
| 149 | RAB5B | 20175 | -6.220 | -0.4260 | Yes | ||
| 150 | EMP1 | 20196 | -6.261 | -0.4193 | Yes | ||
| 151 | ARNT | 20257 | -6.406 | -0.4143 | Yes | ||
| 152 | CHES1 | 20432 | -6.813 | -0.4140 | Yes | ||
| 153 | STAT2 | 20472 | -6.911 | -0.4074 | Yes | ||
| 154 | EDARADD | 20526 | -7.013 | -0.4013 | Yes | ||
| 155 | MMP14 | 20535 | -7.043 | -0.3931 | Yes | ||
| 156 | NOTCH2 | 20736 | -7.587 | -0.3930 | Yes | ||
| 157 | CAP1 | 20757 | -7.665 | -0.3846 | Yes | ||
| 158 | ID3 | 21036 | -8.725 | -0.3867 | Yes | ||
| 159 | ARF3 | 21039 | -8.729 | -0.3762 | Yes | ||
| 160 | FLI1 | 21086 | -8.875 | -0.3675 | Yes | ||
| 161 | JMJD1C | 21097 | -8.917 | -0.3571 | Yes | ||
| 162 | ATF7IP | 21131 | -9.063 | -0.3476 | Yes | ||
| 163 | NRAS | 21157 | -9.167 | -0.3376 | Yes | ||
| 164 | PACS1 | 21177 | -9.247 | -0.3272 | Yes | ||
| 165 | MADD | 21274 | -9.747 | -0.3197 | Yes | ||
| 166 | DACT1 | 21277 | -9.787 | -0.3079 | Yes | ||
| 167 | CUGBP1 | 21293 | -9.860 | -0.2966 | Yes | ||
| 168 | PICALM | 21330 | -10.065 | -0.2860 | Yes | ||
| 169 | CD69 | 21606 | -12.413 | -0.2835 | Yes | ||
| 170 | TCF12 | 21627 | -12.637 | -0.2690 | Yes | ||
| 171 | RAG1 | 21632 | -12.680 | -0.2538 | Yes | ||
| 172 | RORC | 21644 | -12.946 | -0.2385 | Yes | ||
| 173 | SEMA4A | 21668 | -13.174 | -0.2235 | Yes | ||
| 174 | SLC37A2 | 21685 | -13.574 | -0.2077 | Yes | ||
| 175 | CBL | 21707 | -13.898 | -0.1918 | Yes | ||
| 176 | SLC12A6 | 21710 | -13.946 | -0.1749 | Yes | ||
| 177 | DNAJC14 | 21734 | -14.348 | -0.1585 | Yes | ||
| 178 | LCK | 21778 | -15.166 | -0.1420 | Yes | ||
| 179 | HIPK1 | 21822 | -16.460 | -0.1239 | Yes | ||
| 180 | CRY1 | 21920 | -22.141 | -0.1014 | Yes | ||
| 181 | DGKA | 21928 | -24.490 | -0.0719 | Yes | ||
| 182 | EGR2 | 21933 | -25.685 | -0.0408 | Yes | ||
| 183 | SCN2B | 21941 | -34.007 | 0.0003 | Yes |